Transcriptional alterations in glioma result primarily from DNA methylation–independent mechanisms
- Franck Court1,
- Elisa Le Boiteux1,
- Anne Fogli1,2,
- Mélanie Müller-Barthélémy1,3,
- Catherine Vaurs-Barrière1,
- Emmanuel Chautard3,4,
- Bruno Pereira5,
- Julian Biau4,6,
- Jean-Louis Kemeny7,
- Toufic Khalil8,
- Lucie Karayan-Tapon9,10,11,
- Pierre Verrelle12,13,14 and
- Philippe Arnaud1
- 1Laboratoire Génétique Reproduction et Développement (GReD), Université Clermont Auvergne, CNRS, INSERM, BP 38, Clermont-Ferrand 63001, France;
- 2Biochemistry and Molecular Biology Department, Clermont-Ferrand Hospital, Clermont-Ferrand 63003, France;
- 3Pathology Department, Jean Perrin Center, Clermont-Ferrand 63011, France;
- 4Université Clermont Auvergne, INSERM, U1240 IMoST, Clermont-Ferrand 63011, France;
- 5Biostatistics Department, Délégation à la Recherche Clinique et à l'Innovation, Clermont-Ferrand Hospital, Clermont-Ferrand 63003, France;
- 6Radiotherapy Department, Jean Perrin Center, Clermont-Ferrand 63011, France;
- 7Pathology Department, Université Clermont Auvergne and Clermont-Ferrand Hospital, Clermont-Ferrand 63003, France;
- 8Department of Neurosurgery, Clermont-Ferrand Hospital, Clermont-Ferrand 63003, France;
- 9INSERM, U1084, Poitiers 86021, France;
- 10Poitiers University, Poitiers 86000, France;
- 11Department of Cancer Biology, Poitiers Hospital, Poitiers 86021, France;
- 12INSERM, U1196 CNRS UMR9187, Curie Institute, Orsay 91405, France;
- 13Radiotherapy Department Curie Institute, Paris 75005, France;
- 14Université Clermont Auvergne, Clermont-Ferrand 63000, France
Abstract
In cancer cells, aberrant DNA methylation is commonly associated with transcriptional alterations, including silencing of tumor suppressor genes. However, multiple epigenetic mechanisms, including polycomb repressive marks, contribute to gene deregulation in cancer. To dissect the relative contribution of DNA methylation–dependent and –independent mechanisms to transcriptional alterations at CpG island/promoter-associated genes in cancer, we studied 70 samples of adult glioma, a widespread type of brain tumor, classified according to their isocitrate dehydrogenase (IDH1) mutation status. We found that most transcriptional alterations in tumor samples were DNA methylation–independent. Instead, altered histone H3 trimethylation at lysine 27 (H3K27me3) was the predominant molecular defect at deregulated genes. Our results also suggest that the presence of a bivalent chromatin signature at CpG island promoters in stem cells predisposes not only to hypermethylation, as widely documented, but more generally to all types of transcriptional alterations in transformed cells. In addition, the gene expression strength in healthy brain cells influences the choice between DNA methylation- and H3K27me3-associated silencing in glioma. Highly expressed genes were more likely to be repressed by H3K27me3 than by DNA methylation. Our findings support a model in which altered H3K27me3 dynamics, more specifically defects in the interplay between polycomb protein complexes and the brain-specific transcriptional machinery, is the main cause of transcriptional alteration in glioma cells. Our study provides the first comprehensive description of epigenetic changes in glioma and their relative contribution to transcriptional changes. It may be useful for the design of drugs targeting cancer-related epigenetic defects.
Cancer is a complex disease that results from the disruption of key pathways, including those regulating cell survival and division. Besides genetic lesions, epigenetic alterations also contribute to tumorigenesis mainly by leading to abnormal gene expression (Flavahan et al. 2017).
Together with genome-wide DNA hypomethylation, DNA hypermethylation of CpG islands (CGIs) is a well-defined feature of cancer cells and is believed to be the main cause of aberrant gene repression (Baylin and Jones 2016). CGIs are key regulatory genomic regions of a few hundred base pairs in size characterized by high frequency of CpG dinucleotides. In humans, ∼70% of promoters are associated with CGIs that generally remain unmethylated during somatic development, regardless of the gene expression status (Deaton and Bird 2011). Conversely, it has been shown that in tumors, DNA hypermethylation of their CGI/promoter leads to aberrant silencing of some tumor suppressor genes, such as BRCA1 (Dobrovic and Simpfendorfer 1997), RB1 (Greger et al. 1994), and MLH1 (Herman et al. 1998). However, the primary role of this defect in the widespread cancer-associated genes silencing, and more broadly in cancer biology, is still being questioned. Indeed, an increasing number of studies have shown that in tumors, DNA hypermethylation affects primarily CGI/promoters that control genes already repressed in the matched normal tissue (Gal-Yam et al. 2008; Sproul et al. 2011, 2012; Hinoue et al. 2012; Court and Arnaud 2017). Moreover, in some tumor types, such as glioma or breast cancer, patients with a CpG island methylator phenotype (CIMP), a signature identified in various human malignancies and defined by the concomitant hypermethylation of multiple CGIs (Suzuki et al. 2014), have a better clinical prognosis compared with patients without CIMP (Noushmehr et al. 2010; Fang et al. 2011).
Therefore, other DNA methylation–independent epigenetic alterations at CGI/promoters might contribute to the genome-wide pattern of aberrant gene repression observed in cancer cells. During normal development, promoters/CGIs are dynamically marked by the permissive H3K4me3 and/or the repressive H3K27me3 histone marks (Mikkelsen et al. 2007). When in combination, these marks constitute the so-called bivalent chromatin signature that maintains genes (e.g., developmental genes in stem cells) repressed, but “poised” for activation, because the bivalent mark can resolve into either H3K4me3 or H3K27me3 (Bernstein et al. 2006). Alterations in the control of these chromatin signatures also could lead to gene silencing (Court and Arnaud 2017). This hypothesis is supported by the observation that genes encoding methyltransferases and demethylases that regulate H3K27me3 and H3K4me3 deposition, such as EZH2, KMT2 (MLL) family members, and KDM6A, are translocated or mutated and/or their expression is altered in many malignancies (Suvà et al. 2013; Dawson 2017). Such defects had been documented in a handful of studies. In detail, analyses in established prostate (Gal-Yam et al. 2008; Kondo et al. 2008) and urothelial (Dudziec et al. 2012) cancer cell lines and in colorectal tumor samples (Hahn et al. 2014) highlighted that gene silencing can be mediated just by H3K27me3.
Chromatin-based alterations could also lead to gain of gene expression in tumors. Hahn et al. (2014) showed that genes associated with GCI/promoters displaying a bivalent chromatin signature in normal colon can be ectopically expressed in the matched tumor samples following H3K27me3 loss. Moreover, Bert et al. (2013) identified in prostate cancer cell lines some genomic domains characterized by altered chromatin signatures associated with aberrant gene expression.
Therefore, it is clear that different epigenetic alterations at promoters/CGIs could contribute to the abnormal gene expression pattern of cancer cells. Because of the absence of dedicated integrative studies, many questions remain concerning the bases and extent of these alterations as well as their relative contribution to aberrant loss/gain of gene expression in cancer cells.
Glioma, which is derived from glial cells, is one of the most widespread brain tumor types. In 2007, the World Health Organization (WHO) classified gliomas in four grades (I–IV) according to their histology. Malignant anaplastic astrocytoma (a subset of the WHO grade III gliomas) and glioblastoma multiforme (GBM; WHO grade IV) account for about half of all gliomas, and are the most deadly and aggressive forms. The median survival time after diagnosis of patients with GBM does not exceed 18 mo despite the aggressive treatments. At the molecular level, aggressive gliomas are characterized by expression of wild-type isocitrate dehydrogenase (IDHwt) genes (IDH1 and IDH2), whereas gliomas with better prognosis express mutated IDH (IDHmut) (Cohen et al. 2013). Consequently, the recently released 2016 WHO classification of diffuse gliomas (Louis et al. 2016), which we used in this study, is primarily based on the IDH1 mutation status (IDHmut vs. IDHwt). IDH1 mutation results in CIMP-positive tumors (Turcan et al. 2012) with a better clinical prognosis compared with CIMP-negative tumors (Noushmehr et al. 2010; Cohen et al. 2013). Other epigenetic regulators also could be involved in glioma development/progression, for instance the polycomb repressors EZH2 and BMI1 (Häyry et al. 2008; Bruggeman et al. 2009; Suvà et al. 2009) and KMT2 family members that are mutated in a subset of GBM (Parsons et al. 2008; Brennan et al. 2013). Here, we used glioma as a model to investigate the molecular bases of transcriptional alterations of CGI/promoter-associated genes in cancer.
Results
CGI methylation poorly contributes to transcriptional alterations in glioma
For this study, we used 70 clinically well-characterized primary glioma samples (the patients’ demographic and main molecular and clinical features are provided in Supplemental Table S1). We classified samples according to their IDH1 status (n = 55 IDHwt, and n = 15 IDHmut). This first level of classification, upstream of the 1p/19q codeletion status according to the 2016 WHO classification (Louis et al. 2016), clearly discriminated two tumor classes relative to aggressiveness, with a significant survival advantage for patients with IDHmut tumors (HR = 0.32, 95% CI [0.14–0.71], P = 0.005) (Supplemental Table S1; Supplemental Fig. S1A).
Genome-wide analysis of DNA methylation at CGIs, using the Infinium HumanMethylation450 (HM450K) BeadChip Arrays, showed that DNA methylation defects were more widespread in IDHmut samples, constituting a Glioma CIMP (G-CIMP) subclass (Supplemental Fig. S1B). In agreement with the literature (Noushmehr et al. 2010; Turcan et al. 2012; Louis et al. 2016), in our cohort, aggressive gliomas were characterized by IDHwt and absence of G-CIMP, whereas IDHmut gliomas showed a G-CIMP profile and had a better prognosis.
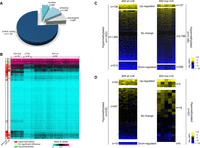
To more precisely define the CGI/promoter alterations in our glioma samples, we analyzed the DNA methylation profiles of 14,714 genes with a single CGI-rich promoter that could be assessed using the HM450K array. Most of these genes (76.0%) were protein-coding genes, and the others were antisense transcripts (10.7%), long intergenic noncoding RNAs (lincRNA; 6.5%), and pseudogenes (6.8%) (Fig. 1A). Because some CGI/promoters can control more than one gene, these 14,714 genes are associated with 11,795 CGI/promoters. About 90% of these CGI/promoters were unmethylated in nontumor control brain samples (mean β-value <0.2), and most of them remained unmethylated also in glioma samples. Among these GCI/promoters, 11.6% (n = 1369; associated with 1623 genes) were aberrantly hypermethylated in IDHwt samples, and 22.8% (n = 2692; 3198 genes) in IDHmut samples, contributing to their CIMP-positive status. Conversely, CGI/promoter hypomethylation, although more limited than hypermethylation, was more common in IDHwt (n = 198 CGI/promoters associated with 235 genes; 1.7%) than in IDHmut (n = 14 CGI/promoters associated with 22 genes; 0.12%) samples (Fig. 1B; Supplemental Fig. S1C,D).
Aberrant methylation at CGI/promoters is not the main contributor to transcriptional alteration in glioma. (A) Classification of the 14,714 genes analyzed in this study. (B) DNA methylation level (mean β-values) of the 11,795 CGI/promoters (rows) analyzed in IDHmut and IDHwt glioma and control (normal brain tissue) samples (columns). Left columns show their hypermethylation or hypomethylation status in IDHmut and IDHwt glioma samples compared with controls. (C,D) Differential expression status of genes associated with hypermethylated (C) or hypomethylated (D) CGI/promoters in IDHwt (left) and IDHmut (right) glioma samples compared with controls.
To evaluate the consequences of these DNA methylation changes, we analyzed eight IDHwt and five IDHmut glioma samples by RNA-seq. Most genes with aberrantly hyper- or hypomethylated CGI/promoters (79.9% in the IDHwt and 87.4% in the IDHmut group) showed no significant transcriptional change compared with control brain samples (|log2 Fc|>2; P < 0.05) (Fig. 1C,D). The number of genes with altered expression was similar between glioma subtypes despite their different DNA methylation profiles. Among the genes with aberrantly hyper- or hypomethylated CGI/promoters, 223 and 265 were down-regulated (82 in common), and 150 and 140 were up-regulated (44 in common) in IDHwt and IDHmut glioma samples, respectively, compared with controls (Fig. 1C,D). These findings indicate that aberrant CGI/promoter methylation minimally affects gene transcription in glioma. However, the deregulated genes included some putative tumor suppressors, such as RASL10A and HTATIP2 (Schmidt et al. 2012; Dong et al. 2015), and some putative oncogenes, such as HOXD9 and CXCL1, the overexpression of which was, counterintuitively, associated with methylation gain (Supplemental Table S2).
Gene transcription alterations are more widespread in IDHwt glioma samples
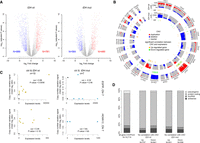
Detailed analysis of the transcriptional landscape of tumor samples showed that transcriptional alterations were more widespread in IDHwt than in IDHmut glioma samples (1670 and 1024 deregulated genes, respectively; FDR < 0.05; |log2 Fc|>2) (Fig. 2A), particularly for CGI/promoter-associated genes (Supplemental Fig. S2A).
Extent of transcriptional alterations in IDHwt and IDHmut glioma samples. (A) Volcano plot analysis of differential gene expression in IDHwt (left) or IDHmut (right) glioma samples. Blue and red dots represent genes that were significantly down- or up-regulated, respectively, compared with healthy controls (n = 14,714 genes analyzed). (B) Circular karyotype showing the duplication (red) and deletion (blue) frequencies at the 14,714 analyzed genes in IDHwt (outer circles) and IDHmut (inner circles) samples. Genes showing a significant correlation between CNV and expression are symbolized by an orange (up-regulated) or green (down-regulated) line. (C) Correlation analysis between CNV and expression levels for the EGFR and HOXA13 genes in IDHwt (yellow dots, left) and IDHmut (blue dots, right) glioma samples. Black dots indicate value in healthy controls. EGFR overexpression correlated with increased copy number in IDHwt glioma samples. (D) Classification of the genes with expression alterations that did not correlate with CNV.
Copy number variation (CNV) analyses in the same samples showed that, as previously reported, Chromosome 7 gain and Chromosome 10 loss characterized IDHwt samples (Louis et al. 2016), whereas the 1p/19q codeletion was mainly present in IDHmut samples (Fig 2B). By integrating these data with the gene expression profiles, we identified 92 genes in IDHwt and 37 genes in IDHmut samples, respectively, in which expression alteration correlated with CNV (P < 0.05) (Fig. 2B; Supplemental Fig. S2B,C; Supplemental Table S3). For instance, up-regulation of epidermal growth factor receptor (EGFR) and the histone methyltransferase EZH2 (both located on Chromosome 7) correlated with increased copy number in IDHwt samples. Conversely, overexpression of HOXA13, also located on Chromosome 7, did not correlate with CNV (Fig. 2C; Supplemental Fig. S2C).
Altogether, affected genes without CNV (mostly protein-coding genes) represented ∼11% of all CGI/promoter-associated genes in IDHwt samples (841 down-regulated and 737 up-regulated genes) and 6.7% in IDHmut samples (556 down-regulated and 431 up-regulated) (Fig. 2D; Supplemental Table S2).
Most transcriptional alterations are DNA methylation–independent
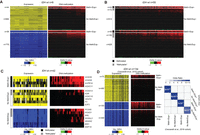
To understand the basis of such transcriptional alterations, we next focused our analyses on IDHwt glioma samples. We used paired RNA-seq and HM450K data from 8 IDHwt samples to concomitantly determine the DNA methylation and transcriptional changes in the 1578 affected genes. In these IDHwt samples, we could identify four main transcriptional defect types (Fig. 3A): gain or loss of gene expression associated with CGI/promoter hypermethylation (referred to as “Meth+/Exp+” and “Meth+/Exp−”defects, respectively), and gain or loss of gene expression without changes in CGI/promoter methylation status (i.e., the CGI/promoter remained unmethylated; referred to as “No Meth/Exp+” and “No Meth/Exp−” defects, respectively). More than 93% of aberrantly repressed genes did not display any significant DNA methylation alteration at their CGI/promoter (No Meth/Exp−). About 47% of the affected genes showed gain of expression that was associated with DNA hypermethylation at their CGI/promoter in 6% of them (Meth+/Exp+).
Four expression defect classes. (A) Integrative analysis of differential gene expression and methylation in eight IDHwt glioma samples identified four main defect classes: gain of expression with gain of methylation (Meth+/Exp+), gain of expression with CGI/promoter remaining unmethylated (No Meth/Exp+), loss of expression with gain of methylation (Meth+/Exp−), and loss of expression with the CGI/promoter remaining unmethylated (No Meth/Exp−). (B) Differential DNA methylation analysis in all IDHwt glioma samples (n = 55) versus controls (n = 8) (delta of the mean β-value). Glioma samples were grouped in the four classes of expression defects defined in A. The methylated and methylable status of genes is indicated in the left column. (C) Integrative analysis of differential expression and methylation at selected Meth+/Exp+ (upper), No Meth/Exp+ (middle), and No Meth/Exp− (lower) genes in 42 IDHwt glioma samples compared with controls (n = 8). (D) Integrative analysis of differential gene expression and methylation in an independent cohort of 135 IDHwt glioma samples (validation cohort) also identified the four main defect classes. Odds ratio and significance of the overlap (Fisher's exact test) between the data of the validation cohort and our cohort, for each defect category, are shown on the right panel.
To evaluate the robustness of this classification, we first analyzed the HM450K data of all 55 IDHwt glioma samples of our cohort and confirmed that DNA hypermethylation was associated with genes classified in the Meth+/Exp+ and Meth+/Exp− groups, and absence of methylation with genes classified in the No Meth/Exp+ and No Meth/Exp− groups. The only exception was a subset of “methylable” genes that gained methylation in some samples (Fig. 3B). Next, we concomitantly assessed, in 42 IDHwt glioma samples, the DNA methylation and expression by RT-qPCR of randomly selected genes from the Meth+/Exp+, No Meth/Exp+, and No Meth/Exp− groups. Genes in the Meth+/Exp+ group were methylated and ectopically expressed in all analyzed IDHwt glioma samples. The seven genes from the No Meth/Exp− group were aberrantly repressed in all analyzed samples, and their CGI/promoter mostly unmethylated (Fig. 3C). For instance, the candidate tumor suppressor gene BIN1 was unmethylated in all analyzed samples. PCSK6 and HOXD1 provided examples of “methylable” genes. They were methylated in a subset of samples, but their expression was repressed in all of them. The six genes from the No Meth/Exp+ group were all overexpressed; and most of them, including the tumor progression-associated VEGFA and E2F2 genes, tended to be unmethylated in all analyzed samples. This group also included some “methylable” genes (Fig. 3B), such as KDR (the tyrosine kinase receptor for VEGFA) that was overexpressed in all samples, and its CGI/promoter was methylated only in a subset of glioma. These “methylable” genes displayed an overall significant gain of DNA methylation in the 55 IDHwt samples. Therefore, we reclassified them from their initial No Meth/Exp+ and No Meth/Exp− groups (based on the analysis of eight IDHwt samples) into the Meth+/Exp+ and Meth+/Exp− groups, respectively, for the subsequent analyses.
To assess the reproducibility of these observations, we performed the same analyses in an independent cohort of 134 IDHwt samples described in Ceccarelli et al. (2016). In these samples, we confirmed the existence of the four main transcriptional defect groups, with proportions similar to those observed in our cohort. Overall, the number of affected genes was smaller in this validation cohort (970 vs. 1564), possibly because of the different RNA-seq strategies used. Indeed, Ceccarelli et al. (2016) used mainly nonstranded RNA-seq in which sense–antisense overlapping transcripts were discarded from the analyses, whereas we used a stranded RNA-seq approach that considered independently sense and antisense transcripts. Nonetheless, all four defect categories significantly overlapped in the two cohorts (minimal P-value <1 × 10−40; Fisher's exact test), demonstrating the robustness of our observations (Fig. 3D).
Finally, to further characterize these defects, we studied the methylation pattern of the 681 genes that were transcriptionally affected in both IDHwt and IDHmut glioma samples (377 down-regulated and 303 up-regulated in both groups, and one down-regulated in IDHwt and up-regulated in IDHmut). Most of these genes displayed the same methylation pattern in IDHwt and IDHmut samples. However, “methylable” genes and a subset of unmethylated genes in IDHwt gliomas (symbolized by a blue column in Supplemental Fig. S3) tended to be methylated in IDHmut samples, suggesting that different molecular pathways can lead to the same aberrant gene expression pattern.
Altogether, this integrative analysis identified four classes of transcriptional defects at CGI/promoter genes in IDHwt glioma: aberrant loss and gain of gene expression without DNA methylation defect (most genes), and gene expression defects associated with aberrant DNA methylation either in all samples or in some samples (i.e., at methylable genes). In summary, among the 1578 CGI/promoter-associated genes affected in IDHwt glioma samples, most belonged to the No Meth/Exp− group (n = 628), followed by the No Meth/Exp+ (n = 612), Meth+/Exp− (n = 208), and Meth+/Exp+ (n = 116) (Supplemental Fig. S4). The classification for each gene is provided in Supplemental Table S2.
The four defect classes include mainly genes with a chromatin bivalent signature in stem cells
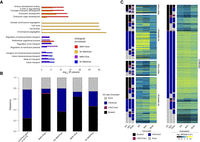
Gene Ontology analysis (Fig. 4A; Supplemental Fig. S5) showed that the Meth+/Exp− and No Meth/Exp− groups were enriched in genes involved in transmembrane ion transport, specifically in synapsis and neurons. Conversely, the No Meth/Exp+ group was enriched in genes implicated in general processes, such as cell cycle, cell division, and chromosome segregation. Finally, the Meth+/Exp+ group was highly enriched in genes encoding homeodomain proteins, including the HOX family, and implicated in embryonic development.
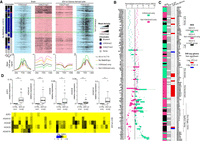
Genes with bivalent chromatin signature in ES cells are more prone to be deregulated in IDHwt glioma. (A) Gene Ontology terms (biological processes) enriched in genes from the four defect categories. For each category, the four highest terms are shown. (B) Distribution of genes of each defect category according to their chromatin signature in human ES cells: (none) gray; (bivalent) black; (H3K4me3-only) blue; (H3K27me3-only) purple. As reference, the distribution of the 14,714 genes analyzed in this study according to their chromatin signatures in human ES cells is shown in the left panel. (C) Expression level and chromatin signatures of genes of the four defect categories in human ES cells, neural progenitor cells (NPCs), and healthy brain. For comparison, the same analysis is provided on the right panel for genes without expression defect (unaffected) in IDHwt glioma samples.
Studies on the molecular bases of CGI hypermethylation have shown that genes with a bivalent chromatin signature in stem cells are more likely to gain aberrant methylation in cancer cells (Ohm et al. 2007; Deneberg et al. 2011; Court and Arnaud 2017). To evaluate whether such an instructive program could apply to some or all genes of the four defect categories, we evaluated the GCI/promoter chromatin signature of genes in these four categories in human embryonic stem (ES) cells, in NPCs, and in brain samples. In agreement with previous findings, most genes in the Meth+ groups showed bivalent signatures in ES cells and NPC. This was true regardless of their aberrant expression pattern (i.e., both Meth+/Exp− and Meth+/Exp+). Similarly (but more unexpectedly), ∼36% and 48% of genes in the No Meth/Exp+ and No Meth/Exp− groups, respectively, showed a bivalent chromatin signature in ES cells (compared with 24% of all studied genes) (Fig. 4B; Supplemental Fig. S5).
In agreement with the resolution of the bivalent signature during development/cell differentiation, the chromatin signature tended to change toward an exclusive H3K4me3 signature, but also to a “none” signature (i.e., depleted for both H3K4me3 and H3K27me3) in brain samples (Fig. 4C). In comparison, most of the transcriptionally unaffected genes maintained their H3K4me3-only signature from ES cell to brain samples. Accordingly, the transcriptionally affected genes displayed a dynamic expression pattern from ES cells to NPC and brain. This was true also for the subset of genes in the No Meth/Exp+ and No Meth/Exp− groups that maintained an exclusive H3K4me3 signature in ES cells, NPC, and brain, but showed loss and gain of expression in brain, respectively (Fig. 4C).
Altogether, these findings suggest that genes with a bivalent chromatin signature in ES cells and/or with a dynamic expression pattern during neural differentiation are more prone to be deregulated in IDHwt glioma.
CGI/promoter hypermethylation is associated with gain of expression
Because bisulfite treatment cannot distinguish between methylation and 5-hydroxymethylation (5-hmC), we evaluated whether the DNA methylation level detected with the HM450K array at genes classified in the Meth+/Exp+ group could be explained by 5hmC deposition. By using publicly available 5hmC data for IDHwt tumors (Johnson et al. 2016), we found that CGI/promoter regions of Meth+/Exp+ genes were devoid of 5hmC (median level = 1.2%), indicating that the signal detected with the HM450K array was attributable to DNA methylation (Supplemental Fig. S6A). Note that a similar observation is made at the three other defect groups (Supplemental Fig. S7).
To determine how DNA hypermethylation and expression gain could coexist in this group of genes, we next assessed their CGI/promoter chromatin signature using publicly available ChIP-seq data for IDHwt glioma cells. Compared with healthy brain (control), H3K27me3 level was strongly decreased, whereas H3K4me3 level was increased in ∼65% of these genes and totally depleted in the others (Fig. 5A).
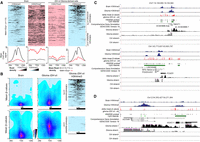
Expression from genes with methylated CGI/promoter. (A) Data mining–derived ChIP-seq read density data for H3K27me3 (pink) and H3K4me3 (blue) in “Meth+/Exp+” genes in a ±2 kb window centered on their TSS, in healthy brain (left) and IDHwt-derived cell lines (right). The mean ChIP-seq signal values are shown on the lower panels for “Meth+/Exp+” genes (red line) and for the 14,714 analyzed genes (black line) that were used as normalized reference. (B) Heatmap showing CpG sites density and their mean methylation level in a ±2 kb window centered on the TSS of “Meth+/Exp+” genes and enriched (upper) or depleted (lower) for H3K4me3 in IDHwt glioma samples compared with healthy controls. The ChIP-seq read density obtained in IDHwt-derived cell lines is shown on the right panels. (C) Genome Browser view at the TWIST1 and FOXD3 loci to show H3K4me3 enrichment, differential DNA methylation, and the oriented RNA-seq signal. These two loci are representative of genes that initiate from an H3K4me3-marked TSS embedded in a methylated CGI/promoter in IDHwt samples. (D) HOXC11 is representative of genes in which expression initiates from an alternative TSS in IDHwt glioma samples.
Analysis of the HM450K data on the localization of hypermethylated sites relative to the transcription start site (TSS) showed that at H3K4me3-enriched genes, the gain of DNA methylation occurred at the border of the CGI/promoter, while the TSS area remained unmethylated (Fig. 5B). Analysis of individual loci in glioma samples using stranded RNA-seq data suggested that transcription initiated from H3K4me3-marked TSS embedded in methylated CGIs (Fig. 5C; Supplemental Fig. S6B). This was observed in several genes that promote gliomagenesis, including TWIST1 (Mikheev et al. 2018), CTHRC1 (Liu et al. 2017a), and FOXD3-AS1 (Chen et al. 2016). The H3K4me3 and DNA methylation signals were mutually exclusive (Fig. 5C; Supplemental Fig. S6B), in agreement with their documented antagonism (Weber et al. 2007).
In H3K4me3-depleted genes, DNA methylation was spread along the entire CGI/promoter, including the TSS (Fig. 5B), suggesting that transcription from these genes could arise from an alternative TSS. Analysis of RNA-seq data supported this hypothesis because genes, such as HOXC11 and NR2F2, showed transcription signal from H3K4me3-enriched regions located away from the documented TSS (Fig 5D; Supplemental Fig. S6C). However, for few genes, such as HEYL and C15orf48 (Supplemental Fig. S6D), transcription apparently initiated from a methylated CGI through an unknown mechanism.
Altogether, these approaches support the hypothesis that in Meth+/Exp+ genes, transcription could be allowed by the absence of DNA methylation at the TSS or the use of alternative TSS.
E2F- and HOX-target genes are frequently overexpressed in glioma
The publicly available ChIP-seq data indicated that genes in the No Meth/Exp+ group were enriched in H3K4me3 and depleted in H3K27me3 in IDHwt glioma cells compared with healthy brain (Fig. 6A). We observed this H3K4me3 gain also in the subset of genes that were constitutively marked by the H3K4me3-only signature in ES cells, NPC, and brain (Fig. 6A, green rectangle). To understand the basis of their overexpression in glioma, we determined whether specific motifs were enriched at their CGI/promoters. We found that overall, No Meth/Exp+ genes were putative targets of transcription factors associated with cell cycle pathways, including the Krüppel-like factors (KLF)/specificity protein (SP) and E2F families (Fig. 6B,C). Moreover, most of the H3K4me3-only genes showed specific motif enrichment for the E26 transformation-specific (ETS) and nuclear transcription factor Y (NFY) families. The other No Meth/Exp+ genes were putative targets of homeodomain transcription factors, including HOX proteins (Fig. 6B).
Transcription factor binding motifs in the promoters of genes overexpressed in glioma samples. (A) Data mining–derived ChIP-seq read density data for H3K27me3 (pink) and H3K4me3 (blue) at “No Meth/Exp+” genes in a ±2 kb window centered on their TSS in healthy brain (left) and IDHwt-derived cell lines (right). The mean ChIP-seq signal values are shown on the lower panel for all “No Meth/Exp+” genes (orange line) and for those that are (dotted green line) or not (dotted pink line) marked by H3K4me3-only in ES cells, NPC, and brain. The black line, used as normalized reference, shows the value for all analyzed genes. (B) Transcription factor motif enrichment in the CGI/promoter of “No Meth/Exp+” genes, calculated using i-cis Target and represented as a normalized enrichment score (NES). Enrichment is shown for genes that are (green squares) or are not (pink squares) marked by H3K4me3-only in ES cells, NPC, and brain. When a transcription factor possesses several binding motifs, data are presented as a box plot. (C) Expression status, assessed by RNA-seq, of the transcription factors identified in B. The middle column shows their expression status in healthy control (n = 5) (white, not expressed; gray, expressed: fpkm > 1) and the right column their expression in IDHwt glioma samples (n = 8). The left column shows the motif enrichment in all “No Meth/Exp+” genes (black), and those marked (green) and not marked (pink) by H3K4me3-only in ES cells, NPC, and brain. (D) Expression versus controls of selected overexpressed transcription factor identified in C assessed by RT-qPCR in 42 IDHwt glioma samples. Details for each sample are provided in the lower panel (P-value by Mann–Whitney U test).
Most of these transcription factors were expressed in healthy controls and remained expressed in glioma samples (Fig. 6C). However, a subset was specifically overexpressed in IDHwt glioma samples, including HOX genes, E2F2, E2F7, ETS1, ETV1, and ETV4 (Fig. 6C, RNA-seq data). We confirmed these findings by RT-qPCR analysis of selected genes in 42 IDHwt samples (Fig. 6D). Genes encoding aberrantly overexpressed transcription factors mostly belonged to the Meth+/Exp+ (e.g., HOXA2 and HOXD8) and No Meth/Exp+ (e.g., E2F2) groups (Supplemental Table S2). This observation suggests that the initial overexpression of a few key transcription factors could lead to overexpression of most genes belonging to the No Meth/Exp+ group.
The repressive signature of silenced genes is related to their transcriptional status in healthy brain
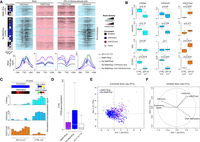
To understand how Meth+/Exp− and No Meth/Exp− genes were transcriptionally repressed, we compared their chromatin signature in healthy brain and in glioma cells using publicly available ChIP-seq data. This highlighted a marked H3K27me3 enrichment in both groups in glioma samples compared with controls (Fig. 7A). The only exception was the subset of No Meth/Exp− genes that displayed the constitutive H3K4me3-only signature in ES cells, NPC, and brain (Fig. 7A, light blue rectangle) and maintained this signature in glioma cells. H3K27me3 enrichment in glioma cells led to a bivalent chromatin signature in the large subset of genes that were also marked by H3K4me3 (Fig. 7A). At CGIs/promoters of Meth+/Exp− genes, H3K4me3 tended to be reduced and H3K27me3 was enriched (Fig. 7A), suggesting that unlike normal cells (Brinkman et al. 2012), both H3K27me3 and DNA methylation can coexist at CGI/promoters in glioma cells.
Gene repression is associated with H3K27me3 gain. (A) Data mining–derived ChIP-seq read density data for H3K27me3 (pink) and H3K4me3 (blue) at “Meth+/Exp−” and “No Meth/Exp−” genes in a ±2 kb window centered on their TSS, in healthy brain (left) and IDHwt-derived cell lines (right). The mean ChIP-seq signal values are shown in the lower panels for “Meth+/Exp−” (purple line) and “No Meth/Exp−” (blue line) genes. Genes in the “No Meth/Exp−” group were further subdivided in genes marked (dotted light blue line) and not marked (dotted dark blue line) by H3K4me3-only in ES cells, NPC, and brain. The black line used as normalized reference shows the value for all analyzed genes. (B) ChIP analysis of H3K9ac, H3K4me3, and H3K27me3 at selected genes in IDHwt (n = 7) and control (n = 5) samples. The precipitation level was normalized to that obtained at the TBP promoter (for H3K4me3 and H3K9ac) and at the SP6 promoter (for H3K27me3; P-values calculated with the Mann–Whitney U test). (C) Detail for each sample of the ChIP analysis at the PCSK6 locus. Heatmaps of the expression and methylation values are in the upper panel. (D) Expression level of “Meth+/Exp−” (purple column) and “No Meth/Exp−” (blue column) genes and of all analyzed genes (white column) in healthy controls. (E,F) Principal component analysis. (E) Two-dimensional scatter plot of the values of each “Meth+/Exp−” (purple dots) and “No Meth/Exp−” gene (blue dots) along the first (Dim 1) and second (Dim 2) principal component. For each class defect, the centroids are shown by colored squares. (F) H3K4me3 and expression levels in healthy brain are the variables that most contributed to and were significantly correlated with the first principal component.
ChIP analysis of selected genes from the No Meth/Exp− group (PCSK6, MAL, SH3GL3, and NKAIN2) confirmed the marked H3K27me3 gain associated with reduced or unchanged H3K4me3 and H3K9ac levels, according to the studied locus, in the seven glioma samples tested (Fig. 7B,C; Supplemental Fig. S8A). Also, bisulfite analysis of the H3K27me3-immunopreciptated fraction confirmed that both DNA methylation and H3K27me3 coexisted at CGI/promoters in glioma samples, as exemplified by the methylable PCSK6 gene (Supplemental Fig. S8B).
Several studies have highlighted that the propensity of genes to be hypermethylated in cancer cells is related to their transcriptional status in the normal tissue (Gal-Yam et al. 2008; Sproul et al. 2011, 2012; Hinoue et al. 2012; Court and Arnaud 2017). Accordingly, we observed that the transcriptional status in brain discriminated between Meth+/Exp− and No Meth/Exp− genes. Specifically, DNA methylation–independent silencing (No Meth/Exp−) mainly affected genes that were highly expressed in normal brain. Conversely, poorly expressed genes tended to be hypermethylated (Fig. 7D). To more systematically assess the bases of the differences between these groups, we performed principal component analysis (PCA) for all genes in the Meth+/Exp− and No Meth/Exp− groups. The first principal component accounted for ∼44.5% of the variance and allowed separating the two groups (centroid values along the axis: +0.984 and −0.326, respectively) (Fig. 7E). The observation that the first principal component was significantly correlated with the permissive H3K4me3 mark (R = 0.79; P < 3.4 × 10−126) and the expression level (R = 0.71; P < 1 × 10−129) in healthy brain (Fig. 7F) supported the hypothesis that the expression status in healthy cells contributes to the choice of silencing pathway used in cancer cells. Additional analyses using normalized RNA-seq data for 21 different tissues showed that genes from the No Meth/Exp− group were strongly expressed specifically in adult brain tissues (Supplemental Fig. S8C,D).
Thus, besides DNA methylation, H3K27me3 enrichment and chromatin bivalency emerge as major causes of aberrant gene repression in aggressive glioma cells. The choice between these alternative silencing pathways is related to the expression level of the affected genes in healthy tissues/cells.
The four classes of expression defects are observed also in IDHmut glioma samples
We next extended our analyses to IDHmut glioma. Integrative analysis of differential gene expression and methylation identified the four main defect classes also in these samples (Supplemental Fig. S9A–C; Supplemental Table S2). DNA methylation–independent defects were the most frequent, although less prominent than in IDHwt samples. Specifically, the 556 repressed genes were equally distributed between the No Meth/Exp− (n = 294) and the Meth+/Exp− (n = 262) groups (Supplemental Fig. S4).
These observations were validated in an independent cohort of 415 IDHmut samples derived from Ceccarelli et al. (2016), with a highly significant overlap in the composition of each defect group between cohorts (minimal P-value <1 × 10−40; Fisher's exact test) (Supplemental Fig. S9D). We also observed that, as in IDHwt samples, genes with a bivalent chromatin signature in ES cells and/or with a dynamic expression pattern during neural differentiation were more prone to be deregulated in IDHmut glioma (Supplemental Fig. S10). Then, to understand the molecular bases of these alterations, and in the absence of publicly available ChIP-seq data on IDHmut samples, we performed ChIP analyses at selected genes from the Meth+/Exp− group (PCSK6 and MAL) and No Meth/Exp− group (SH3GL3 and PCDH10) in five IDHmut samples. We found that H3K27me3 enrichment was associated with reduced or unchanged H3K4me3 and H3K9ac in function of the studied locus (Supplemental Fig. S11A). This confirms that, like in IDHwt samples, gain of H3K27me3 and chromatin bivalency are, besides DNA methylation, the main hallmarks of repressed genes in IDHmut glioma cells. Finally, by considering the expression level in healthy brain and performing principal component analysis, we observed that the expression status in healthy cells contributed to the choice of silencing pathway used in IDHmut cells, with genes repressed independently of DNA methylation (No Meth/Exp− group) being mainly genes that were highly expressed in brain (Supplemental Fig. S11B–D).
Altogether, these approaches show that the main observations made in IDHwt glioma also apply to IDHmut glioma.
Discussion
Here, we used glioma, one the most widespread brain tumor types, as a model to evaluate the relative contribution of DNA methylation–dependent and –independent mechanisms to transcriptional alteration at CGI/promoter-associated genes in cancer cells. Our study showed that H3K27me3 level changes are the predominant molecular defect at both aberrantly repressed and expressed genes. Moreover, our findings support that H3K27me3 dynamics deregulation, particularly when present in a bivalent context, is the main cause of transcriptional alteration in glioma cells.
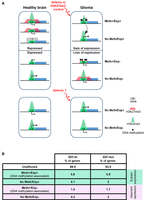
Some studies have described H3K27me3-based transcriptional repression in cancer cells (Gal-Yam et al. 2008; Kondo et al. 2008; Dudziec et al. 2012; Statham et al. 2012; Hahn et al. 2014). In colorectal tumors, ectopic gene expression has been associated with aberrant loss of H3K27me3 from CGI/promoters with bivalent chromatin signature (Hahn et al. 2014). Moreover, gene expression changes in primary human clear cell renal cell carcinomas can be attributed to chromatin accessibility alterations, independently of DNA methylation (Becket et al. 2016). Our study further extends these observations and provides, for the first time, a comprehensive description of these alterations in glioma samples and their relative contribution to transcriptional alteration in such tumors. Specifically, our integrative analyses identified and quantified four main types of transcriptional defects in glioma (Fig. 8) that recapitulate the DNA methylation- and H3K27me3-based molecular signatures at aberrantly repressed and expressed CGI/promoter-associated genes. We detected these defects in IDHwt and also in IDHmut glioma samples (Fig. 8B), indicating that they occur regardless of the G-CIMP status and tumor aggressiveness. Additional studies are required to determine whether the relative distribution of these defects can discriminate different IDHmut subpopulations (e.g., classified according to the 1p/19q codeletion status) and can be associated with specific clinical features. Moreover, these observations prompt to investigate whether they might apply to cancer cells in general.
Working model. (A) In glioma, alterations in the control of the H3K27me3 signature could be one of the main contributors to the four types of transcriptional defects observed at CGI/promoter-controlled genes (upper). In this model, genome-wide hypomethylation induces H3K27me3 redistribution that could lead to ectopic expression of genes that are normally repressed by polycomb proteins, including some genes encoding transcription factors. These overexpressed transcription factors could then promote the aberrant expression of their target genes (dotted arrow). Similarly, alterations in the interplay between the polycomb complex and the transcriptional machinery could affect H3K27me3 fate during ES and/or neural stem cell differentiation. Specifically, this alteration could lead to the aberrant maintenance of bivalency and silencing at a subset of genes that are normally specifically expressed in brain. At genes that are normally poorly expressed in healthy brain, this process is associated with gain of DNA methylation in glioma. Beside defects in the H3K27me3 signature, we also identified a subset of genes that are apparently constitutively associated with H3K4me3-only, regardless of their expression status in brain and glioma (lower). The mechanisms underlying their transcriptional deregulation remain to be determined. (B) Percentage of unaffected and affected CGI/promoter-controlled genes for each of the four described defects in IDHwt and IDHmut glioma samples from our cohort.
Our study revisited the relationship between aberrant transcription and DNA methylation in cancer cells. First, we confirmed that gene expression deregulation is very rarely associated with CGI/promoter DNA hypomethylation in glioma samples, in agreement with the general unmethylated status of CGI/promoters in healthy cells. Unexpectedly, we found that DNA hypermethylation is not the main cause of transcriptional repression at CGI/promoter genes, and that it can be associated also with gain of expression. Indeed, ∼16% of ectopically expressed genes were associated with a hypermethylated CGI/promoter in IDHwt glioma samples. Specifically, in many genes, ectopic expression was associated with CGI/promoters that gained methylation at their borders, whereas the H3K4me3-marked TSS was methylation-free. At other genes, extensive methylation of their main CGI/promoter was associated with the use of an alternative promoter. It is not known whether there is a causal link between these events. These two distinct signatures have also been described in prostate cancer cell lines (Bert et al. 2013), suggesting that an association between CGI/promoter DNA hypermethylation and gain of gene expression is common in cancer.
Besides the concomitant gain of expression and DNA methylation, this group of genes was characterized also by a reduction of H3K27me3 level in glioma cells (compared with controls), suggesting that the interplay between these repressive marks is a driving force in their transcriptional deregulation. In the mouse, widespread DNA methylation depletion triggers redistribution of H3K27me3 (Brinkman et al. 2012; Reddington et al. 2013) that in turn leads to a loss of H3K27me3 and ectopic expression at a subset of polycomb target genes, including Hox clusters (Reddington et al. 2013). Reciprocally, analyses in mouse ES cells showed that the area close to CGI/promoters of polycomb target genes is protected from aberrant DNA methylation gain by polycomb proteins (Li et al. 2018). Therefore, the Meth+/Exp+ defect might affect a subset of polycomb target genes that are particularly sensitive to the H3K27me3 redistribution induced by the genome-wide hypomethylation of cancer cells. Noteworthy, in the aggressive IDHwt glioma, this group is enriched in homeodomain genes, and more specifically in HOX genes. Deregulation of HOX genes contributes to the tumorigenic potential of glioblastoma stem cells by activating a network of downstream genes (Gallo et al. 2013). Accordingly, we observed that many ectopically expressed genes from the No Meth/Exp+ group are putative HOX transcription factor targets. Altogether, our observations support a domino effect model to account for the gain of expression of CGI/promoter genes in aggressive glioma. In this model, genome-wide hypomethylation leads to ectopic expression (and methylation gain) of Meth+/Exp+ genes (especially HOX genes) that then promote gain of expression of target genes from the No Meth/Exp+ group (Fig. 8A). Additional studies are required to test this model and specifically whether No Meth/Exp+ genes are bona fide HOX transcription factor targets.
Another key finding of our study is that a bivalent chromatin signature in stem cells may not only predispose genes to hypermethylation, as widely documented (Ohm et al. 2007; Deneberg et al. 2011; Court and Arnaud 2017), but more globally to transcriptional alteration in cancer cells. We observed that genes with CGI/promoters marked by a bivalent chromatin signature in ES cells and NPC were more prone to be deregulated in glioma samples, regardless of the transcriptional defect. This was particularly true for genes with DNA methylation-associated defects, irrespective of their association with gain or loss of expression (Meth+/Exp+ and Meth+/Exp−), and to a lesser extent, for genes with DNA methylation–independent defects (No Meth/Exp+ and No Meth/Exp−). This observation suggests that defects in the control of the bivalent chromatin signature, and more specifically of H3K27me3 dynamics, upon differentiation, are one of the main causes of transcriptional deregulation of CGI/genes in cancer cells. Accordingly, we found that aberrant gene repression in glioma samples affected mainly genes with brain-specific expression, and thus more sensitive to bivalency alterations upon neural stem cell differentiation. Besides the functional aberrations of H3K27me3 and H3K4me3 writers and erasers documented in many tumors (Suvà et al. 2013; Dawson 2017), these control defects could also result from transcriptional changes in key tissue-specific transcription factors or cofactors. Studies in mouse ES cells and tissues showed that their transcriptionally inactive status is sufficient to promote the recruitment of the polycomb responsive complex PRC2 and H3K27me3 deposition at CGI/promoters (Mendenhall et al. 2010; Klose et al. 2013; Riising et al. 2014; Jadhav et al. 2016; Maupetit-Méhouas et al. 2016). This suggests that the fate of bivalent chromatin domains upon development/differentiation relies on the interplay between PRC2 and the availability of the ad hoc transcriptional machinery. Our observation that the gene transcription strength in healthy brain influences the choice of silencing mechanism at repressed genes in glioma precisely argues for an alteration in this interplay. Specifically, we propose that following the alteration of a subset of brain-specific cofactors, the resulting weakened transcriptional machinery cannot efficiently counteract PRC2 recruitment upon differentiation, leading to aberrant maintenance of chromatin bivalency and to silencing of a subset of genes that are normally specifically expressed in brain. Moreover, gain of function of factors that promote, directly or indirectly, the recruitment of PRC2 at CGI could facilitate this process. This includes for instance the histone demethylase KDM2B (Farcas et al. 2012; Wu et al. 2013; Blackledge et al. 2014) that is critical in various cancers, including glioma (Staberg et al. 2018; Yan et al. 2018). At normally poorly expressed genes, these events associated with the initial weak level of H3K4me3, a mark that prevents recruitment of DNA methyltransferases, would facilitate the subsequent gain of DNA methylation (Fig. 8A).
In addition to genes in which their chromatin signature was altered in glioma, we also identified a subset of genes with an apparent constitutive H3K4me3-only signature in healthy (ES cells to brain) and glioma samples, and that showed either gain of expression (No Meth/Exp+) or aberrant repression (No Meth/Exp−) in glioma samples (Fig. 8A). Additional studies are required to establish the molecular bases of these observations. Specifically, it would be interesting to determine whether the regulation of these genes in normal and pathological contexts relies exclusively on the availability of ad hoc transcription factor(s), or whether it is also associated with not yet identified repressive histone marks. Moreover, No Meth genes that are ectopically expressed in glioma samples were also expressed in ES cells and NPC (Fig. 4). Similarly to genes with bivalent chromatin signature in ES cells whose aberrant repression in glioma recapitulated their repression in stem cells (Fig. 4), this group of genes could contribute to glioma aggressiveness by maintaining tumor cells in a stem-cell-like state.
Our study also provides a framework to explain the counterintuitive observation that patients with CIMP-positive IDHmut have a better clinical outcome than patients with CIMP-negative IDHwt. Our data indicate that CIMP is observed mainly at genes that are already repressed in healthy brain. Consequently, the number of deregulated genes with CGI/promoters associated with DNA methylation defects is similar between glioma subtypes. Conversely, the higher frequency (about two times) of DNA methylation–independent transcriptional alterations in IDHwt than in IDHmut samples could contribute to the prognosis difference between glioma subtypes. The CIMP-positive status, by promoting stable gene repression, could also act as a protective mechanism against cancer progression, by limiting the tumor epigenetic plasticity and its ability to adapt to environmental changes, such as metastasis formation or treatment (Sproul and Meehan 2013). Specifically, among the many genes that gain expression in IDHwt samples and that are maintained repressed through gain of methylation in IDHmut samples (Supplemental Table S4), a dozen are oncogenes with some documented for their role in glioma biology, such as spalt like transcription factor 4 (SALL4) (Liu et al. 2017b) and the long noncoding RNA MIR155 host gene (MIR155HG) (Wu et al. 2017).
In conclusion, our study on the extent and consequences of epigenetic alterations in glioma indicates that transcriptional deregulations rely mainly on chromatin-based DNA methylation–independent mechanisms. It also shows that the gene expression level in healthy tissue influences the type of silencing pathway used for repression in cancer cells, whereby highly expressed genes are more likely to be repressed by H3K27me3 rather than DNA methylation. Besides providing an original framework to understand the epigenetic basis of carcinogenesis, these observations are also important for the design of drugs to target epigenetic defects in cancer.
Methods
Tumor and control samples
Glioma samples (n = 70) resected between 2007 and 2014 were obtained from Clermont-Ferrand University Hospital Center, France (anonymized samples from the “Tumorotheque Auvergne Gliomes,” ethical approval DC-2012-1584). The ethics committees and the respective competent authorities approved this study. The study protocols conform to the World Medical Association Declaration of Helsinki.
Immediately after surgery, samples were snap frozen and stored in liquid nitrogen. Random sections of each tumor were analyzed under a light microscope after hematoxylin–eosin staining to determine the extent of necrosis and the percentage of tumor cells. All glioma samples included >50% of tumor cells. Gliomas were classified according their IDH1 mutation status: IDHwt (n = 55) and IDHmut (n = 15). IDH1 genotyping was performed by EpigenDx (pyrosequencing assays ADS1703 and ADS1704) (Fogli et al. 2016).
Eight control brain samples (healthy controls, mean age of 27.3 yr, standard deviation ± 2 yr) were removed by autopsy 4–16 h after accidental death (Brain and Tissue Bank of Maryland). These samples, identified by the Brain and Tissue Bank of Maryland as corpus callosum (n = 5) and frontal cortex (n = 3), corresponded to white matter enriched in astrocytes and oligodendrocytes from which gliomas originate.
Tumor and control samples were homogenized into powder by cryogenic grinding, equally distributed in at least three vials before use for matched genomic DNA, RNA, and chromatin extraction. All samples were stored at −80°C until use. Overall survival (OS) was calculated as the number of days between the surgery date and the patient's death. Tumor resection was classified as gross total resection, when no enhanced contrast was detected 48 h postsurgery, or as partial resection, when enhanced contrast was still detected 48 h postsurgery.
The demographic and clinical features are presented in Supplemental Table S1. The related statistical analyses were performed using the Stata software, version 13 (StataCorp) and the R software, version 3.3.1 (R Core Team 2016). To test the prognostic value of the patients’ characteristics in univariate analyses, OS curves were compared between groups using the log-rank test. Results are expressed as hazard ratios (HRs) and 95% confidence intervals (CIs).
Validation cohorts, obtained from The Cancer Genome Atlas (TCGA) research network, were described in Ceccarelli et al. (2016). We retained IDHmut (n = 415) and IDHwt (n = 134) samples for which matched DNA methylation (HM450K array) and RNA expression (RNA-seq) data were available. Clinical and molecular data on these patients were retrieved from the cBioPortal for Cancer Genomics (https://www.cbioportal.org/) (Cerami et al. 2012; Gao et al. 2013); processed RNA-seq and methylation data were obtained from the TCGA web site (https://portal.gdc.cancer.gov/) and analyzed as described below. The TCGA ID for each sample is provided in the Supplemental Table S5.
Genome reference
Arrays (HM450K and Cytoscan HD), as most of the databases used in this study, are based on the hg19 reference. Moreover, compared with GRCh37/hg19, a higher fraction of HM450K probes displays low mapping quality with the GRCh38 assembly (Zhou et al. 2017). Therefore, we used the hg19 as the genome reference in this study. Realigning to GRCh38 assembly should not significantly affect the conclusions of this work as we determined that among the 11,795 CGI/promoters, all being single CGI/promoters within the associated gene(s), studied here, >93% remained single CGI/promoters (i.e., overlap ±1 kb with the TSS area) within the associated gene(s) in GRCh38. This proportion is maintained when considering only affected CGI/promoters.
Selection of the genes to be analyzed
The positions of genes and CpG islands (CGI: defined using standard criteria: GC content ≥50%; length >200 bp; ratio Obs/Exp CpG >0.6) were downloaded from GENCODE annotation release V19, and CpG island tracks of UCSC hg19 assembly, respectively. For each gene, the promoter region was defined as TSS ±1 kb. To assess the relationship between CGI/promoter methylation status and gene expression, we first identified the genes associated with only one promoter with CGI features (n = 15,350). Because our cohort included both men and women, we then excluded genes located on the X or Y Chromosomes. We finally retained 14,714 genes in which the CGI/promoter is covered by at least two probes in the HM450K Illumina array.
DNA methylation analyses
DNA extraction
DNA was isolated from frozen tissue samples using the QIAamp DNA Mini Kit (Qiagen) according to the manufacturer's recommendations.
Gene-specific bisulfite sequencing
Bisulfite conversion, PCR amplification, cloning, and sequencing were performed as previously described (Arnaud et al. 2006). Details on the primers used are in Supplemental Table S6.
HM450K analysis
The HM450K array data for controls and gliomas sample (eight normal brain, 55 IDHwt, and 15 IDHmut gliomas) were analyzed as previously described (Maupetit-Mehouas et al. 2018). Specifically, DNA bisulfite conversion and array hybridization were performed by Integragen SA, using the Illumina Infinium HD methylation protocol. β-Values were computed using the GenomeStudio control interplate normalization and background subtraction (version 2011.1, manifest files: HumanMethylation450_1 5017482_v.1.2.bpm). For each sample, β-values with a detection P-value >0.01 were excluded. All probes with a detection P-value >0.01 or lacking signal in >5% of our samples were excluded. Finally, 26,507 probes containing common SNPs (dbSNP 147) in their last 5 bp or in the CpG sites were discarded. Because our patient cohort included both men and women, probes on the X and Y Chromosomes were also excluded from the analysis. After the implementation of these quality filters, a total of 443,691 CpG methylation values were considered suitable for the downstream analysis. Methylation level at the 14,714 genes (11,795 CGIs) was given by the mean β-value of all probes located in their CGI. Differential methylation analyses were performed using the limma R package (Ritchie et al. 2015), as previously described (Court et al. 2014). These analyses concerned the entire groups (eight controls vs. 55 IDHwt, and eight controls vs. 15 IDHmut) or only the samples with matched RNA-seq data (three controls vs. eight IDHwt and three controls vs. five IDHmut). The HM450K probes were considered differentially methylated when the FDR was <0.05 and when the β-value difference between groups was >0.1. To consider only robust methylation variations, CGI/promoters were classified as hypermethylated or hypomethylated only if they included at least two probes differentially methylated (gain or a loss of methylation) in their CGI.
To test the robustness of this strategy we performed again the analyses by using Wilcoxon test, instead of limma (Ritchie et al. 2015), and by including the filters described in Zhou et al. (2017). For both IDHwt and IDHmut samples, we identified the same four groups of defects. At the statistical level, the affected probes displayed >91% of homology with both tests. In addition, the percentage of affected genes for each defect only marginally changed between our initial conditions and these conditions (Supplemental Fig. S12).
5hmC analysis by data mining
5hmC data from 30 IDHwt glioma samples were retrieved from Johnson et al. (2016) (GSE73895). β-values, derived from HM450K arrays hybridized with DNA after conversion with oxidative bisulfite (oxBS) and bisulfite only (BS), were computed as described above. The level of 5hmC for each probe and sample was obtained by subtracting the β-values of oxBS samples from their corresponding BS pair.
Expression analysis
RNA extraction
RNA was isolated from frozen tissue samples using the RNeasy Mini Kit (Qiagen), according to the manufacturer's recommendations.
RT-qPCR expression analyses
RT-qPCR assays were performed using a microfluidic-based approach as previously described (Maupetit-Méhouas et al. 2016). First-strand cDNA was preamplified for 14 cycles with the pool of primers used for RT-qPCR and the TaqMan PreAmplification Master Mix (Life Technologies 4488593). Primer sequences are in Supplemental Table S6. RT-qPCR assays were then performed and validated using Fluidigm 96.96 Dynamic Arrays and the Biomark HD system (Fluidigm) according to the manufacturer's instructions. The relative expression level was quantified with the 2-dCt. The housekeeping genes PPIA, TBP, and RPL13A were used to normalize transcript expression. Each analysis was performed in duplicate.
RNA-sequencing
RNA-seq was performed using total RNA after ribosomal RNA depletion (Ribo-Zero rRNA Removal Kit, Illumina) from three brain, eight IDHwt, and five IDHmut glioma samples. RNA processing, rRNA depletion, library preparation, and sequencing on an Illumina HiSeq 2500 apparatus were performed by Integragen SA according to the manufacturer's protocol (mean of 90 million of paired reads per sample). Stranded RNA-seq reads were mapped to the human genome (hg19) using TopHat2 (version 2.1.0) and a transcript annotation file from GENCODE (Release 19) (Kim et al. 2013). The average alignment rate was ∼94.5% with a concordant pair alignment rate of 92%. Only properly paired reads were considered for downstream analysis. The read count per gene was obtained with the HTseq-count script (option: -m intersection-nonempty -s reverse), and the FPKM gene expression level was determined with Cuffquant and Cuffnorm from the Cufflinks suite (version 2.2.1) based on GENCODE V19 transcript annotation (Trapnell et al. 2010; Anders et al. 2015). Strand-specific RNA-seq signals were derived from the RNA-seq alignments using SAMtools, genomeCoverageBed, and bedGraphToBigWig tools, and visualized using the UCSC Genome Browser (Li et al. 2009; Kent et al. 2010; Quinlan and Hall 2010). Differential expression analyses between controls and glioma samples were based on read counts using the DESeq2 and edgeR R packages (Robinson et al. 2010; Love et al. 2014). Genes were considered as differentially expressed between groups when |log2-fold change| >2 with an adjusted P-value <0.05 in both statistical approaches.
Gene expression data mining
Gene expression levels in several human tissues were obtained from the Roadmap Epigenomics project (https://egg2.wustl.edu/roadmap/web_portal/processed_data.html#RNAseq_uni_proc). The transcription levels in different brain regions were retrieved from the Gene Expression Omnibus (GEO) database (accession number GSE33587).
Chromatin analyses
Chromatin immunoprecipitation from glioma samples
Anti-H3K9ac (Millipore 06-942), -H3K4me3 (Diagenode 03-050), and -H3K27me3 (Millipore 07-449) antibodies were used to assess the respective marks at selected genes by ChIP of native chromatin isolated from glioma samples and controls as previously described (Maupetit-Méhouas et al. 2016). Input and antibody-bound fractions were quantified by real-time PCR amplification with the SYBR Green mixture (Roche) using a LightCycler 480II (Roche) instrument. Background precipitation levels were determined by performing mock precipitations with a nonspecific IgG antiserum (Sigma-Aldrich C-2288) and were only a fraction of the precipitation levels obtained with the specific antibodies. The bound/input ratios were calculated and normalized to the precipitation level at the TBP promoter for the anti-H3K9ac and -H3K4me3 ChIPs and at the SP6 promoter for the anti-H3K27me3 ChIP. The primers used are described in Supplemental Table S6.
ChIP-seq data mining associated with chromatin analyses
ChIP-seq data from NPC and brain samples were from the NIH Roadmap Epigenomics project (http://www.roadmapepigenomics.org/) detailed as follows:
-
For H9-derived NPCs: input (GSM772805), H3K4me3 (GSM77 2736), and H3K27me3 (GSM772801).
-
In brain: input (GSM772991), H3K4me3 (GSM772996), H3K27me3 (GSM772993), and H3K9me3 (GSM670005).
ChIP-seq data for the H3K4me3 and H3K27me3 profiles in glioma-derived cells were obtained from the GEO database (accession numbers: GSM1121888 and GSM1121885, respectively).
To describe the histone modification enrichment in each defect group, the ChIP-seq read densities around TSS (± 2 kb) were represented by a heatmap in which each line represents one single promoter. The mean signal around TSS (±2 kb) for each defect group was compared with the mean signal for all genes included in this study (n = 14,714) to correct for the bias attributable to the use of different data sets.
Gene classification according to their chromatin signature
The gene classification according to their chromatin signature (bivalent, H3K4me3-only, H3K27me3-only, and none) in human ES cells is from Court and Arnaud (2017). For NPC and brain samples, this classification was performed as previously described (Court and Arnaud 2017). Briefly, data for input, H3K4me3, and H3K27me3 ChIPs were aligned to the hg19 genome assembly. Peaks were then called with MACS 1.4.2 using the input as control for peak detection (Zhang et al. 2008). Chromatin signatures were classified using an in-house R scripts (Supplemental Code). Specifically, a bivalent region was defined by the overlapping of H3K4me3 and H3K27me3 peaks for at least 1 kb. H3K4me3-only and H3K27me3-only regions were identified as 1-kb peaks for H3K4me3 or H3K27me3 that did not overlap. The rest of the genome was considered as having a “none” chromatin signature. To attribute a chromatin signature to CGI/promoter regions, only the signature spreading across the CGI region was retained.
Copy number variation analyses
CNV analyses were performed using the Genome-Wide Human CytoScan HD Array (Affymetrix) and the samples analyzed by RNA-seq (three controls, eight IDHwt, and five IDHmut glioma samples). DNA and array data were processed by the Genomic Platform/Curie Institute according to the manufacturer's protocol. Arrays were scanned using an Affymetrix GeneChip Scanner 3000 7G. Scanned data files were analyzed with Affymetrix Chromosome Analysis Suite v3.1 (ChAS) (Affymetrix) using the CytoScanHD_Array.na33.annot.db annotation file on the hg19 genome. To find CNVs, single samples were analyzed using the reference model file “CytoScanHD_Array.na33.r2.REF_MODEL.” To prevent the detection of false-positive CNVs, only alterations that involved at least 50 consecutive probes and spanning >100 kb were considered in the analysis. To evaluate genetic alterations at gene level, the mean log2 ratio of the CNV fragment overlapping with a CGI/promoter region was assigned to each gene. For genes with a CGI/promoter region not covered by a CNV fragment, the mean log2 ratio was considered as null. Positive and negative mean log2 ratios were used to categorize duplicated and deleted regions, respectively. To identify genes, the expression alteration of which correlates (P < 0.05) with CNV changes, the mean log2 ratio of the CNV values and the normalized pseudo RNA-seq counts for each transcriptionally affected gene in the same sample were compared using the Pearson's correlation.
Principal component analysis
The PCA was done with the FactoMineR package using molecular features (histone modifications, DNA methylation, and expression) associated with genes from Meth+/Exp− and No Meth/Exp−the groups in brain (Le et al. 2008). For each gene, the histone modification levels at the CGI/promoter region (TSS ±1 kb) were based on the average ChIP-seq signal for H3K4me3, H3K27me3, and H3K9me3 obtained in brain samples. The DNA methylation levels were determined by the log2 value of the mean β-value, obtained from eight brain control samples, for all the probes located in the CGI of that gene. For the transcriptional level, the average FPKM value from three brain control samples was log2 transformed. Because the different variables used were defined by different measure units, they were standardized all before the PCA analysis (i.e., centered and scaled).
Functional annotations
InterPro protein functional classification analysis was performed using the functional annotation tools in DAVID 6.8 (https://david.ncifcrf.gov/) (Huang et al. 2009). Gene Ontology analyses were done with the GeneRanker tools of the Genomatix suite (http://www.genomatix.de/). To identify putative regulatory features linked to the transcriptional defect groups, the CGI positions in genes were analyzed with i-cis Target (https://gbiomed.kuleuven.be/apps/lcb/i-cisTarget/) (Herrmann et al. 2012). The comparative analysis tools were then used to identify specific binding sites. Because transcription factors could have multiple position weight matrices (PWM), all normalized enrichment scores (NES) given by i-cis Target for a factor were displayed in a box plot. The tumor suppressor gene and oncogene lists were obtained from www.ongene.bioinfo-minzhao.org, www.cta.lncc.br, and www.uniprot.org with the keywords “tumor suppressor” (KW-0043) and “proto-oncogene” (KW-0656).
Data access
Data generated in this study have been submitted to the NCBI Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) under the following accession numbers: GSE123678 for the HM450K DNA methylation data; GSE123682 for the Cytoscan HD data; and GSE123892 for the oriented RNA-seq data. Custom R scripts used to perform this study are available as Supplemental Code.
Acknowledgments
We thank the Clermont-Ferrand hospital's Neurosurgery department (Prof. J.J. Lemaire) for help in completing this study, the Platform “Gentyane” (http://gentyane.clermont.inra.fr/) for technical help with the Fluidigm 96.96 Dynamic Arrays, and Zachary Arnaud for precious input. We also thank Dr. David Monk and all members of P.A.’s team for critical reading of the manuscript. This study was funded by the Plan Cancer-INSERM (CS14085) (to L.K.-T., P.V., and P.A.), the Cancéropole CLARA (Oncostarter “Gliohoxas”) (to P.A.), the Fonds de dotation Patrick Brou de Lauriére (to P.A.), the Ligue Contre le Cancer comité de l'Ardéche et du Puy de Dôme (to F.C. and P.A.). F.C. and E.L.B. had a fellowship from Université Clermont Auvergne and La Ligue Nationale Contre le Cancer, respectively. This research is supported by the French government IDEX-ISITE initiative 16-IDEX-0001 (CAP 20-25).
Author contributions: F.C. and P.A. initiated and supervised the study. F.C., A.F., C.V.-B., E.C., L.K.-T., P.V., and P.A. designed the study. T.K. collected glioma samples from the operating room. J.-L.K., J.B., E.C., and P.V. characterized the glioma samples and updated the clinical follow-up of the patients. J.B. and E.C. collected the patient data. B.P. conducted the patients’ data statistical analyses. F.C., E.L.B., A.F., M.M.-B., and C.V.-B. performed the experiments. F.C. performed the bioinformatics analyses. F.C., E.L.B., A.F., M.M.-B., C.V.-B., E.C., and P.A. analyzed the data. F.C. produced the figures with C.V.-B.’s and P.A.’s input. P.A. wrote the paper with F.C.’s input. All authors read and approved the final manuscript.
Footnotes
-
[Supplemental material is available for this article.]
-
Article published online before print. Article, supplemental material, and publication date are at http://www.genome.org/cgi/doi/10.1101/gr.249219.119.
- Received February 19, 2019.
- Accepted August 27, 2019.
This article is distributed exclusively by Cold Spring Harbor Laboratory Press for the first six months after the full-issue publication date (see http://genome.cshlp.org/site/misc/terms.xhtml). After six months, it is available under a Creative Commons License (Attribution-NonCommercial 4.0 International), as described at http://creativecommons.org/licenses/by-nc/4.0/.