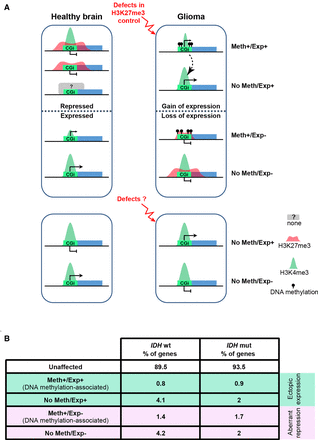
Working model. (A) In glioma, alterations in the control of the H3K27me3 signature could be one of the main contributors to the four types of transcriptional defects observed at CGI/promoter-controlled genes (upper). In this model, genome-wide hypomethylation induces H3K27me3 redistribution that could lead to ectopic expression of genes that are normally repressed by polycomb proteins, including some genes encoding transcription factors. These overexpressed transcription factors could then promote the aberrant expression of their target genes (dotted arrow). Similarly, alterations in the interplay between the polycomb complex and the transcriptional machinery could affect H3K27me3 fate during ES and/or neural stem cell differentiation. Specifically, this alteration could lead to the aberrant maintenance of bivalency and silencing at a subset of genes that are normally specifically expressed in brain. At genes that are normally poorly expressed in healthy brain, this process is associated with gain of DNA methylation in glioma. Beside defects in the H3K27me3 signature, we also identified a subset of genes that are apparently constitutively associated with H3K4me3-only, regardless of their expression status in brain and glioma (lower). The mechanisms underlying their transcriptional deregulation remain to be determined. (B) Percentage of unaffected and affected CGI/promoter-controlled genes for each of the four described defects in IDHwt and IDHmut glioma samples from our cohort.











