Orc4 spatiotemporally stabilizes centromeric chromatin
- Lakshmi Sreekumar1,8,
- Kiran Kumari2,3,4,
- Krishnendu Guin1,
- Asif Bakshi1,9,
- Neha Varshney1,10,
- Bhagya C. Thimmappa1,11,
- Leelavati Narlikar5,
- Ranjith Padinhateeri2,
- Rahul Siddharthan6 and
- Kaustuv Sanyal1,7
- 1Molecular Biology and Genetics Unit, Jawaharlal Nehru Centre for Advanced Scientific Research, Bangalore 560064, India;
- 2Department of Biosciences and Bioengineering, Indian Institute of Technology Bombay, Mumbai 400076, India;
- 3IITB-Monash Research Academy, Mumbai 400076, India;
- 4Department of Chemical Engineering, Monash University, Melbourne 3800, Australia;
- 5Department of Chemical Engineering, CSIR-National Chemical Laboratory, Pune 411008, India;
- 6The Institute of Mathematical Sciences/HBNI, Taramani, Chennai 600113, India;
- 7Graduate School of Frontier Biosciences, Osaka University, Suita, Osaka 565-0871, Japan
Abstract
The establishment of centromeric chromatin and its propagation by the centromere-specific histone CENPA is mediated by epigenetic mechanisms in most eukaryotes. DNA replication origins, origin binding proteins, and replication timing of centromere DNA are important determinants of centromere function. The epigenetically regulated regional centromeres in the budding yeast Candida albicans have unique DNA sequences that replicate earliest in every chromosome and are clustered throughout the cell cycle. In this study, the genome-wide occupancy of the replication initiation protein Orc4 reveals its abundance at all centromeres in C. albicans. Orc4 is associated with four different DNA sequence motifs, one of which coincides with tRNA genes (tDNA) that replicate early and cluster together in space. Hi-C combined with genome-wide replication timing analyses identify that early replicating Orc4-bound regions interact with themselves stronger than with late replicating Orc4-bound regions. We simulate a polymer model of chromosomes of C. albicans and propose that the early replicating and highly enriched Orc4-bound sites preferentially localize around the clustered kinetochores. We also observe that Orc4 is constitutively localized to centromeres, and both Orc4 and the helicase Mcm2 are essential for cell viability and CENPA stability in C. albicans. Finally, we show that new molecules of CENPA are recruited to centromeres during late anaphase/telophase, which coincides with the stage at which the CENPA-specific chaperone Scm3 localizes to the kinetochore. We propose that the spatiotemporal localization of Orc4 within the nucleus, in collaboration with Mcm2 and Scm3, maintains centromeric chromatin stability and CENPA recruitment in C. albicans.
The kinetochore is a multiprotein complex that assembles on centromeric chromatin and provides the chromosomal platform for spindle microtubule binding during chromosome segregation (Musacchio and Desai 2017). In a majority of the eukaryotes, centromeres are epigenetically specified by the centromeric histone variant CENPA (Cse4 in yeast), in a manner independent of the underlying DNA sequence (Yadav et al. 2018a). Despite performing an evolutionarily conserved function of assembling the kinetochore, the establishment of centromeric chromatin and its subsequent propagation through many generations depends on a variety of factors that are often species specific. These include centromere-specific DNA sequence elements, replication timing, and transcriptional status of centromere DNA, as well as the influence of proteins involved in DNA replication, repair, recombination, and RNA interference (Yadav et al. 2018a). More recently, spatial chromosomal arrangements have been implicated as a determinant of centromere identity (Burrack et al. 2016; Sreekumar et al. 2019; Guin et al. 2020).
A particularly striking feature of centromere DNA is its replication timing. Whereas metazoan cells replicate centromeres during late S phase (Ten Hagen et al. 1990), centromeres replicate at early S phase in fungi (Pohl et al. 2012). This temporal distinction in the centromere DNA replication in two major eukaryotic kingdoms can be attributed to the differential timing of firing of DNA initiation sites or DNA replication origins within the genome. Replication origins initiate DNA replication with the help of the prereplication complex (pre-RC), two major components of which are the origin recognition complex (Orc1-6) and the minichromosome maintenance complex (Mcm2-7) (Leonard and Méchali 2013). Although ORC proteins are associated with replication origins throughout the cell cycle (Bell and Stillman 1992; Dutta and Bell 1997), MCM proteins move along the replication fork (Dutta and Bell 1997; Bell and Dutta 2002; Forsburg 2004). Within the nucleus, regions that have similar replication timing tend to cluster to facilitate coregulation (Duan et al. 2010; Gong et al. 2015; Eser et al. 2017). Several studies indicate that fungal centromeres cluster either at specific stages or throughout the cell cycle, toward the nuclear periphery and close to the spindle pole bodies (SPBs) to facilitate chromosomes adopting the Rabl configuration (Rabl 1885; Jin et al. 2000; Kozubowski et al. 2013; Guin et al. 2020). Conservation in replication timing and clustering patterns of centromeres not only preserves the kinetochore integrity but may help in determining the site of centromere formation in subsequent cell divisions.
A growing body of evidence suggests that DNA replication origins and replication initiation proteins crosstalk with centromeres. In fungal species such as Saccharomyces cerevisiae, Schizosaccharomyces pombe, and Yarrowia lipolytica, a centromere-linked replication origin helps in the early replication of centromeres (Vernis et al. 1997; Patel et al. 2006; Koren et al. 2010). In S. cerevisiae, kinetochores orchestrate early S phase replication of centromeres (Natsume et al. 2013). Following DNA replication, new molecules of CENPA are efficiently targeted to the replicated centromere DNA by a CENPA-specific chaperone, the Holliday junction recognition protein (HJURP) in mammals or Scm3 in yeast (Kato et al. 2007). HJURP stabilizes soluble CENPA-H4 dimers before they are incorporated into centromeric nucleosomes (Dunleavy et al. 2009; Foltz et al. 2009) with the help of the Mis18 complex (Fujita et al. 2007). The timely deposition of CENPA by HJURP/Scm3 occurs at distinct stages in the mitotic cell cycle, for instance, during S phase in S. cerevisiae (Pearson et al. 2004), during G2 in S. pombe (Shukla et al. 2018), and during G1 in humans (Foltz et al. 2009). In humans, DNA replication is also used as an error correction mechanism for loading new CENPA molecules (Nechemia-Arbely et al. 2019). This temporal regulation of CENPA deposition necessitates the study of factors common to both the local assembly of CENPA and genome-wide replication timing patterns.
Candida albicans is a diploid budding yeast and a human pathogen that has eight pairs of chromosomes. One of the striking features of the C. albicans genome is that each of its eight chromosomes has a 3- to 5-kb unique and different DNA sequence enriched with CENPA (Sanyal and Carbon 2002; Sanyal et al. 2004; Baum et al. 2006). Centromeres themselves do not harbor an active replication origin (Mitra et al. 2014) but are the earliest replicating regions on each chromosome (Koren et al. 2010). They constitutively cluster toward the nuclear periphery close to the SPBs at a defined space, forming a CENPA-rich zone or CENPA cloud (Sanyal and Carbon 2002; Thakur and Sanyal 2013; Guin et al. 2020). The constitutive kinetochore ensemble gets disintegrated and CENPA molecules are degraded if any of the essential kinetochore proteins are depleted (Thakur and Sanyal 2012). The epigenetic specification of centromeres in this organism has been exemplified by the efficient activation of neocentromeres at centromere-proximal regions (Thakur and Sanyal 2013). Additionally, a physical interaction between homologous recombination proteins and CENPA has been proved to render stability to centromeric chromatin (Mitra et al. 2014). In the absence of the functional RNAi machinery, heterochromatin factors, CENPA loading factors like Mis18, and conserved centromere DNA sequence cues, what determines centromeric chromatin establishment and its subsequent propagation in C. albicans remains an enigma. Hence, this study focuses on dissecting factors that help in the stability of centromere chromatin in C. albicans.
Results
Orc4 binds to discrete regions uniformly across the C. albicans genome
To examine the replication landscape of the C. albicans genome, we sought to determine the genome-wide occupancy of Orc4. Orc4 in C. albicans is a 564-aa-long protein (Padmanabhan et al. 2018) that contains the evolutionarily conserved AAA+ ATPase domain (Supplemental Fig. S1A; Walker et al. 1982). We raised polyclonal antibodies in rabbits against a peptide from the N terminus of native Orc4 (aa 20–33) of C. albicans (Supplemental Fig. S1B). Western blot of the whole-cell extract of C. albicans SC5314 (ORC4/ORC4) yielded a strong specific band at the expected molecular weight of ∼64 kDa when probed with custom-made and purified anti-Orc4 antibodies (Supplemental Fig. S1C). Indirect immunofluorescence microscopy using anti-Orc4 antibodies revealed that Orc4 was strictly nuclear-localized at all stages of the C. albicans cell cycle (Fig. 1A), a feature conserved in the ORC proteins of S. cerevisiae (Dutta and Bell 1997).
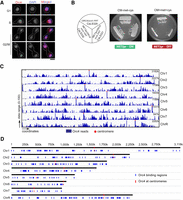
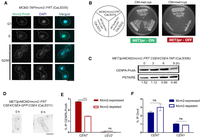
Orc4, an essential subunit of the origin recognition complex, is nuclear-localized and binds to discrete loci in the C. albicans genome. (A) Nuclear localization of Orc4 in C. albicans SC5314 (ORC4/ORC4) cells as evidenced by staining with anti-Orc4 antibodies and DAPI. Scale bar, 5 µm. (B) The promoter of MET3 in C. albicans, expressed in the absence of methionine and cysteine and repressed in the presence of both amino acids, was used for the conditional expression of ORC4. CaLS329 (ORC4/orc4::FRT) with one deleted copy of ORC4 and two independent transformants, CaLS330 and CaLS331 (MET3prORC4/orc4::FRT), where the remaining wild-type copy was placed under the control of the MET3 promoter, were streaked on plates containing permissive (CM-met-cys) or nonpermissive (CM + 5 mM met + 5 mM cys) media and photographed after 48 h of incubation at 30°C. (C) ChIP-sequencing analysis revealed that Orc4 was bound to discrete genomic sites in C. albicans. The total Orc4 reads (blue histogram) were obtained by subtracting the relative number of sequencing reads from the whole-cell lysate from the Orc4 ChIP sequence reads and aligning them to the reference genome C. albicans SC5314 assembly 21. Red dots indicate centromeres. (D) Orc4 binding regions (blue) on each of the eight C. albicans chromosomes including all eight centromeres (red).
Orc4 is an evolutionarily conserved essential subunit of ORC across eukaryotes (Chuang and Kelly 1999; Dai et al. 2005). A conditional mutant of orc4 in C. albicans CaLS330 (MET3prORC4/orc4::FRT), constructed by deleting one allele and replacing the endogenous promoter of the remaining ORC4 allele with the repressible MET3 promoter of C. albicans (Care et al. 1999), was unable to grow in nonpermissive conditions (Fig. 1B). Hence, Orc4 is essential for viability in C. albicans. We confirmed the depletion of Orc4 protein levels from the cellular pool by performing a western blot analysis in the Orc4 repressed compared with expressed conditions (Supplemental Fig. S1D). Subsequently, we used the purified anti-Orc4 antibodies as a tool to map its binding sites across the C. albicans genome. Orc4 chromatin immunoprecipitation (ChIP) sequencing in asynchronously grown cells of C. albicans yielded a total of 417 discrete binding sites with 414 of these belonging to various genomic loci, whereas the remaining three mapped to mitochondrial DNA (Figs. 1C,D; Supplemental Dataset S1). We validated a subset of highly and weakly Orc4-enriched regions by ChIP-qPCR assays (Supplemental Fig. S1E). All centromeres were highly enriched with Orc4. Although most of the binding loci (more than 300) spanned ∼1 kb in length, all eight centromeres had an Orc4 occupancy spanning 3–4 kb (Supplemental Fig. S1F).
Orc4 displays differential DNA binding modes that are spatiotemporally positioned in the genome
We used the de novo motif discovery tool DIVERSITY (Mitra et al. 2018) on the C. albicans Orc4 binding regions. DIVERSITY allows for the fact that the profiled protein may have multiple motifs/modes of DNA binding. Here, DIVERSITY reported four binding modes (Fig. 2A). Mode A is a strong motif GAnTCGAAC, present in 50 regions, 49 of which were found to be located within tRNA gene bodies (tDNAs) and one within the tDNA regulatory region. The most enriched nucleotides of mode A correspond to the third hairpin loop of the tRNA clover structure. The other three modes were low complexity motifs, TGATGA (mode B), CAnCAnCAn (mode C), and AGnAG (mode D). Each of the 417 binding regions was associated with one of these four modes. Mode C has been identified in a previous study (Tsai et al. 2014) in which ORC binding sites in the C. albicans genome were mapped using a microarray-based approach. ORC binding regions of these two studies share the maximum overlap at mode A (Supplemental Fig. S2A). Taken together, these results suggest that Orc4 in C. albicans is not specified by a single DNA binding site but rather displays differential DNA binding modes.
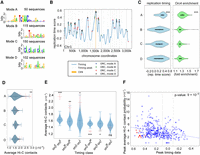
Orc4 is associated with four different DNA binding modes that are spatiotemporally located across the genome. (A) The four different modes identified by DIVERSITY (modes A, B, C, D) and their distribution across the 414 Orc4 binding regions in the C. albicans genome have been listed. (B) Orc4 ChIP-seq peaks denoted as asterisks, colored according to the four modes identified by DIVERSITY, were overlaid on the replication timing profile of Chr 1 in C. albicans from a previous study (Koren et al. 2010). A higher time score indicates replication at early S phase. Color-coded stars indicate each of the four motifs identified by DIVERSITY, which covers all the chromosomal sites. Light gray lines indicate local maxima in replication time. (CEN) Centromeres. (C) Violin plots depicting the replication timing scores (blue) (Koren et al. 2010) and Orc4 enrichment (green) of all the Orc4 peaks classified according to each of the four modes. (***) P-value < 0.0001. (D) Average Hi-C interactions (Burrack et al. 2016) of Orc4 peaks with other peaks in the same mode. Solid red indicates mean; dotted red, standard error; violins are from 1000 sets of randomized data (randomly selected genomic regions with the same size and chromosomal distribution as the peaks in that mode). (**) P-value < 0.001. (E) Mean Hi-C interactions (solid red) with standard error (dotted red) within and across each of the three timing classes (orcE, orcM, and orcL). These indicated higher interaction values within orcE and within orcL domains. Blue violins indicate mean interactions across 1000 randomizations, as in D. (***) P-value < 0.0001; (**) P-value < 0.001; (*) P-value < 0.01; (ns) P-value > 0.5. (F) A scatter plot of Hi-C contacts, replication timing, and Orc4 fold enrichment values of Orc4 binding regions. Each blue/red dot is an individual Orc4 peak, with its color intensity corresponding to its ChIP-seq enrichment value; red dots are peaks overlapping eight centromeres. The y-axis (peak average Hi-C contacts) represents the average of 10 best contacts for each peak of Orc4. The Pearson correlation coefficient is 0.27.
To categorize the replication timing of Orc4-bound regions, we used the available fully processed replication timing profile of the C. albicans genome (Koren et al. 2010). Based on the replication time of the entire genome, the first one-third (33.3%) of the replicating regions were classified as early, the second one-third were mid, and the remaining were late replicating regions. By comparing the Orc4 sites to this profile, we found 218 early or orcE sites (∼52% of the total), 127 mid or orcM sites (∼31%), and 69 late or orcL sites (∼17%) (Fig. 2A). We then overlaid the DIVERSITY modes onto the timing profile (Fig. 2B; Supplemental Fig. S2B). We observed a significantly early replication timing of the tRNA associated modes (mode A) (Fig. 2C). The other three modes (B, C, D) displayed no significant bias toward an early replication score. Moreover, we could correlate early replication timing with increased enrichment of Orc4 in these regions (Fig. 2C). Besides, all the modes were located toward the local maxima of the timing peaks (Fig. 2B).
To map the interactions made by the Orc4 binding regions with each other, we used high-throughput chromosome conformation capture (Hi-C) data from a previous study in C. albicans (Burrack et al. 2016). All the Orc4 binding regions were aligned in increasing order of their replication timing (early to late), and similar analyses were performed for the whole genome of C. albicans. We observed that the overall “only-ORC” interactions were higher than the whole-genome “all” interactions, suggesting that Orc4 binding regions interacted more frequently than the genome average (Supplemental Fig. S2C,D). Hi-C analysis also revealed that the mode A sites formed stronger interactions among themselves than all the other modes (Fig. 2D). Because mode A is associated with tRNA genes (tDNA), a comparative analysis of the contact probabilities of mode A with the rest of the tDNAs in the genome revealed a higher interaction of mode A tDNAs over the rest (Supplemental Fig. S2E,F), but that was not significant. Additionally, there was a significant increase in interaction frequencies within similarly timed domains (orcE-orcE; orcM-orcM; orcL-orcL) compared with interactions across domains (Fig. 2E). These interactions were conserved and showed a similar trend even when the peaks corresponding to centromeres were removed from the analysis (Supplemental Fig. S2G). Upon arraying the Orc4 peaks according to their replication timing scores reported previously (Koren et al. 2010) against the average Hi-C interaction frequencies, we could observe a weak but significant correlation between contact probability and replication timing (Fig. 2F). We also found higher Orc4 enrichment in the orcE-orcM regions as the majority of the Orc4 peaks lay in the middle of the pack (Fig. 2F). Taken together, our analyses suggest that Orc4-bound regions with a similar replication timing tend to associate together, and this association is independent of the physical clustering of centromeres, the strongest higher-order chromosomal interactions in the genome (Sreekumar et al. 2019).
Polymer modeling of C. albicans chromosomes reveals a replication timing–driven spatial positioning of Orc4 within the nucleus
Hi-C analysis is not sufficient to reveal the positioning of a particular locus within the nucleus. The intra- and inter-chromosomal interaction frequencies can be converted to linear distance approximations and averaged across populations to generate computational models that yield an ensemble of genomic conformations (Berger et al. 2008; Gürsoy et al. 2017). To study the higher-order chromosomal organization of the C. albicans genome, we resorted to polymer modeling of chromosomes using the contact probability data from the published Hi-C experiment (Burrack et al. 2016). To do this, we used a statistical approach in which each bead-pair is either bonded or not bonded based on the available contact probability data (Supplemental Table S1). At first, we compared the experimental Hi-C data (Supplemental Fig. S3A) with the contact probability data obtained from our simulation of 1000 different configurations (Supplemental Fig. S3B) to ensure that our simulation could satisfactorily recover the input contact matrix. Contact probability for a bead-pair (i,j) from the simulation is calculated by averaging the bonding function bij over 1000 realizations. The function bij = 1, in the case of a contact (rij < 1.5 l0) and zero otherwise. Here, the rij is the spatial distance between bead i and bead j, and l0 is the natural extension of the connector spring. The contact probability from the simulation was found to be in close agreement with the Hi-C data, showing the reliability of the model (Supplemental Fig. S3A,B). From our simulations, we could predict the average spatial distance between any two beads within the genome. For a given contact probability, the corresponding average spatial distance could be computed (Supplemental Fig. S3C). We fixed the position of one of the centromere beads as the reference and hence sought to determine the spatial (three-dimensional [3D]) location of each of these beads.
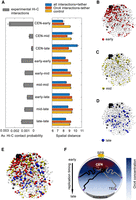
To examine the genome-wide distribution of Orc4 in the nuclear space, we mapped the Orc4 ChIP-seq peaks to the corresponding coarse-grained beads. By using our simulation, we computed 3D locations of the Orc4 binding sites mapped onto the early, mid, and late replicating regions of the genome (as categorized previously). From the experimentally obtained contact probability, it was observed that orcE sites show stronger interactions with centromeres compared with orcM and orcL sites (Fig. 3A). Our simulations show the corresponding distances between the above-mentioned regions, where the average distance between the orcE sites with centromeres is significantly shorter than the average distance between centromeres and orcM/orcL regions. To examine the contribution of Orc4 in these interactions, we simulated only the Orc4 interactions in a system in which none of the centromere beads were attached to the nuclear membrane, in which case the centromere cluster can explore the nucleus without constraint. When compared with the control in which we simulated the polymer chain without any interactions (without tethering to SPB), we observed that Orc4 interactions alone can initiate clustering of centromeres and establish signatures of prominent Hi-C interactions. This can facilitate centromeric interactions to early, mid, and late replicating regions in the genome (Fig. 3A; Supplemental Fig. S3D,E). To visualize the location of binding sites of orcE, orcM, and orcL with respect to centromeres and telomeres, we chose one random configuration from an ensemble of 1000 configurations. The orcE regions are relatively closer to centromeres (Fig. 3B), the orcM sites are farther away (Fig. 3C), and orcL sites are the farthest from centromeres (Fig. 3D). We also examined the linear distance of orcE, orcM, and orcL from centromeres for all chromosomes (Supplemental Fig. S3F,G), and specifically for Chr 2 (metacentric) and Chr 6 (acrocentric) (Supplemental Fig. S3H,I), and found this trend to be comparable with the trend observed in 3D distances. Hence, there is a replication time–driven spatial distribution of Orc4 along the chromosomes with the highest concentration near centromeres that decreases toward telomeres (Fig. 3E). Taken together, the nonrandom distribution of Orc4 in the nucleus of C. albicans from our computational model is largely suggestive of a specific spatial organization driven by the replication timing of Orc4 occupied loci (Fig. 3F; Supplemental Movie S1). The interactions mediated by Orc4 help to stabilize the centromeric cluster even in the absence of a nuclear envelope tether, making Orc4 an efficient genome organizer in this organism.
Early replicating Orc4-bound regions are spatially proximal to the clustered centromeres. (A, left) The average contact probability for the indicated region for each of the timing classes (across and within orcE, orcM, and orcL domains) showed stronger CEN-orcE interactions. (Right) The average spatial distances between the indicated regions calculated from the Langevin simulation for each of the timing classes (across and within the orcE, orcM, and orcL domains) when all Hi-C interactions (blue), only the Orc4-Orc4 interactions (red), and control (yellow) simulation for a random configuration are performed. (B–E) Snapshot of the 3D configuration of the C. albicans genome (one out of 1000 realizations) from the simulations shows all chromosomes in light gray, centromeres in black, and telomeres in dark gray. The orcE regions are shown in red (B), orcM in yellow (C), and orcL in blue (D); E represents all the Orc4 binding sites (orcE, orcM, and orcL). (F) Schematic of a budding yeast nucleus showing the typical Rabl configuration in which the clustered centromeres are anchored near SPBs and telomeres are often away from the centromere cluster and occasionally interacting with the nuclear envelope. In C. albicans, the highest spatial enrichment of Orc4 is near the centromere cluster. Orc4 concentration gradually diminishes toward the opposite pole. Concomitantly, early replicating regions are located toward centromeres and the late regions are toward telomeres. (CEN) Centromeres; (TEL) telomeres.
Constitutive localization of Orc4 at centromeres stabilizes CENPA
The existence of a high Orc4-enriched zone around the centromere cluster and the strong enrichment of Orc4 at all centromeres identified by ChIP-seq analysis prompted us to examine its biological significance in C. albicans. Upon comparison of the Orc4 enrichment with CENPA occupancy in C. albicans, there was a striking overlap in the binding regions of these two proteins, indicating a strong physical association of Orc4 at centromeres (Fig. 4A; Supplemental Fig. S4A; Supplemental Table S2). To validate its role in centromere establishment, we examined whether its binding is required for centromere formation at a nonnative locus. C. albicans can efficiently activate neocentromeres at centromere-proximal regions when a native centromere is deleted (Thakur and Sanyal 2013). Orc4 was found to be enriched at the neocentromere hotspots nCEN7-I and nCEN7-II when the 4.5-kb CENPA-rich region of CEN7 was deleted but was absent at these loci in a wild-type strain with unaltered CEN7 (Supplemental Fig. S4B). This suggests that Orc4 helps in centromere establishment, and further hints toward the possible role of replication initiator proteins in specifying centromeres in C. albicans.
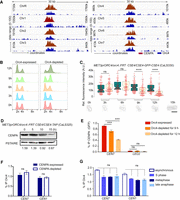
Centromeric localization of Orc4 stabilizes CENPA. (A) A 30-kb region harboring each centromere (x-axis) was plotted against the subtracted ChIP sequencing reads (y-axis) for CENPA (red) and Orc4 (blue). (B) Flow cytometric analyses of orc4 mutants in Orc4-expressed versus Orc4-depleted conditions at the indicated time of incubation. (C) Relative fluorescence intensities of CENPA (GFP) clusters in orc4 mutant CaLS330 grown at indicated time points in permissive (green) and nonpermissive (yellow) conditions show a significant reduction of GFP upon depletion of Orc4. Scale bar, 10 µm. t-test; (****) P-value < 0.0001, (ns) P-value > 0.05; n ≥ 100. (D) Western blot of the whole-cell lysate of CaLS325 (METprORC4/orc4::FRT CSE4/CSE4-TAP) using anti–Protein A antibodies showed a time-dependent decrease in CENPA levels upon Orc4 depletion when normalized with the loading control, PSTAIRE. (E) ChIP-qPCR using anti-GFP (CENP-A) antibodies revealed reduced CENPA enrichment at the centromere upon Orc4 depletion in CaLS330 when grown in nonpermissive media for 9 and 12 h. Two-way ANOVA; (***) P-value < 0.001; (ns) P-value > 0.05; n = 3. (F) Orc4 ChIP-qPCR revealed no significant reduction in centromeric Orc4 when CENPA was depleted for 8 h in YP with dextrose in the strain CAKS3b (cse4/PCK1prCSE4) (Sanyal and Carbon 2002). Two-way ANOVA; (ns) P-value > 0.05; n = 3. (G) ChIP-qPCR revealed the Orc4 enrichment at CEN7 in various stages of the cell cycle: hydroxyurea treated (S phase), nocodazole treated (metaphase), cdc15 mutant (late anaphase). Percentage IP values for Orc4 ChIP at CEN7 were normalized with noncentromeric regions enriched with Orc4. One-way ANOVA; (***) P-value < 0.001; (ns) P-value > 0.05; n = 3.
Next, we sought to characterize the effect of Orc4 depletion on the cell cycle. Flow cytometric analysis showed a G2/M peak 12 h post depletion of the protein, indicative of a late S phase or mitotic arrest in the orc4 mutant CaLS330 (Fig. 4B). Upon assaying for CENPA localization in the orc4 conditional mutant (Fig. 4C), we found a significant drop in CENPA (GFP) signal intensity 9 h post depletion. This was supported by western blot (Fig. 4D) and ChIP-qPCR (Fig. 4E) analyses. However, depletion of CENPA did not alter the levels of centromeric Orc4 (Fig. 4F), indicating that Orc4 regulates the stability of centromeric chromatin and not vice versa. We then determined the enrichment levels of Orc4 at centromeres by ChIP-qPCR assays in cells arrested at the S phase, metaphase, and late anaphase stages in comparison with asynchronous cells and observed no significant difference across all stages (Fig. 4G). These results suggest that Orc4 is constitutively localized to the kinetochore across the cell cycle in C. albicans. Spot dilution assays to determine the viability of orc4 mutant (CaLS330) after prolonged depletion until 24 h revealed no observable drop in the viability of orc4 mutant (Supplemental Fig. S4C) but an increased rate of chromosome missegregation in >60% of the cells (Supplemental Fig. S4D). To rule out the possibility that general replication stress can lead to loss of CENPA, we treated the cells with an S phase inhibitor, hydroxyurea (HU), for 2 h and quantified the mean GFP intensity in the strain YJB8675 (CSE4-GFP-CSE4/CSE4) (Joglekar et al. 2008). Compared with the untreated control, we could not detect any significant difference in the GFP intensity upon HU treatment (Supplemental Fig. S4E). This was further corroborated by performing a western blot (Supplemental Fig. S4F) and ChIP-qPCR analyses under the same conditions (Supplemental Fig. S4G). These results together suggest that Orc4 helps to establish centromere identity and that its constitutive centromeric localization is essential to maintain CENPA chromatin stability.
Mcm2 is essential for chromosome segregation and CENPA stability
Having established Orc4, a pre-RC subunit, as a regulator of CENPA chromatin, we sought to examine the role of another pre-RC subunit, Mcm2, on CENPA stability in C. albicans. Apart from its canonical helicase activity during replication initiation, Mcm2 is known to bind to canonical and variant histone H3 in vitro (Huang et al. 2015). BLAST analysis using S. cerevisiae Mcm2 as the query sequence revealed that ORF19.4354 translates to a 101.2-kDa protein that contains the conserved Walker A, Walker B, and the R finger motifs together, constituting the MCM box (Supplemental Fig. S5A,B; Forsburg 2004). We tagged Mcm2 with Protein A at the C terminus in the C. albicans strain BWP17 (Wilson et al. 1999) and then deleted the untagged allele of MCM2 to generate a singly Protein A–tagged strain, CaLS334 (MCM2-TAP/mcm2::FRT). Western blot analysis with the tagged protein lysate yielded a specific band at the expected molecular weight of 135 kDa, which could not be detected in the untagged lysate (Supplemental Fig. S5C). By indirect immunofluorescence microscopy, Mcm2–Protein A was found to colocalize with the nucleus in the G1 and S phases of the cell cycle (Fig. 5A). The localization was confirmed by microscopic examination of a C-terminally GFP-tagged strain of Mcm2, CaLS341 (MCM2-GFP/MCM2) (Supplemental Fig. S5D). In large budded cells, Mcm2 could also be visualized in the cytoplasm, along with a diffused nuclear signal (Fig. 5A; Supplemental Fig. S5D). We could localize Mcm2 in the nuclei of large budded cells where nuclear separation has occurred (late anaphase) between the mother and daughter buds (Fig. 5A; Supplemental Fig. S5D), reminiscent of MCM proteins in S. cerevisiae (Yan et al. 1993). We performed Mcm2 ChIP-qPCR analysis with primers corresponding to a few of the Orc4 binding regions on each of the eight chromosomes and could detect four out of eight sites to be significantly enriched with Mcm2 over a control region (Supplemental Fig. S5E). Further analysis with more binding sites will reveal if there is a sequence preference for Mcm2 in the genome of C. albicans.
Mcm2 is essential for cell viability and CENPA stability in C. albicans. (A) Intracellular localization of Mcm2–Protein A in CaLS335 (MCM2-TAP/mcm2::FRT) cells stained with anti–Protein A antibodies and DAPI. Scale bar, 5 µm. (B) CaLS310 (MCM2/mcm2::FRT), where one copy of MCM2 was deleted, and two independent transformants, CaLS311 and CaLS312 (MET3prMCM2/mcm2::FRT), where the remaining wild-type copy was placed under the control of the MET3 promoter, were streaked on plates containing permissive (CM-met-cys) and nonpermissive (CM + 5 mM cys + 5 mM met) media and photographed after 48 h of incubation at 30°C. (C) Western blot of the whole-cell lysate of CaLS306 (MET3prMCM2/mcm2::FRT CSE4/CSE4-TAP) using anti–Protein A antibodies revealed a time-dependent decrease in CENPA levels upon depletion of Mcm2 for 3, 6, and 9 h. Normalization was performed using PSTAIRE. (D) CENPA (GFP) cluster delocalizes upon depletion of Mcm2 in CaLS311 (MET3prMCM2/mcm2::FRT CSE4-GFP-CSE4/CSE4). Scale bar, 5 µm. n = 100. (E) ChIP-qPCR using anti–Protein A antibodies revealed a significant reduction of CENPA at CEN7 in CaLS306 when grown in nonpermissive media for 6 h. Two-way ANOVA; (***) P-value < 0.001; (ns) P-value > 0.05; n = 3. (F) ChIP-qPCR in CaLS311 revealed no significant difference in the Orc4 enrichment at the centromeres in the presence or absence of Mcm2. Two-way ANOVA; (***) P-value < 0.001; (ns) P-value > 0.05; n = 3.
To determine the essentiality of Mcm2 in C. albicans, we constructed a conditional mutant of mcm2, CaLS311 (MET3prMCM2/mcm2::FRT), by deleting one allele and replacing the endogenous promoter of the remaining MCM2 allele with the MET3 promoter (Fig. 5B; Care et al. 1999). Mcm2 was found to be essential for viability in C. albicans. Western blot analysis in the Mcm2-expressed versus Mcm2-repressed conditions confirmed the depletion of the protein levels of Mcm2 by 6 h (Supplemental Fig. S5F). We observed a drastic decline in the viability (Supplemental Fig. S5G) and an increased rate of missegregation of chromosomes in the mcm2 mutant after 6 h of depletion (Supplemental Fig. S5H). We wanted to probe the cause of chromosome missegregation by examining the effect of Mcm2 depletion on CENPA stability. Depletion of Mcm2 led to a concomitant reduction in CENPA protein levels (Fig. 5C), like the previous observation upon Orc4 depletion, shedding light on a previously unknown candidate to preserve kinetochore stability. We also observed declustering of the kinetochore architecture in 90% of the mcm2 mutant cells (Fig. 5D). ChIP-qPCR analysis revealed a >50% reduction in the centromere-bound CENPA following Mcm2 depletion (Fig. 5E). The centromeric occupancy of Orc4 was unaltered upon Mcm2 depletion (Fig. 5F). Hence, Mcm2 is required for CENPA stability but is dispensable for the centromeric binding of Orc4.
The CENPA chaperone Scm3 loads CENPA during late anaphase/telophase in C. albicans
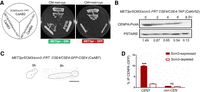
Having identified previously unknown factors of the pre-RC regulating CENPA stability, we wanted to examine the de novo loading of CENPA at the kinetochore in C. albicans. To address this, we sought to characterize the homolog of the CENPA chaperone Scm3/HJURP in C. albicans. BLAST analysis using S. cerevisiae Scm3 as the query sequence revealed that ORF19.668 translates to a protein of ∼72 kDa containing the CENPA-interacting Scm3 domain (Sanchez-Pulido et al. 2009) that was found to be conserved in C. albicans (Supplemental Fig. S6A). Additionally, there were three separate C2H2 zinc finger clusters present toward the C terminus of Scm3 in C. albicans that were absent in S. cerevisiae (Supplemental Fig. S6B; Aravind et al. 2007). We constructed a depletion mutant of scm3 by replacing the endogenous promoter of the intact copy with the MET3 promoter in a heterozygous null strain, CaAB3 (MET3prSCM3/scm3::FRT; as described in previous sections) and found Scm3 to be essential for viability in C. albicans (Fig. 6A). We observed the gradual degradation of CENPA protein levels by western blot following the depletion of Scm3 (Fig. 6B). Microscopic examination of CENPA revealed declustering of kinetochores in ∼75% of the cells post 8 h of Scm3 depletion, a phenotype observed upon depletion of any of the essential kinetochore proteins in C. albicans (Fig. 6C; Thakur and Sanyal 2012). Additionally, ChIP-qPCR analysis revealed a drastic reduction of CENPA from centromeres upon Scm3 depletion (Fig. 6D). Hence, Scm3 stabilizes CENPA in C. albicans.
Scm3 is essential for stability of CENPA in C. albicans. (A) The C. albicans strain CaAB2 (SCM3/scm3::FRT), where one copy of SCM3 has been deleted, and two independent transformants CaAB3 and CaAB4 (MET3prSCM3/scm3::FRT), where the remaining allele was placed under the control of the MET3 promoter, were streaked on plates containing permissive (CM-met-cys) and nonpermissive (CM + 1 mM met + 1 mM cys) media and photographed after 48 h of incubation at 30°C. (B) Western blot showing protein levels of CENPA upon depletion of Scm3 for the indicated time points normalized to PSTAIRE. (C) Dissociation of the CENPA (GFP) cluster marking disintegration of the kinetochore in CaAB7 (MET3prSCM3/scm3::FRT CSE4-GFP-CSE4/CSE4) cells grown in nonpermissive conditions for the indicated time. Scale bar, 5 µm. n = 100. (D) ChIP-qPCR using anti-GFP (CENPA) antibodies revealed reduced CENPA enrichment at CEN7 in CaAB7 when grown in nonpermissive media for 8 h. Two-way ANOVA; (***) P-value < 0.001; (ns) P-value > 0.05; n = 3.
To study the subcellular localization of Scm3, we generated a strain CaLS342 (SCM3-2× GFP(URA3)/scm3::SAT1 NDC80-RFP(ARG4)/NDC80) in which one allele of SCM3 was deleted; the remaining copy of SCM3 was tagged at the C terminus with 2×GFP, and a kinetochore protein Ndc80 was tagged with RFP. Microscopic examination of CaLS342 revealed Scm3 signals as a distinct punctum in the nucleus colocalizing with Ndc80 during late anaphase/telophase and G1 stages of the C. albicans cell cycle (Fig. 7A). Scm3 localization was undetected at other stages as well as in nocodazole-treated (metaphase-arrested) cells (Fig. 7A).
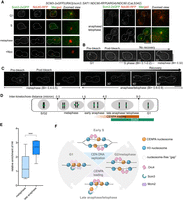
Scm3 loads CENPA during late anaphase/telophase in C. albicans. (A) Localization of Scm3 in CaLS342 (SCM3-2×GFP(URA3)/scm3::SAT1 NDC80-RFP(ARG4)/NDC80) cells coexpressing Scm3-2xGFP and a kinetochore marker, Ndc80-RFP, at various stages of the cell cycle. During late anaphase/telophase through G1 stage of the next cycle, Scm3 colocalizes with the kinetochore cluster. Absence of Scm3 at the unsegregated kinetochore cluster in the metaphase-arrested CaLS342 cells upon treatment with 20 µg/mL nocodazole (NOC; bottom, left panel). Scale bar, 5 μm. (B) A representative image of a G1 cell subjected to targeted photobleaching of CENPA (GFP) centromeric cluster. Images were captured at the indicated time points. GFP fluorescence could not be recovered even after 30 min of budding. Stages have been categorized according to the budding index (BI) and inter-kinetochore distance, as shown in Supplemental Figure S7B. Scale bar, 5 μm. n = 6. (C) A representative image of a metaphase cell subjected to targeted photobleaching of CENPA (GFP) centromeric cluster. Images were captured at the indicated time points. Fluorescence recovery could be observed after 30 min post bleaching (anaphase). Scale bar, 5 μm. n = 6. (D) Schematic showing cell cycle stages in C. albicans with the distance of separation of sister kinetochores (green circles). Correlation of inter-kinetochore distance with the cell cycle stages revealed an increase in the recovery of CENPA (GFP) from anaphase (>4-μm inter-kinetochore distance) and maximum recovery during anaphase/telophase (5- to 9-μm inter-kinetochore distance). These stages overlapped with the timing of Scm3 localization during late anaphase/telophase. (E) The relative enrichment of H4 at CEN7 normalized to LEU2 (a far-CEN locus) when SBC189 (URA3-TETp-CDC15/cdc15Δ::dpl200) (Bates 2018) was grown in presence of nocodazole (metaphase-arrested) and doxycycline (post-anaphase-arrested) shows a higher H4 occupancy at late anaphase. Two-way ANOVA; (***) P-value < 0.001; (ns) P-value > 0.05; n = 3. (F) A model to explain CENPA loading at late anaphase/telophase and centromere stabilization by the constitutive localization of Orc4 in C. albicans. During centromere DNA replication, CENPA molecules are probably partitioned to the replicated chromatids leaving gaps. Until then, centromeric chromatin is protected by localization of Orc4 at S phase, G2/metaphase, and anaphase. Scm3 localizes to the kinetochore toward the end of mitosis, loading new CENPA molecules by late anaphase/telophase, and remains associated with the kinetochore at G1, after which it is undetectable. Although experimentally proved localization of proteins are shown in solid colors, when the centromeric localization of these proteins is speculative, they are shown as a transparent haze.
As a CENPA chaperone, Scm3 localizes to kinetochores only during late anaphase through the next G1 phase of the cell cycle in C. albicans. This is different compared with the constitutive localization of Scm3 at all stages of the cell cycle in S. cerevisiae (Wisniewski et al. 2014), which shows a replication-coupled mode of CENPA loading. Thus, we wanted to examine the precise loading time of CENPA in C. albicans by performing extensive microscopic examination in YJB8675 (CSE4-GFP-CSE4/CSE4) (Joglekar et al. 2008). For this purpose, we categorized cell cycle stages in C. albicans based on budding indices and inter-kinetochore distances (Supplemental Fig. S7A,B). Upon examining a population of unbudded (G1), small budded (S), large budded (metaphase and anaphase/telophase) cells, we observed an increase in fluorescence intensity of CENPA (GFP) in late anaphase/telophase cells (Supplemental Fig. S7C). This increase was significant in comparison to signals obtained from metaphase cells, indicating a loading regimen of CENPA during late anaphase/telophase. The increase in fluorescence intensity can also be explained by the fluorescence maturation of GFP during anaphase, as has been reported previously (Wisniewski et al. 2014). However, earlier studies have suggested the deposition of new kinetochore subunits during anaphase in budding yeasts (Shivaraju et al. 2012; Dhatchinamoorthy et al. 2017). Hence, we performed fluorescence recovery after photobleaching (FRAP) experiments to measure the loading time of CENPA. To examine if there is an S phase loading of CENPA in C. albicans, we first performed photobleaching in G1 (unbudded) cells and could not recover any fluorescence until metaphase (Fig. 7B). Subsequently, photobleaching of the CENPA cluster during metaphase led to an increase in fluorescence intensity as cells progressed to late anaphase (mean recovery = 110%), revealing a late anaphase/telophase loading pathway (Fig. 7C; Supplemental Table S3). This also confirmed the increased fluorescence intensity of anaphase cells, consistent with our previous observation (Supplemental Fig. S7C). The inter-kinetochore distance in late anaphase cells that recovered from bleaching was found to be 5.4–8.6 μm, coinciding with late anaphase/telophase stage (Fig.7D). Hence, Scm3 localization and CENPA FRAP assays reveal that CENPA loading in C. albicans occurs during late anaphase/telophase. Subsequently, we performed ChIP-qPCR using anti-H4 antibodies in metaphase and late anaphase cells (Fig.7E). We observed an increase in the relative H4 enrichment at centromeres (with respect to a noncentromeric control locus, LEU2) at late anaphase compared with metaphase. This result indicates the assembly of new CENPA nucleosomes at late anaphase. Taken together, these results suggest that although centromere DNA replicates early in S phase, new CENPA molecules load at centromeres in late anaphase/telophase of the cell cycle in C. albicans.
Discussion
In the present study, we provide evidence that genome-wide replication time zones and spatial distribution of Orc4 play a critical role in the loading of CENPA and its maintenance at epigenetically regulated centromeres in C. albicans. We examined the genome-wide binding of Orc4 across the C. albicans genome and observed that, apart from many discrete genomic loci, Orc4 was strongly enriched at centromeres. Orc4 occupancy at all eight centromeres overlapped with that of CENPA. A thorough in silico analysis identified that the four distinct DNA modes are associated with more than 400 Orc4-bound sites that are spatiotemporally distributed across the genome. Our polymer model simulation revealed that the early replicating, highly enriched Orc4-bound regions are positioned toward the centromeres, and the mid and late replicating regions are positioned toward the telomeres. This organization of Orc4 facilitates centromere clustering and, along with Mcm2, is essential for CENPA stability. The constitutive centromeric localization of Orc4 was found to be independent of CENPA or Mcm2. To decipher the loading pathway of CENPA, we characterized Scm3, the CENPA chaperone in C. albicans, which was found to be localized to the kinetochore during anaphase/telophase through G1, coinciding with the loading time of new CENPA molecules. Taken together, we propose Orc4 to be a genome organizer in C. albicans, facilitating centromeric chromatin assembly and its maintenance with the help of Scm3 and Mcm2. Our replication timing analysis also helps to explain the presence of an Orc4 concentration gradient, which we propose to be responsible for regulating centromere assembly and function in C. albicans.
Previous attempts to identify replication origins in fungi have relied on examining the genome-wide occupancy of pre-RC subunits. A genome-wide study on the identification of ORC-bound regions in C. albicans (Tsai et al. 2014) used antibodies against the S. cerevisiae ORC complex to report approximately 390 sites, 27% (113/414) of which overlapped with our study. Because we used antibodies against an endogenous protein (CaOrc4) to map its binding sites in C. albicans, we possess a more authentic depiction of genome-wide Orc4 occupancy. A subset of 50 Orc4-bound regions identified in our study was located within the tDNA. tDNAs show a conserved replication timing (Müller and Nieduszynski 2017) and are known to contain binding sites for TFIIIC, TFIIIB, RNA pol III, and the SMC subunits (Glynn et al. 2004; Kogut et al. 2009). tDNAs also cluster near centromeres and recover stalled forks (Thompson et al. 2003). This feature was consistent with the clustering frequency of mode-A-containing Orc4 binding sites in our study, indicative of a higher-order chromatin structure imposed by Orc4. Additionally, the multiple Orc4 binding modes identified in our study, unlike the conserved motifs associated with replication origins in S. cerevisiae (Wyrick et al. 2001), hint toward a differential mode of origin recognition in this organism as has been reported in Pichia pastoris (Liachko et al. 2014).
In eukaryotes, early firing origins are more efficient and are organized into initiation zones (Mesner et al. 2013). We show that the same feature is conserved in C. albicans as well, wherein orcE-orcE and orcL-orcL regions cluster significantly more than orcE-orcL regions. The orcE regions had higher enrichment of Orc4 than orcL regions. One fact that could limit the resolution of our analysis is that the anti-Orc4 antibodies might primarily detect the early regions owing to a higher enrichment of Orc4 at these sites, hence overrepresenting the “early” data set. Similar to previous studies (Mesner et al. 2011, 2013), early firing, but not late firing, origins appear to have been sequenced to saturation. Our analysis also reveals a higher contact probability between orcL-orcL regions than orcE-orcE regions, which could be explained by the fewer number of orcL regions obtained from the tertile distribution. The orcE regions form closely associated units and interact sparsely with orcL, reminiscent of the genome-wide replication landscape in Candida glabrata (Descorps-Declère et al. 2015). Hence, one can speculate the existence of topologically distinct domains of chromosomes that are separated in space and time as the S phase progresses.
Polymer models for chromosomes previously generated using Hi-C contact maps assume an inverse relationship between contact probability and the average distance between the bead-pairs/two loci (Rousseau et al. 2011). However, the present simulation method enables us to predict the average spatial 3D distance between any two regions in the genome using a contact map as the input. Our simulations produce an ensemble of steady-state genome configurations (corresponding to a population of cells) by which a higher-order organization and other statistical properties of the genome can be computed. The spatiotemporal distribution of Orc4 not only contributes to the compartmentalization of replication domains but also facilitates centromere clustering and kinetochore stability. This enables a more holistic understanding of ORC-origin recognition mediated by either DNA, chromatin, conformation, or the more recently explored interactions by multiprotein complexes that phase separate in solution (Strom et al. 2017), a possibility that we wish to explore in the future. Recent evidence of the nonuniform localization of ORCs in flies has been attributed to phase separation owing to their intrinsically disordered regions (IDRs) (Parker et al. 2019). Even though IDRs are absent in ORC subunits of yeast species (Parker et al. 2019), the replication time–driven spatial distribution of Orc4 identified in our study can explain conserved processes like replication origin communication and coordination of origin firing time.
The strong centromeric enrichment of Orc4 and its overlapping binding pattern with CENPA were of particular interest, primarily because of the lack of a consensus DNA sequence at centromeres in C. albicans (Sanyal et al. 2004). Whereas CEN2, CEN3, and CEN7 harbor mode B, CEN5 and CENR harbor mode C, and CEN1, CEN4, and CEN6 harbor mode D, none of the centromeres have mode A. The lack of consensus reinstates a DNA sequence–independent recognition of centromeres by Orc4 that facilitates centromere clustering. Additionally, the association of Orc4 with the neocentromere locus suggests its role in centromere establishment. Hi-C analysis has shown that the neocentromere locus gets included in the centromeric cluster upon its activation (Burrack et al. 2016). Therefore, Orc4, being a constitutive part of the centromeric cluster, also occupies the neocentromeric locus. This is also supported by the high enrichment of Orc4 at early replicating regions. We currently lack direct experimental evidence to show the mechanism by which Orc4 regulates centromere clustering. However, CENPA declustering is well evident upon Orc4 depletion. CENPA is dislodged from the kinetochore and is degraded by the proteasomal machinery whenever kinetochore integrity is compromised in C. albicans (Thakur and Sanyal 2012). Therefore, CENPA reduction and declustering of kinetochores are coupled, making it difficult to look for a mutant that affects clustering alone. To address this issue, we plan to examine the specific domain of Orc4 involved in centromere clustering and perform further Hi-C assays in such a conditional orc4 mutant. It is to be noted here that the gradual drop in CENPA levels upon Orc4 depletion cannot be solely explained by the G2/M arrest observed in the orc4 mutant. ChIP-qPCR assays may not be the most sensitive means to delineate this difference.
The centromeric localization of Orc4 is independent of CENPA or Mcm2, supporting the constitutive centromeric localization of Orc4. This also means that in the event of eviction of CENPA nucleosomes (naturally or owing to a genomic insult), Orc4 protects the centromere integrity. It is more likely that Orc4 is directly bound to centromeric DNA than indirectly recruited by a chromatin-mediated interaction because Orc4 binds to centromere DNA even in the absence of CENPA. Orc2 has been shown to localize to centromeres in human cells (Prasanth et al. 2004), and the role of ORC in heterochromatin organization is well documented (Prasanth et al. 2010). It will be useful to determine if Orc4 in C. albicans acts independently of the rest of the ORC subunits in stabilizing centromeric chromatin, more so because C. albicans lacks conventional heterochromatin machinery.
The temporal sequestration of centromeric chromatin replication relative to bulk chromatin is thought to prevent misincorporation of H3 nucleosomes onto centromere DNA and thereby facilitate efficient incorporation of CENPA nucleosomes. Our results reveal a late anaphase loading pathway for CENPA in which the integrity of kinetochores is maintained with the help of Orc4 and Mcm2. The role of Orc4 is important here because both the replication timing and spatial distribution of Orc4-bound regions not only orchestrate the whole-genome organization but also mediate localized protection of centromere integrity. Our study also reveals that Orc4 loads and stabilizes CENPA in a capacity independent of its canonical function of replication initiation, because CENPA loading occurs much after centromere replication (Fig. 7F). Upon centromere replication during early S phase, CENPA is distributed into the replicated DNA strands, presumably leaving nucleosome-free regions (gaps) that need to be protected until the late anaphase/telophase loading of CENPA by Scm3. In humans and flies, placeholder molecules like H3.3 occupy centromeric nucleosomes until new CENPA is loaded (Dunleavy et al. 2011; Ray-Gallet et al. 2011). Our observations instead hint toward nucleosome-free gaps from early S phase until late anaphase, possibly protected by Orc4. The affinity of ORC toward nucleosome-depleted regions (Lipford and Bell 2001) might help in explaining our model. An increase in the nucleosome content at centromeres at late anaphase compared with metaphase with a concomitant increase of CENPA levels suggests a gap-filling mode of CENPA incorporation in this organism. Our ChIP-seq analysis fails to delineate the precise centromeric occupancy of both Orc4 and CENPA at the base-pair level, the resolution of which will reveal whether Orc4 and CENPA are parallelly or alternately arranged at centromeres.
At present, we have insufficient evidence to pinpoint the exact role of Mcm2 in CENPA loading. Hence the dynamics of Mcm2 at centromeres is speculative. Although the nuclear localization of Mcm2 during late anaphase might suggest an interaction with Scm3, this hypothesis is plagued by the caveat that MCMs in S. cerevisiae display temporal regulation in their subnuclear localization (Yan et al. 1993), where only a fraction of the nuclear MCMs associate with DNA. If the mechanism of Mcm2 shuttling is similar in C. albicans, our placement of Mcm2 in the model for CENPA stability may be simplistic and highly speculative at present. Hence, examining the cell cycle–specific localization of Mcm2 at centromeres will reveal its association with them. In addition, we have insufficient evidence to address whether Mcm2 acts singly or as part of the Mcm2-7 complex. In humans, HJURP copurifies with the Mcm2-7 complex and simultaneously interacts with CENPA (Zasadzińska et al. 2018). Although Scm3 interaction with Mcm2 remains to be studied in C. albicans, the acquisition of the novel module of three C2H2 motifs in Scm3 of C. albicans might suggest a species-specific CENPA loading pathway.
Methods
The chemical reagents, strains, plasmids, and primers used in this study have been provided in Supplemental Tables S4, S5, S6, and S7. Details of strain construction, additional experiments and analyses are mentioned in the Supplemental Information Materials and Methods. Software and algorithms used for analysis have been listed in Supplemental Table S8.
Media and growth conditions
The orc4 and mcm2 mutants were grown either in permissive (CM-methionine-cysteine) or in nonpermissive (CM + 5 mM methionine + 5 mM cysteine) conditions of the MET3 promoter for the indicated time. The scm3 mutants were grown in presence of 1 mM methionine + 1mM cysteine for repression. CAKS3b (Sanyal and Carbon 2002) was grown in YP with succinate (2%) for expressing CENPA and in YP with dextrose (2%) for depleting CENPA for 8 h. SC5314 was grown in YPDU. The cdc15 mutant SBC189 (Bates 2018) was grown in CM and repressed in presence of 20 μg/mL doxycycline for 16 h. To arrest cells in the S phase, YJB8675 (Joglekar et al. 2008) cells were grown in presence of 200 mM HU for 2 h. To arrest cells in metaphase, YJB8675, SBC189, and CaLS342 cells were grown in presence of 20 μg/mL nocodazole for 4 h. For recycling SAT1 marker, colonies were grown in YP with 2% maltose and checked for nourseothricin sensitivity. The S. cerevisiae strain JBY254 (Wisniewski et al. 2014) was grown in YPDU. All cultures were grown at 30°C.
Generation of anti-Orc4 antibodies
The peptide sequence from C. albicans Orc4 (YLPKRKIDKEESSI) was chemically synthesized and conjugated with keyhole limpet hemocyanin. The conjugated peptide (1 mg/mL) was mixed with equal volumes of Freund's complete adjuvant and used as an antigen to inject nonimmunized rabbits as the priming dose. Three subsequent booster doses at an interval of 2 wk (per immunization) were given using Freund's incomplete adjuvant. Following antibody detection using ELISA, a major bleed was performed. The antiserum was collected, IgG-fractionated, and affinity-purified against the free peptide (AbGenex). The specificity of the purified antibody preparation was confirmed by western blot and immunolocalization experiments.
Chromatin immunoprecipitation
For the Orc4 ChIP-sequencing experiment, ∼500 O.D. of asynchronously grown log phase culture of C. albicans SC5314 cells was cross-linked for 1 h using formaldehyde added to a final concentration of 1%. The quenched cells were incubated in a reducing environment in the presence of 9.5 mL of distilled water and 0.5 mL of 2-mercaptoethanol. The protocol for ChIP was followed as described previously (Yadav et al. 2018b). Briefly, the sheared chromatin was split into two fractions, one of which was incubated with purified anti-Orc4 antibodies (5 μg/mL of IP). Following overnight incubation, the IP and mock (no antibody) fractions were further incubated with Protein A–Sepharose beads. The de-cross-linked chromatin fraction was purified and ethanol-precipitated. The DNA pellet was finally resuspended in 20 μL of Milli-Q water. All three samples (I, +, −) were subjected to PCR reactions. For the CENPA ChIP, cells were cross-linked for 15 min with 1% formaldehyde, and IP samples were incubated with 4 μg/mL of anti–Protein A antibodies or 3 μg/mL of anti-GFP antibodies. For Mcm2 ChIP, cells were cross-linked for 1 h with 1% formaldehyde, and IP samples were incubated with 4 μg/mL of anti–Protein A antibodies. For H4 ChIP, cells were cross-linked for 15 min with 1% formaldehyde, and IP samples were incubated with anti-H4 antibodies (Abcam ab10158). The rest of the protocol was the same as described above.
ChIP-seq analysis
Immunoprecipitated DNA and the corresponding DNA from whole-cell extracts were quantified using Qubit before proceeding with library preparation. Around 5 ng ChIP and total DNA were used to prepare sequencing libraries using a NEB next ultra DNA library preparation kit for Illumina (NEB). The library quality and quantity were checked using Qubit HS DNA (Thermo Fisher Scientific) and Bioanalyzer DNA high sensitivity kits (Agilent Technologies), respectively. The QC passed libraries were sequenced on Illumina HiSeq 2500 (Illumina). HiSeq rapid cluster and SBS kits v2 were used to generate 50-bp single-end reads. The reads were aligned onto the C. albicans SC5314 reference genome (v. 21) using Bowtie 2 aligner (v. 2.3.2) (Langmead et al. 2009). More than 95% of the reads mapped onto the reference genome (control: 97.74%; IP: 96.13%). The alignment files (BAM) were processed to remove PCR duplicate reads using the MarkDuplicates module of Picard tools. For peak calling, MACS2 (Feng et al. 2012) was run in the default mode (narrow peaks) with paired-end mode switched on with an effective genome size set as 14,324,316, and other parameters were set to default (default q-value/FDR is ≤0.05). The command used to identify narrow peaks was as follows: “callpeak -c SC-1CHIPcontrol_rmdup.bam -t SC-1CHIPtest_rmdup.bam -f BAMPE -g 14324316 -n Narrowpeak.” To detect peaks that span a broader range, MACS2 was run in the broad mode, with paired-end mode switched on, effective genome size set to 14,324,316, and other parameters set to default (default q-value/FDR is ≤0.1). The command used to identify broad peaks was as follows: “callpeak -c SC-1CHIPcontrol_rmdup.bam -t SC-1CHIPtest_rmdup.bam -f BAMPE -g 14324316 -n Broadpeak ‐‐broad.” These peaks were annotated with the C. albicans SC5314 reference annotation file. Visualization of the BAM files on the reference genome was performed using Integrative Genomics Viewer (IGV) (Robinson et al. 2011; https://software.broadinstitute.org/software/igv/).
ChIP-qPCR analysis
All ChIP-qPCR experiments were performed with three technical replicates for each of the biological triplicates (n). The input and IP DNA were diluted appropriately, and qPCR reactions were set up using specific primers. ChIP-qPCR enrichment was determined by the percentage (%) input method. In brief, the Ct values for input DNA were corrected for the dilution factor (adjusted value = Ct input or Ct IP − log2 of dilution factor) and then the percentage of the input chromatin immunoprecipitated by the antibodies was calculated using the formula: 100 × 2(adjusted Ct input − adjusted Ct IP) (Mukhopadhyay et al. 2008). One-way ANOVA, two-way ANOVA, and Bonferroni posttests were performed to determine statistical significance, wherever applicable. For all the Orc4 ChIP-PCR assays, % IP values at centromeres were either compared with the control region, LEU2, or normalized with % IP values at a centromere unlinked (far-CEN) Orc4 binding region. Relative enrichment of H4 was plotted as a ratio of % IP values of H4 at CEN and LEU2.
Fluorescence recovery after photobleaching
Photobleaching experiments were performed using a Zeiss LSM880 microscope. The overnight culture of YJB8675 was transferred to fresh media and grown until log phase. Cells were washed with PBS and sandwiched between CM with 2% agarose on glass glide and a coverslip. The entire cluster of GFP in a metaphase cell was bleached using 75% laser power of 488-nm laser for 75 iterations with a pinhole of 1 Airy unit. Images were acquired before and after bleaching with Z-stacks (eight slices., 0.5-μm step-size) collected after every 10 min until recovery of the fluorescence. Similar procedures were performed for unbudded cells. The GFP fluorescence intensities were plotted by measuring the pixel values of the signals corrected for the background levels; stacked projection images were processed and quantified using ImageJ. The percentage of recovery of photobleaching was calculated using the formula reported previously (Dhatchinamoorthy et al. 2017): % recovery = (maximum intensity at recovery − postbleaching intensity) × 100 / (prebleaching intensity − postbleaching intensity).
Polymer modeling of chromosomes
The paired-end reads from the Hi-C data (Burrack et al. 2016) were mapped onto the wild-type C. albicans genome assembly 21 following the HiCUP pipeline with default parameters (Wingett et al. 2015). Next, the resulting BAM file was analyzed using the DryHiC R package (Vidal et al. 2018), and ICE normalization (Imakaev et al. 2012) was applied. The contact matrix was converted to a data frame object and written to a file for subsequent analysis. To compute
the 3D organization of the C. albicans genome, chromatin was modeled as a polymer with N beads connected by (N − 1) harmonic springs. We coarse-grained chromatin
into equal-sized beads, each representing 10 kb of the genome and the connecting springs with a natural length l0 (Lieberman-Aiden et al. 2009). To model the haploid yeast genome of C. albicans comprising of eight chromosomes of different lengths, we considered eight polymer chains each consisting of 319, 224, 180,
161, 120, 104, 95, and 229 beads, respectively. The bead corresponding to the midpoint of centromeres of each chromosome was
assigned as the centromere (CEN) bead. In each chain, to represent the connectivity, all neighboring beads were connected
linearly by a harmonic spring having energy (Ganai et al. 2014):
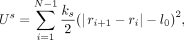 where Us is the spring potential energy, ks is the spring stiffness, ri is the position vector of the ith bead, and l0 is the natural length. The summation here is between nearest neighbors. To mimic the steric hindrance between any two parts
of chromatin, the repulsive part of the Lennard-Jones (LJ) potential energy, given below, was used:
where Us is the spring potential energy, ks is the spring stiffness, ri is the position vector of the ith bead, and l0 is the natural length. The summation here is between nearest neighbors. To mimic the steric hindrance between any two parts
of chromatin, the repulsive part of the Lennard-Jones (LJ) potential energy, given below, was used:
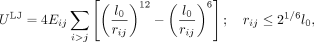 where rij represents the distance between bead i and bead j, Eij represents the strength of attraction, and the sum is over all possible bead-pairs. Hi-C data at a 10-kb resolution was considered
as an input in the current model. We generated an initial configuration by connecting each pair of beads (i, j) with probability Pij as per the Hi-C contact matrix. We chose a uniformly distributed random number r in the interval [0, 1], and the bond was introduced if Pij > r, for each pair of beads. This bond is also a harmonic spring with high stiffness kc and of natural length l0 having energy
where rij represents the distance between bead i and bead j, Eij represents the strength of attraction, and the sum is over all possible bead-pairs. Hi-C data at a 10-kb resolution was considered
as an input in the current model. We generated an initial configuration by connecting each pair of beads (i, j) with probability Pij as per the Hi-C contact matrix. We chose a uniformly distributed random number r in the interval [0, 1], and the bond was introduced if Pij > r, for each pair of beads. This bond is also a harmonic spring with high stiffness kc and of natural length l0 having energy
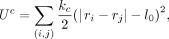 where (i, j) represents summation over the pairs selected probabilistically as described above. All the chromosomes are confined into
a sphere of radius Rs, which represents the confinement arising owing to the nucleus. One of the centromeres (CEN1) was tethered to the nuclear periphery. The resulting polymer was equilibrated via Langevin simulation using LAMMPS (Plimpton 1995). The whole process described above was repeated for 1000 realizations, generating an ensemble of 1000 configurations. Each
of the configurations is equivalent to the chromatin in a single cell.
where (i, j) represents summation over the pairs selected probabilistically as described above. All the chromosomes are confined into
a sphere of radius Rs, which represents the confinement arising owing to the nucleus. One of the centromeres (CEN1) was tethered to the nuclear periphery. The resulting polymer was equilibrated via Langevin simulation using LAMMPS (Plimpton 1995). The whole process described above was repeated for 1000 realizations, generating an ensemble of 1000 configurations. Each
of the configurations is equivalent to the chromatin in a single cell.
Data access
The ChIP-sequencing data generated in this study have been submitted to the NCBI BioProject database (https://www.ncbi.nlm.nih.gov/bioproject/) under accession number PRJNA477284.
Competing interest statement
The authors declare no competing interests.
Acknowledgments
We thank Clevergene Biocorp for generating the Orc4 ChIP-sequencing data. We also thank Dr. R.G. Prakash for the animal facility and B. Suma for confocal microscopy, JNCASR. We thank S. Bates (University of Exeter), C. Wu (Johns Hopkins University), J. Berman (Tel Aviv University), and S. Mitra (University of Oxford) for providing us with yeast strains. We thank A. Koren and J. Berman for sharing the raw data of the replication timing experiment. K.S. acknowledges funding from Tata Innovation Fellowship from the Department of Biotechnology (DBT), Government of India (BT/HRT/35/01/03/2017); J.C. Bose fellowship from Science and Engineering Research Board (SERB-JCB/JNCASR/D0017/2020/00807), Government of India; a grant from DBT (BT/PR14557/BRB/10/1529/2016); and a DBT grant in Life Science Research, Education and Training at JNCASR (BT/INF/22/SP27679/2018), as well as intramural funding from JNCASR. R.S. thanks the PRISM-II project at IMSc and funding by the Department of Atomic Energy (DAE), and R.P. acknowledges support from the Science and Engineering Research Board (SERB) and Department of Science and Technology (DST), India, grant EMR/2016/005965. L.N. is supported by DBT grant BT/PR16240/BID/7/575/2016. L.S. thanks support from the Council of Scientific and Industrial Research (CSIR), Government of India (09/733(0178)/2012-EMR-I), and JNCASR (JNC/AO/PB.022(L)). A.B. is supported by DBT grant BT/PR14840/BRB/10/880/2010. N.V. is supported by CSIR fellowships 09/733 (0253)/219-EMR-I and 9/733 (0161)/2011-EMR-I. K.G. acknowledges CSIR S.P. Mukherjee fellowship SPM-07/733(0181)/2013-EMR-I.
Footnotes
-
[Supplemental material is available for this article.]
-
Article published online before print. Article, supplemental material, and publication date are at https://www.genome.org/cgi/doi/10.1101/gr.265900.120.
- Received May 12, 2020.
- Accepted January 27, 2021.
This article is distributed exclusively by Cold Spring Harbor Laboratory Press for the first six months after the full-issue publication date (see https://genome.cshlp.org/site/misc/terms.xhtml). After six months, it is available under a Creative Commons License (Attribution-NonCommercial 4.0 International), as described at http://creativecommons.org/licenses/by-nc/4.0/.