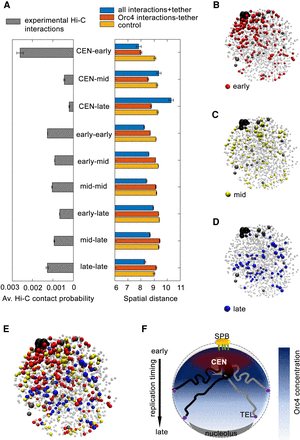
Early replicating Orc4-bound regions are spatially proximal to the clustered centromeres. (A, left) The average contact probability for the indicated region for each of the timing classes (across and within orcE, orcM, and orcL domains) showed stronger CEN-orcE interactions. (Right) The average spatial distances between the indicated regions calculated from the Langevin simulation for each of the timing classes (across and within the orcE, orcM, and orcL domains) when all Hi-C interactions (blue), only the Orc4-Orc4 interactions (red), and control (yellow) simulation for a random configuration are performed. (B–E) Snapshot of the 3D configuration of the C. albicans genome (one out of 1000 realizations) from the simulations shows all chromosomes in light gray, centromeres in black, and telomeres in dark gray. The orcE regions are shown in red (B), orcM in yellow (C), and orcL in blue (D); E represents all the Orc4 binding sites (orcE, orcM, and orcL). (F) Schematic of a budding yeast nucleus showing the typical Rabl configuration in which the clustered centromeres are anchored near SPBs and telomeres are often away from the centromere cluster and occasionally interacting with the nuclear envelope. In C. albicans, the highest spatial enrichment of Orc4 is near the centromere cluster. Orc4 concentration gradually diminishes toward the opposite pole. Concomitantly, early replicating regions are located toward centromeres and the late regions are toward telomeres. (CEN) Centromeres; (TEL) telomeres.











