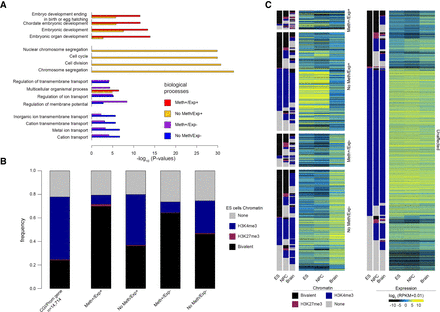
Genes with bivalent chromatin signature in ES cells are more prone to be deregulated in IDHwt glioma. (A) Gene Ontology terms (biological processes) enriched in genes from the four defect categories. For each category, the four highest terms are shown. (B) Distribution of genes of each defect category according to their chromatin signature in human ES cells: (none) gray; (bivalent) black; (H3K4me3-only) blue; (H3K27me3-only) purple. As reference, the distribution of the 14,714 genes analyzed in this study according to their chromatin signatures in human ES cells is shown in the left panel. (C) Expression level and chromatin signatures of genes of the four defect categories in human ES cells, neural progenitor cells (NPCs), and healthy brain. For comparison, the same analysis is provided on the right panel for genes without expression defect (unaffected) in IDHwt glioma samples.











