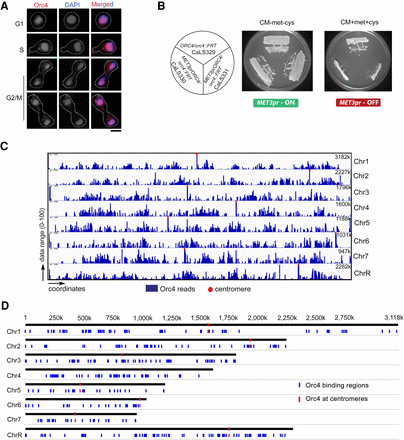
Orc4, an essential subunit of the origin recognition complex, is nuclear-localized and binds to discrete loci in the C. albicans genome. (A) Nuclear localization of Orc4 in C. albicans SC5314 (ORC4/ORC4) cells as evidenced by staining with anti-Orc4 antibodies and DAPI. Scale bar, 5 µm. (B) The promoter of MET3 in C. albicans, expressed in the absence of methionine and cysteine and repressed in the presence of both amino acids, was used for the conditional expression of ORC4. CaLS329 (ORC4/orc4::FRT) with one deleted copy of ORC4 and two independent transformants, CaLS330 and CaLS331 (MET3prORC4/orc4::FRT), where the remaining wild-type copy was placed under the control of the MET3 promoter, were streaked on plates containing permissive (CM-met-cys) or nonpermissive (CM + 5 mM met + 5 mM cys) media and photographed after 48 h of incubation at 30°C. (C) ChIP-sequencing analysis revealed that Orc4 was bound to discrete genomic sites in C. albicans. The total Orc4 reads (blue histogram) were obtained by subtracting the relative number of sequencing reads from the whole-cell lysate from the Orc4 ChIP sequence reads and aligning them to the reference genome C. albicans SC5314 assembly 21. Red dots indicate centromeres. (D) Orc4 binding regions (blue) on each of the eight C. albicans chromosomes including all eight centromeres (red).











