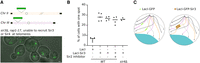Sir3 mediates long-range chromosome interactions in budding yeast
- Myriam Ruault1,5,
- Vittore F. Scolari1,2,5,
- Luciana Lazar-Stefanita2,3,6,
- Antoine Hocher1,7,
- Isabelle Loïodice1,
- Romain Koszul2,4 and
- Angela Taddei1,4
- 1Institut Curie, PSL University, Sorbonne Université, CNRS, Nuclear Dynamics, 75005 Paris, France;
- 2Institut Pasteur, Unité Régulation Spatiale des Génomes, CNRS, UMR 3525, C3BI USR 3756, F-75015 Paris, France;
- 3Sorbonne Université, collège Doctoral, F-75005 Paris, France;
- 4Cogitamus Laboratory, F-75005 Paris, France
-
↵5 These authors contributed equally to this work.
Abstract
Physical contacts between distant loci contribute to regulate genome function. However, the molecular mechanisms responsible for settling and maintaining such interactions remain poorly understood. Here, we investigate the well-conserved interactions between heterochromatin loci. In budding yeast, the 32 telomeres cluster in 3–5 foci in exponentially growing cells. This clustering is functionally linked to the formation of heterochromatin in subtelomeric regions through the recruitment of the silencing SIR complex composed of Sir2/3/4. Combining microscopy and Hi-C on strains expressing different alleles of SIR3, we show that the binding of Sir3 directly promotes long-range contacts between distant regions, including the rDNA, telomeres, and internal Sir3-bound sites. Furthermore, we unveil a new property of Sir3 in promoting rDNA compaction. Finally, using a synthetic approach, we demonstrate that Sir3 can bond loci belonging to different chromosomes together, when targeted to these loci, independently of its interaction with its known partners (Rap1, Sir4), Sir2 activity, or chromosome context. Altogether, these data suggest that Sir3 acts as a molecular bridge that stabilizes long-range interactions.
The 3D organization of a genome in space and time can potentially impact its functions. However, the mechanisms underlying this organization and its dynamics remain poorly understood. Contacts between distant loci can, for instance, influence and/or regulate biological functions, as exemplified by the enhancer–promoter interactions observed in metazoa. Another example observed from yeast to man is provided by the clustering of repeated sequences sequestering silencing factors and thus forming repressive subcompartments (Meister and Taddei 2013).
The formation of these subcompartments in the nuclear space has several functional consequences. First, it allows for a more robust and specific repression of the associated sequences. Subcompartments allow a local concentration of silencing factors and, consecutively, sequester these factors from the outside, limiting their action elsewhere in the genome. Second, the spatial proximity of the clustered sequences favors recombination events between them (Batté et al. 2017). Finally, because these sequences can be located on different chromosomes or separated by long distances along the same chromosome, subcompartments affect the global genome folding in 3D space. Therefore, understanding the molecular mechanisms driving and regulating these long-range interactions provides insights on several DNA mechanisms, including genetic regulation, the maintenance of chromosomal stability, or also compaction and folding throughout the cell cycle. Chromatin tethering or colocalization involves a variety of direct or indirect molecular mechanisms, such as cohesin-mediated loop formation (Nasmyth 2005; Rao et al. 2014) or anchoring of heterochromatin to the nuclear lamina (Falk et al. 2019). However, protein-mediated direct bridging of distant loci positioned on the same or different chromosomes remains scantily described experimentally.
In exponentially growing cells, the 32 telomeres of Saccharomyces cerevisiae cluster within 3 to 5 foci mainly found at the nuclear periphery, where the silencing information regulators Sir2, Sir3, and Sir4, forming the SIR complex, concentrate (Gotta et al. 1996). These clusters represent a well-documented example of a repressive nuclear subcompartment (Kueng et al. 2013; Gartenberg and Smith 2016). This organization is regulated by the physiological state of the cells, as illustrated in long-lived quiescent cells, in which telomeres regroup into a unique, large cluster in the center of the nucleus (Guidi et al. 2015).
At the molecular level, the SIR complex is recruited at the telomeric and subtelomeric TG1-3 repeats by the binding of the transcription factor Rap1, whose C-terminal end contains binding domains for the silencing factors Sir3 and Sir4 (Gartenberg and Smith 2016). The SIR complex is also recruited at the silent mating type loci (HM loci) through different DNA-binding proteins, such as Orc1 and Abf1, which—like Rap1—also have other functions in the cell (Haber 2012).
Furthermore, genome-wide ChIP experiments revealed that Sir3 can also be found at the promoters of some subtelomeric seripauperin genes (Radman-Livaja et al. 2011) and at a handful of discrete and telomere-distal loci not associated with specific functions (so far) (Sperling and Grunstein 2009; Radman-Livaja et al. 2011; Takahashi et al. 2011; Teytelman et al. 2013; Mitsumori et al. 2016; Hocher et al. 2018).
In addition, Sir2 is found at the rDNA locus, where it protects rDNA repeats from recombination. Although Sir3 was initially thought to associate to the nucleolus only in aged cells (Kennedy et al. 1997; Sinclair 1997), it was later found to interact with the rDNA by chromatin immunoprecipitation (ChIP) in exponentially growing cells (Radman-Livaja et al. 2011).
Once recruited, the SIR complex has the ability to spread along the chromatin fiber and to repress the transcription of underlying genes by the RNA polymerase II. This spreading results from the association of the histone deacetylase activity of Sir2 with the affinity of Sir3 for unmodified nucleosomes, linked by Sir4 interactions with both Sir2 and Sir3. The N-terminal BAH (bromo-adjacent homology) domain of Sir3 plays a major role in binding nucleosomes through an extensive surface of interaction, which is stabilized by the constitutive N-terminal acetylation of Sir3 (Armache et al. 2011; Arnaudo et al. 2013). According to these structural studies, acetylation of H4K16 or methylation of H3K79 would reduce nucleosome binding affinity for Sir3. Reciprocally, H4K16 deacetylation by Sir2 increases the affinity of the Sir2/3/4 complex, favoring its spreading on adjacent nucleosomes (Oppikofer et al. 2011). In vivo, SIR spreading is limited by the titration of SIR proteins, especially Sir3, and by H3K79me3, which prevents Sir3 binding in euchromatic regions. Indeed, overexpression of Sir3 leads to the formation of extended silent domains (ESD) covering subtelomeric regions and delimited by transition zones where H3K79me3 levels abruptly increase (Hocher et al. 2018).
A consequence of the limiting amount of SIR proteins is that loci associated with these proteins compete for these limiting pools (Buck and Shore 1995; Smith et al. 1998; Michel 2005). Telomere clustering, by increasing the local concentration within these subcompartments, could thus favor SIR spreading within subtelomeric regions. Reciprocally, telomere clustering as well as long-range interactions between HM loci depend on SIR recruitment at these loci (Gotta et al. 1996; Miele et al. 2009). Furthermore, Sir3 overexpression leads to increased telomere clustering in addition to increased SIR spreading (Ruault et al. 2011). A Sir3 point mutation (A2Q) enables disentanglement of telomere clustering and SIR spreading. This mutation prevents the N-terminal acetylation of Sir3 and thus its ability to bind nucleosomes and spread along subtelomeric regions, yet Sir3-A2Q promotes telomere clustering (Sampath et al. 2009; Ruault et al. 2011; Hocher et al. 2018). This led us to propose that Sir3 is the bridging factor between telomeres (Ruault et al. 2011; Meister and Taddei 2013).
Here, we directly test this hypothesis by exploring the role of Sir3 on the global organization of the genome by combining genome-wide capture of chromosome conformation (Hi-C) and microscopy.
Results
Sir3 impacts genome organization
We previously showed that Sir3 is a limiting factor for telomere clustering. Indeed, whereas the telomere-bound protein Rap1 fused to GFP in living cells forms 3–5 foci at the nuclear periphery in wild-type cells, only weak residual foci can be detected in sir3Δ cells (see Fig. 1A; Ruault et al. 2011). These weak foci possibly result from the random encounter of telomeres at the nuclear periphery (Zimmer and Fabre 2011; Hozé et al. 2013), due to Sir3-independent telomere anchoring at the nuclear envelope (Tham et al. 2001; Hediger et al. 2002; Schober et al. 2009). On the contrary, Sir3 overexpression leads to the grouping of telomeres in larger foci, or hyperclusters, located in the center of the nucleus (Ruault et al. 2011). Moreover, overexpression of the Sir3-A2Q mutant, whose N-terminal substitution blocks its acetylation, thus preventing silencing (Wang et al. 2004) and spreading in subtelomeric regions (Hocher et al. 2018), also resulted in the formation of telomere hyperclusters (Fig. 1A; Ruault et al. 2011). Whereas telomere clusters tend to dissociate in wild type during mitosis (Fig. 1A), hyperclusters mediated by Sir3 overexpression persist throughout the cell cycle (right panel of Fig. 1A).
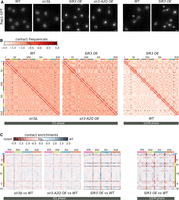
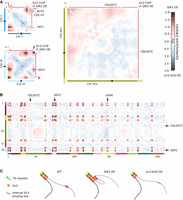
Sir3 impacts genome organization. (A) Representative fluorescent images of the telomere-associated protein Rap1 tagged with GFP in exponentially growing strains expressing different Sir3 levels: wild type (yAT2583), no Sir3 (yAT2584), high Sir3 levels (yAT2476), or high Sir3-A2Q levels (yAT2822). The two panels on the far right: representative mitotic cells (M) in the wild-type strain or in the strain overexpressing Sir3. (B) Normalized chromosome contact maps (bin: 50 kb) of cells expressing various levels of Sir3 synchronized either in G1 or G2/M. (C) Ratio plots of pairs of contact maps of four representative chromosomes. Blue: enrichment of contacts in WT; red: enrichment in mutants.
To explore the influence of Sir3 on the 3D organization of the genome, we generated genome-wide Hi-C contact maps (Lieberman-Aiden et al. 2009) of cells either depleted for Sir3 or carrying different sir3 alleles (Methods). To prevent unwanted influence(s) of the loop organization of mitotic chromosomes characterized in cycling cells (Garcia-Luis et al. 2019; Dauban et al. 2020), Hi-C libraries were obtained from G1 elutriated daughter cells (Diamond 1991). The contact maps of wild-type cells revealed an enrichment of contacts between telomeres as well as between centromeres (Fig. 1B, red dots on the map), as previously reported (Duan et al. 2010; Cournac et al. 2012). The enrichment in telomere–telomere contacts almost disappears in the absence of Sir3 (sir3Δ background) (black arrowheads), whereas centromere–centromere contacts remain apparent (black * on the map). In the strain overexpressing Sir3, inter-subtelomeres contact frequency was strongly increased (Fig. 1B) at the expense of contacts with other parts of the genome (white lines in between red dots), whereas frequency of inter-centromeres contacts appeared unchanged compared to a WT strain (Fig. 1B, black * on the map; Supplemental Fig. S1A). These differences can be visualized more quantitatively by computing the log2-ratio plot between mutants and WT maps (Fig. 1C). In these maps, the color scale reflects the balance of contacts between the two conditions: the bluer, the more contacts in the WT; the redder, the more contacts in the mutant. The comparison of WT and sir3Δ maps results in the visualization of inter-subtelomeres contact ratio as blue pixels dots, as expected from the drop in inter-subtelomeres contact frequency observed in the sir3Δ strain compared to WT. Conversely, the red stripes expanding from and bridging subtelomeric regions indicate that the declustered telomeres are now able to contact more other regions of the genome in the absence of Sir3. This reflects the fact that telomeres are now less constrained in the absence of clustering (see also Muller et al. 2018 for a similar effect during meiosis prophase).
In strains overexpressing the silencing-deficient Sir3-A2Q protein, the frequency of inter-telomere contacts also increased compared to WT, in agreement with the microscopy, although not to the extent of the strains overexpressing Sir3 (red dots in Fig. 1B). The ratio map of the Sir3-A2Q-overexpressing cells versus WT cells shows an inverted pattern compared to the sir3Δ versus WT, with red dots at the level of the telomeres and blue lines in between telomeres (Fig. 1C). A similar but stronger pattern was observed when comparing WT and Sir3-overexpressing strains in G1. This pattern was even more contrasted for G2/M cells, as telomere contacts are barely detectable in the wild-type strain whereas they remained high in number upon Sir3 overexpression (Fig. 1B,C right panels; Supplemental Fig. S1B). Altogether, these data show that contacts between subtelomeres decrease in the absence of Sir3, whereas they increase upon Sir3 or Sir3-A2Q overexpression, in good agreement with microscopy.
Sir3 binding coincides with inter-subtelomere contacts
To determine whether Sir3 overexpression has an impact on global chromosome compaction, we plotted the contact probability curve, p(s). As expected, the curves display an enrichment in contacts for distances ∼10 kb to 100 kb in G2/M compared to G1 samples (Fig. 2A), reflecting the condensation of chromosomes during mitosis (Lazar-Stefanita et al. 2017; Schalbetter et al. 2017). However, different amounts of Sir3 did not significantly modify the compaction of the genome measured at the global level in G1 phase, as assessed by p(s) curves (Fig. 2A).
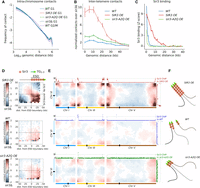
Sir3 spreading coincides with telomere–telomere contacts. (A) Contact probability as a function of genomic distance Pc(s) (log scale) for different strains either in G1 or G2. (B) Accumulated normalized contacts over 40-kb subtelomeric windows (0 = X-core elements). (C) Sir3 enrichment over 40-kb subtelomeric windows (data from Hocher et al. 2018). (D) Pile-up contact maps ratio for various pairs of strains of subtelomeric windows centered on the extended silent domains (ESD; dotted line) determined in the SIR3 overexpression (OE) condition. The windows extend over 50 kb in both directions from the ESD. (E) Representative inter-chromosomal ratio maps of the same pairs of strains as in D processed through Serpentine binning. Sir3 enrichment observed by ChIP (Hocher et al. 2018) in the indicated conditions is plotted along the top and right axis. (F) Schemas illustrating the behavior of telomeres and subtelomeric regions in the SIR3 OE, WT, and sir3-A2Q OE conditions.
We then focused on telomere–telomere contacts. We first compared the frequency of inter-telomere contacts as a function of the distance to telomeres, with the frequency of inter-centromere contacts as a function of the distance to centromeres (Supplemental Fig. S2A). The frequency of contacts shows a very similar decay when the distances from telomeres or centromeres increase in wild-type cells. This probably reflects a similar flexibility of chromosome arms in these regions. In Sir3- or Sir3-A2Q-overexpressing cells, we observed a higher frequency of inter-telomere contacts, whereas inter-centromere contacts remain unchanged. However, assessing subtelomeric interactions in the telomere proximal regions was limited by the high variability of the alignment coverage in subtelomeric regions. We thus plotted the ratio of inter-subtelomere contacts between a given strain and a sir3Δ strain as a function of the distance to the telomeres to conceal these sequence biases (Fig. 2B). The ratio of the signal in the sir3Δ strain measures the effects of the Sir3 protein on the contact probability and concomitantly conceals the Sir3-independent biases present in all strains. For both the wild-type or the Sir3-A2Q-overexpressing strains, ratios were the highest near the telomeres and decreased slowly when moving away from it. In good agreement with our microscopy data, inter-subtelomere contacts were higher in the Sir3-A2Q-overexpressing strain than in the wild type. Sir3 overexpression led to an even higher ratio of inter-subtelomere contacts that decay slowly with the distance to telomere over the first 10 kb before decreasing sharply up to 35 kb toward the centromeres. The extent of high level of inter-subtelomere contacts is comparable to the extent of Sir3 spreading as analyzed by ChIP (Fig. 2C; Hocher et al. 2018), suggesting that Sir3-bound regions mediate these contacts.
Indeed, in wild-type cells, Sir3 binds to DNA 2.6 kb away, on average, from the last telomeric element. In strains overexpressing Sir3, it can be detected as far as 20 kb away from the telomere, generating large Sir3-bound regions, dubbed “extended silent domains” or ESDs (Fig. 2C; Hocher et al. 2018). In contrast, Sir3-A2Q detection is limited to the TG repeats, even when overexpressed.
To assess the contribution of Sir3 binding to trans-subtelomere contacts, we computed the cumulated amount of trans contacts between subtelomeric loci as a function of their distance from their respective ESD (Methods). We observed a general increase of trans-subtelomere interactions for all loci located between the telomeres and the ESD boundaries, which we identify as the signature of the telomere hypercluster. Past the ESD border, trans interactions decrease sharply, further suggesting that in Sir3-overexpressed conditions, Sir3-bound regions (ESDs) define a specific compartment isolated from the rest of the chromosomes. We also notice a persistence of trans contacts along the diagonal reflecting a co-alignment of subtelomeric regions adjacent to the ESDs (seen as the increased contact frequency between regions at equivalent genomic distances from the ESD boundary). These regions are possibly maintained in close proximity by the constraint imposed by the telomere hypercluster to the chromatin polymeric backbone. This co-alignment of chromosome arms is also visible in the cumulated inter-chromosome contact maps of the 100-kb subtelomeric regions and is very similar to the one observed in pericentromeric regions in G2/M phase (Supplemental Fig. S1B; Schalbetter et al. 2017). In wild-type cells or Sir3-A2Q-overexpressing cells, we observed a progressive decrease of contacts from the telomere proximal region to the telomere distal region, probably reflecting the bonding of subtelomeres through shorter regions upstream of the ESD boundary (Fig. 2D,E; Supplemental Fig. S2A). Also, in WT cells, the intensity of the contacts is generally lower, reflecting the presence of more than one telomere cluster, possibly due to heterogeneity at the single cell level as a result of random encounters of different couples of telomeres (Fig. 1A).
At the individual chromosome level, to visualize the correlation between Sir3 spreading and contacts between different subtelomeres, we built differential contact maps at a higher level of detail using the Serpentine algorithm (Baudry et al. 2020), which adapts the resolution on each position of the contact map to the data coverage (Fig. 2E; see Supplemental Fig. S2B for maps comprising all chromosomes). The Serpentine map comparing the Sir3-overexpressing cells with sir3Δ showed a strong increase of inter-subtelomere contacts between ESDs of Chromosome VI and Chromosomes I, III, and V (dark red regions at the corners of the maps, Fig. 2E). As a consequence, these subtelomeric regions make less contact with non-subtelomeric regions. In the wild-type versus sir3Δ Serpentine maps, we observed a similar, though less pronounced, trend.
Thus, contacts between subtelomeric regions coincide with Sir3 binding, suggesting that Sir3 binding, which is the epigenetic information setting on the linear chromosomes, also defines the boundary of the telomere clusters in 3D (Fig. 2F).
Sir3 but not Sir3-A2Q accumulates at the rDNA upon overexpression
We next asked whether non-subtelomeric Sir3-bound regions interact with subtelomeres. Sir3 binds to the rDNA through a mechanism that requires Sir2 deacetylase activity (Hoppe et al. 2002; Radman-Livaja et al. 2011). In agreement with this work, live microscopy showed that a functional endogenously expressed yeGFP-tagged version of Sir3 colocalizes with an endogenously expressed TagRFP-T-tagged rDNA-bound protein Net1 (Fig. 3A; Supplemental Fig. S3A–C).
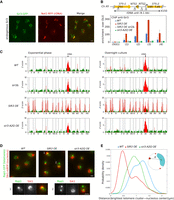
Sir3 but not the silencing-deficient mutant, Sir3-A2Q, associates with the rDNA and promotes rDNA-telomere contacts. (A) Representative fluorescent images of a double-tagged Sir3-yeGFP/Net1-TagRFP-T strain (yAT2803). Cells were grown in CSM 2% glucose and imaged in exponential phase. (B) The S. cerevisiae rDNA locus is composed of ∼150 tandem copies of a 9.1-kb repeating unit, each encoding two transcribed region RDN5 and RDNA37 (comprising RDN18, RDN5.8, and RDNA25 genes). The graph represents Sir3 occupancy along the rDNA locus probed by ChIP-qPCR using an anti-Sir3 antibody (Ruault et al. 2011) in WT (yAT232), GAL1p-SIR3 (yAT208), GAL1p-sir3-A2Q (yAT1205), and GAL1p-SIR3 sir2Δ (yAT772) strains. Primer pair 1 amplifies a region of the RDN25 locus, primer pair 2 amplifies a region in NTS1 (Non-Transcribed Spacer 1) region, primer pair 3 amplifies a region in NTS2, and primer 4 amplifies a region in the ETS1 (External Transcribed Spacer 1) region (see Supplemental Table S2 for primer sequences). Strains were grown in YPGal for 48 h. The bar graph represents the Sir3 enrichment over the mitochondrial locus OLI1. Error bars show the SEM of three independent experiments, each analyzed in triplicate qPCRs. (C) 4C extraction profiles representing contact maps between the rDNA repeats and the rest of the genome; profiles are shown for all strains in both exponential and overnight cultures. Red-highlighted loci correspond to bins with z-value greater than 2.5. (D) Representative fluorescent images of a double-tagged strain Rap1-GFP/Sik1-mRFP in strains expressing an endogenous level of Sir3 (yAT340), high levels of Sir3 (yAT341), or high levels of the separation-of-function mutant Sir3-A2Q (yAT1198) after an overnight culture in CSM 2% galactose. Magnification of representative nuclei (1 and 2) are presented at the bottom of the panel. (E) Distance between the brightest Rap1-GFP cluster and the nucleolus center is plotted for a wild type (yAT340, n = 581), a strain overexpressing Sir3 (yAT341, n = 627), and a strain overexpressing Sir3-A2Q (yAT1198, n = 590) using the NucLoc software (Berger et al. 2008). Cells were grown in CSM 2% galactose overnight before imaging. Scale bar: 1 µm in all panels.
To identify precisely the region bound by Sir3 in a rDNA repeat unit, we performed ChIP experiments (Methods). Sir3 was found preferentially enriched in the nontranscribed regions (NTS1 and NTS2) and, to a lesser extent, around the RDN25 transcriptional unit and the promoter of the 37s rDNA precursor transcript (Fig. 3B). Probing Sir3 binding in a strain overexpressing Sir3, we observed a strong accumulation of Sir3 over the rDNA unit, with a stronger signal at NTS1 and NTS2, showing that Sir3 concentration was limiting for binding not only at telomeres but also at the rDNA locus. Consistent with previous work (Hoppe et al. 2002), we did not observe Sir3 binding in the absence of Sir2, even upon Sir3 overexpression. In contrast to wild-type Sir3, the Sir3-A2Q mutant that has lost its affinity to bind nucleosomes was not detected at the rDNA by ChIP. Together, these results suggest that Sir3 accumulates at the rDNA through its affinity for unacetylated nucleosomes resulting from Sir2 activity (Fig. 3B).
Sir3 promotes the interaction of telomeres with the rDNA locus
A 4C-like analysis using the rDNA locus as a viewpoint (Methods) showed that when Sir3 is expressed at its endogenous level in exponential phase, the rDNA locus makes significant long-range contacts with only two loci, located outside Chromosome XII (Fig. 3C). When the rDNA locus was less active (overnight cultures), we observed that long-range contacts between the rDNA and other regions of the genome increased to 11, corresponding mostly to rDNA–telomere interactions (10 out of 11). In contrast to the contacts observed in exponential phase, those were lost in the absence of Sir3 (Fig. 3C). These observations suggest that the activity of the rDNA prevents these Sir3-dependent contacts during exponential phase. In strains overexpressing Sir3, the rDNA locus now shows during exponential phase significant contacts with all telomeres, indicating that the increased amount of Sir3 at both types of loci favored their trans contacts. When Sir3-A2Q was overexpressed, the loci in contacts with the rDNA remained similar to WT exponentially growing cells, but significant contacts with telomeres were not detected after an overnight culture. Therefore, rDNA-telomere contacts require Sir3 binding to the rDNA. Note that these analyses are averaged over a cell population and do not necessarily reflect the contacts of the rDNA in each independent cell of the population.
To observe these contacts at the single cell level in strains expressing different sir3 alleles, we imaged cells expressing Rap1-GFP (telomere foci) and Sik1-mRFP (involved in pre-rRNA maturation and thus staining the nucleolus). Images were taken from overnight cultured cells enriched in rDNA–telomere interactions. In a wild-type strain, telomere foci are located at the nuclear periphery and do not interact with the nucleolus (Fig. 3D). In contrast, telomeres in a strain overexpressing Sir3 group together and position in close contact with the nucleolus. More specifically, the telomere hypercluster is set within the nucleolus, excluding the Sik1 protein (magnification shown at the bottom of Fig. 3D). The Sir3-A2Q-mediated telomere hypercluster, on the other hand, did not interact with the nucleolus, in agreement with Hi-C maps. We next measured the 3D-distance between the brightest telomere clusters and the nucleolus at a single cell level using NucLoc (Berger et al. 2008). The distance between the brightest cluster of telomeres and the center of mass of the nucleolus was shorter in strains overexpressing Sir3 (with a median distance of 0.6 µm) compared to the wild-type strain (1.2 µm) (Fig. 3E). In strains overexpressing the Sir3-A2Q mutant, the distance was intermediate (median distance: 0.9 µm), reflecting the localization of the hypercluster at the center of the nucleus without interacting with the nucleolus, again in agreement with 4C maps.
Also in agreement with 4C plots, the physiological state of the cells influenced the level of association between the telomere hypercluster and the nucleolus. Indeed, these two subnuclear compartments that are tightly associated after an overnight culture in Sir3-overexpressing cells dissociate upon dilution in fresh medium and re-associate when cells reach saturation (Supplemental Fig. S3D,E).
The number of rDNA repeats also influenced telomere hypercluster-rDNA contacts: fewer contacts were measured in cells carrying a short rDNA (25 copies) compared with cells with a high copy number (190 repeats) in the Sir3-overexpressing mutant (Supplemental Fig. S3F).
These results show that contacts between the rDNA and telomeres are modulated by (1) the amount of available Sir3, and (2) the transcriptional activity at the rDNA, which prevents these interactions.
Sir3 overexpression compacts the rDNA and inverts the spatial organization of the nucleolus
Because Sir3 has a strong impact on the spatial organization of the telomeres, we asked whether the accumulation of Sir3 at the rDNA locus could modify its spatial organization. In wild-type cells, the rDNA labeled by Net1-GFP appears as a filament (Fig. 4A,B), as previously reported (Straight et al. 1999). In Sir3-overexpressing cells, rDNA compaction varies with the physiological state of the nucleolus (Supplemental Fig. S4A), with an increasing compaction from early to late exponential phase. As expected from cells grown overnight, the compacted rDNA is overlapping with the telomere hypercluster in most cells (Fig. 4A, bottom panel). In contrast, cells overexpressing the Sir3-A2Q mutant, that cannot bind the rDNA, show a filamentous rDNA (Fig. 4A), suggesting a direct role for Sir3 in compacting the rDNA.
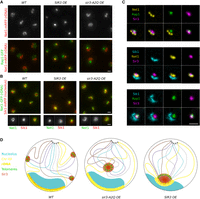
Sir3 overexpression impacts the rDNA spatial organization and its compaction. (A) Representative fluorescent images of Rap1-GFP/Net1-mRFP strains expressing either the endogenous level of Sir3 (yAT3729), high levels of Sir3 (yAT3730), or high levels of the Sir3-A2Q mutant (yAT3733). Cells were grown in CSM with 2% galactose and imaged after an overnight culture. (B) Representative fluorescent images of a double-tagged strain Net1-GFP/Sik1-mRFP in strains expressing the endogenous level of Sir3 (yAT1004), high levels of Sir3 (yAT1008), or high levels of the Sir3-A2Q mutant (yAT1541). Cells were grown in CSM 2% galactose and imaged after an overnight culture. In the bottom panels, close-up of selected nuclei from the field are shown (GFP-channel, RFP channel, and merge). (C) Representative fluorescent deconvolved images of triple-tagged strains, individual and merge channels are shown. Cells were grown overnight in CSM 2% glucose and are all expressing high Sir3 levels (Sir3 under the control of the constitutive GPD promoter). From top to bottom: Net1-BFP2 Rap1-GFP Sir3-mCherry (yAT3901), Sik1-BFP2 Net1-GFP sir3-mCherry (yAT2213), and Sik1-BFP2 Rap1-GFP Sir3-mCherry (yAT3666). (D) Schema representing the nuclear organization of a wild type, a strain overexpressing the Sir3-A2Q separation-of-function mutant, and a strain overexpressing the wild-type Sir3. Scale bar: 1 µm in all panels.
We next investigated the consequence of overexpressing Sir3 on nucleolus organization by imaging strains expressing the rDNA-bound protein Net1 labeled with GFP, as well as Sik1-mRFP, staining the nucleolus. The GFP-labeled rDNA filament surrounds the Sik1 signal in wild-type cells or in cells overexpressing Sir3-A2Q (Fig. 4B). This organization was inverted in Sir3-overexpressing cells, with the Net1-GFP signal located in the center of the nucleolus and surrounded by the Sik1 signal. The compaction of the rDNA as well as the inversion of the nucleolar organization were both independent of the formation of telomere hyperclusters, as they were observed in sir4Δ cells overexpressing Sir3 (Supplemental Fig. S4B,C).
Finally, to get a better idea of the arrangement of the telomere hypercluster and the nucleolus when they interact, that is, when the activity of the nucleolus is low (overnight cultures), we made triple-labeled strains that allowed us to image three-way combination of the telomere hypercluster (Rap1), Sir3, the rDNA (Net1), and the nucleolus (Sik1), in Sir3-overexpressing strains (Fig. 4C). In those conditions, the Sir3 signal from the telomeres was indistinguishable from the rDNA-bound one, corresponding to a large focus that overlapped with both the Net1 and Rap1 signals (Fig. 4C, upper panel; Supplemental Fig. S4D), with no overlap between these two signals. Furthermore, the Sir3 signal overlapping with Net1 was always facing the Sik1 signal (i.e., the nucleolus), whereas the Sir3 signal colocalizing with Rap1 was found at the opposite side of the nucleolus (Fig. 4C, lower panel; Supplemental Fig. S4D). Therefore, the subtelomeres (enriched in Sir3 and deprived of Rap1) are in contact with the Sir3-bound rDNA, whereas the extremity of the telomeres (enriched with Rap1) protrudes toward the nuclear interior.
These results led us to propose the model presented in Figure 4D: in strains overexpressing Sir3, Sir3 binds and coalesces the telomeres into a hypercluster, while accumulating at the rDNA as well and increasing its compaction. When the activity of the nucleolus is low, the regions of the rDNA that are Sir3-bound interact with the telomere hypercluster, bringing them together into a “super” Sir-enriched subcompartment. The part of the nucleolus dedicated to pre-RNA processing is excluded from this subcompartment and is self-organizing between the Sir3-enriched subcompartment and the nuclear periphery.
Ectopic Sir3-bound regions coincide with long-range and trans contacts
In addition to the telomeres and the rDNA, small peaks of Sir3 binding are detected by ChIP at a few discrete sites located internally on chromosomal arms (Sperling and Grunstein 2009; Takahashi et al. 2011; Mitsumori et al. 2016; Hocher et al. 2018). Although some of these sites are known ChIP artifacts (Teytelman et al. 2013), others are not, as no enrichment was seen in the Sir3-A2Q-expressing strain, and Sir3 enrichment increased upon Sir3 overexpression (Hocher et al. 2018).
The Serpentine maps revealed that some of those internal sites are also in contact with other Sir3-bound loci, especially in cells that have slowed down cell division (i.e., overnight cultures). This was clearly visible on the Serpentine map of Chromosome I and VI when comparing cells overexpressing Sir3 to the ones overexpressing the Sir3-A2Q mutant protein that is not recruited at these internal sites (Fig. 5A). Especially, the YAT1 and IGD1 genes, respectively located on Chromosome I and VI (38 kb and 60 kb from the closest telomere), showed increased contact frequencies with the subtelomeres belonging to the same chromosome. This was also true for the YDL007C-A ORF located 13 kb away from the centromere of Chromosome IV, which interacts with the subtelomere IVL, thus counteracting the global Rabl-like organization of this chromosomal arm (Fig. 5A). The Serpentine map indeed also revealed increased cis interactions along the left arm of Chromosome IV, possibly reflecting a loop bridging YDL007C-A and the telomere in some of the cells. Further, we observed increased contact frequencies between the SIR3 gene, where Sir3 is recruited upon overexpression, and the subtelomeres located 56 kb downstream on the right arm of Chromosome XII (Supplemental Fig. S5A), possibly hinting at a negative feedback mechanism regulating SIR3 expression. This mechanism would be different from one previously proposed (Renauld et al. 1993), where Sir3 spreading extends up to SIR3, as we do not observe such a continuous spreading of Sir3 from the telomeres up to the SIR3 locus (Supplemental Fig. S5A).
Sir3-bound regions coincide with long-range contacts. (A) Intra-chromosomal ratio plots of Chromosome I, VI, and IV-L contact maps generated in sir3-A2Q OE and SIR3 OE conditions and processed by Serpentine binning. The ChIP deposition profile of Sir3 in OE conditions are plotted along the top and right axis. Intra-chromosomal loci enriched in Sir3 are indicated by the closest gene name. (B) Serpentine binning plots recapitulating the enrichment in contacts in SIR3 OE conditions with respect to sir3-A2Q OE conditions, showing Chromosomes I, IV, V, and VI interacting with the rest of the genome. (C) Schematic representation of the behavior of internal Sir3 binding sites in different conditions.
However, not all Sir3-bound sites showed preferential interactions with telomeres, maybe owing to other constraints associated with cis-associated loci. Some of the internal Sir3-bound sites, that interact with the subtelomeres of their own chromosome in cis, also interact in trans with other subtelomeres. Figure 5B presents the contacts between Chromosomes IV, V, and VI and the rest of the genome and emphasizes the ability of internal Sir3-bound sites to contact any subtelomere of the genome. No other trans contacts outside Sir3-bound regions were observed in these maps (Supplemental Fig. S5B). However, Hi-C maps of the sir3Δ strain show apparent contacts between the FLO1, FLO9, and FLO11 genes located, respectively, in telomere-proximal regions of Chromosome I's right arm, left arm, and IX's right arm (Supplemental Fig. S5C). Further analysis at the read level revealed that the apparent interaction between FLO1 and FLO9 could stem from an alignment artifact. However, this is not the case for FLO1–FLO11 interaction (Supplemental Fig. S5D). In strains expressing Sir3 or Sir3A2Q, this interaction probably also occurs but is almost impossible to distinguish from Sir3-mediated interactions between subtelomeres.
In summary, we observed that upon Sir3 overexpression, increased Sir3 binding at internal sites smaller than 4 kb is accompanied by increased contacts between these sites and subtelomeric regions, suggesting a direct involvement of Sir3 (Fig. 5A–C). We do not observe increased interactions between these sites and subtelomeres when comparing the wild-type and sir3Δ strains. However, we detect an increased compaction of these loci resulting in a higher frequency of contact between these sites and their flanking regions (Supplemental Fig. S5E). It is possible that Sir3 retains a role of segregation and silencing on these sites without affecting the large-scale chromosome architecture in the tested wild-type conditions. Alternatively, increased bridging between these loci and telomeres might be present in the cell population but is not significant enough to be detectable by our assay.
Artificial arrays of Sir3 are sufficient to promote trans interactions
To directly test whether Sir3 is sufficient to bond two ectopic loci together, independently of silencing and clustering of telomeres, we hijacked the GFP-LacI/LacO array system to target Sir3 to specific loci harboring LacO arrays by expressing GFP-LacI fused to Sir3 (Fig. 6A). The fusion of GFP-LacI in the N terminus of Sir3 abolishes its silencing function, but its clustering function remains fully functional (Supplemental Fig. S6A,B). LacO arrays were introduced at LYS2 (Chr II) and LEU2 (Chr III) loci in a strain unable to form telomere foci (rap1-17 sir3Δ) to prevent the recruitment of GFP-LacI-Sir3 to telomeres. We measured the pairing of the two arrays in different strains after an overnight culture. In a strain expressing GFP-LacI, the two foci were paired in 6% of the cells (Fig. 6B). When we expressed GFP-LacI-Sir3, the percentage of paired loci increased to 28%. This increased pairing was independent of Sir2 activity, as 25% of the cells had a paired array when grown in the presence of the splitomicin Sir2 inhibitor. The Sir3-induced increased pairing was also independent of Sir4, because the percentage of pairing was similar in a WT and sir4Δ strain expressing GFP-LacI-Sir3. Because these experiments were performed in strains expressing a rap1 allele truncated for the Rap1–Sir3 interaction domain, we can infer that the Sir3-mediated pairing is also Rap1-independent. Therefore, this synthetic approach demonstrated that Sir3 can bond loci belonging to different chromosomes together, independently of its interaction with Rap1, and Sir4 and Sir2 activity, or chromosome context (Fig. 6C). This strongly suggests that Sir3–Sir3 interactions provide the physical link that holds together Sir3-bound loci.
Arrays of Sir3 are sufficient to promote trans interactions. (A) Schema describing the strains used in the assay. sir3Δ rap1-17 strains carrying a lac operator (lacO) array at the LYS2 locus on Chromosome II and a lacO array at the LEU2 locus on Chromosome III are expressing either GFP-LacI or GFP-LacI-Sir3. Merge image of a transmitted-light image and the GFP channel fluorescent image of a strain expressing the GFP-LacI-Sir3 construct (yAT1470). (B) Graphic showing the percentage of cells with one spot in different strains and conditions: GFP-LacI (yAT1476), GFP-LacI-Sir3 (yAT1470) in the presence or in the absence of splitomycin (a Sir2 inhibitor) and GFP-LacI-Sir3 sir4Δ (yAT1864). Cells were grown in CSM 2% glucose to mid-exponential phase of growth before being imaged. (C) Schema representing the nuclear organization of strains expressing either the GFP-LacI (left) or the GFP-LacI-Sir3 (right).
Discussion
Here, we investigated the impact of the silencing factor Sir3 on the regulation of the yeast genome 3D folding. We show that Sir3 is both necessary and limiting for telomere clustering.
Sir3 mediates interactions between telomeres
Hi-C contact maps clearly show that interactions between subtelomeric regions decrease in the absence of Sir3 and increase upon Sir3 overexpression. These interactions do not require Sir3 spreading because overexpressing Sir3-A2Q, whose binding is limited to the telomeric TG repeats, is sufficient to increase subtelomeric contacts. However, contacts rapidly decrease away from telomeres in the Sir3-A2Q strain, whereas in the strain overexpressing the WT version of Sir3, they remain strong up to over ∼20 kb along the subtelomeres. In addition, when Sir3 is overexpressed, the increased contacts coincide with Sir3 enrichment along the subtelomeres. This suggests that the (limited) subtelomeric contacts observed in the Sir3-A2Q strain are an indirect consequence of the Sir3-dependent bonding of chromosomes by their telomeres, whereas the spreading of WT Sir3 directly promotes the subtelomeric contacts. Extended Sir3 spreading is also associated with increased interaction in cis possibly reflecting the compaction resulting from heterochromatin formation.
Sir3 impacts nucleolar organization
Another consequence of overexpressing Sir3 is its impact on the organization of the nucleolus. Although Sir3 association with the rDNA has been documented (Hoppe et al. 2002; Radman-Livaja et al. 2011), its function at this locus remained elusive. Our experiments show that increasing Sir3 association with the rDNA increases its compaction, especially when the rDNA activity is low (i.e., in slow growing cells). This compaction is accompanied by an inversion of the nucleolar organization: whereas the rDNA is found at the periphery of the nucleolus in wild-type cells, it becomes nested within the nucleolus upon Sir3 overexpression (Fig. 4). In addition, in these conditions both Hi-C and microscopy show that the rDNA associates with telomeres in a Sir3-dependent manner. Although we could not observe this association by microscopy in wild-type cells, such contacts were nevertheless detected in the present Hi-C data for 10 telomeres (out of 32) in wild-type cells with low rDNA activity. These observations show that Sir3 influences the overall genome organization not only at the level of subtelomeric regions but also more broadly.
It is noteworthy that Sir3 recruitment at the rDNA was first reported as a hallmark of aged cells (Kennedy et al. 1997; Sinclair 1997), possibly owing to a lack of sensitivity to detect Sir3 in the nucleolus of young cells. Sir3 was especially visible in the fragmented nucleoli resulting from the instability of the rDNA in old cells. It will be important to test whether this is related to the ability of Sir3 to compact the rDNA and whether this new property of Sir3 has impacts on the stability of this locus.
Tethering Sir3 is sufficient to bridge chromatin loci
In addition to the trans contacts between large Sir3-covered regions, such as subtelomeres, and the rDNA, we also observed contacts between Sir3-bound subtelomeric domains and at least five discrete, few-kb-long regions enriched in Sir3 and positioned internally on chromosomal arms (Sperling and Grunstein 2009; Takahashi et al. 2011; Mitsumori et al. 2016; Hocher et al. 2018). Although weaker than contacts between subtelomeres, they can counteract the canonical Rabl-like configuration (where telomeres and centromeres are on opposite sides of the nucleus), drawing together the centromere proximal locus of Chromosome IV and its subtelomeres. These internal sites also interact with subtelomeres from other chromosomes in trans (Fig. 5). Albeit these sites do not show specific contacts with telomeres in wild-type cells, we could detect an increased compaction of those loci.
Although the functional significance of these internal Sir3 binding sites remains to be elucidated, they correspond to intergenic regions or genes that are not expressed in our culture conditions. Of note, one example is the YAT1 gene that has the highest G + C content of the budding yeast genome (58%) (Chávez et al. 2001). This is also one of the rare ORFs containing a cluster of meiotic double-strand break hot spots (Pan et al. 2011). Furthermore, this gene is prone to form R-loops when expressed under the control of the strong GAL1 promoter (Bonnet et al. 2017). This could be related to our previous findings that Sir3 is recruited at sites of chromatin stress, favoring their perinuclear anchoring (Dubarry et al. 2011). As for the rDNA, it will be important to test whether Sir3 contributes to the genetic stability of these loci. Finally, we showed that the artificial tethering of Sir3 to two lacO arrays is sufficient to increase their trans association, independently of Sir3 interaction with its known partners (Rap1, and Sir4), Sir2 activity, or chromosome context (Fig. 6).
Dynamics of Sir3-mediated long-range contacts
Telomere clusters are dynamic. They can split and fuse (Schober et al. 2008; Hozé et al. 2013) and dissolve during mitosis to reform in G1 phase. We observed that Sir3 overexpression prevents this dissolution probably by stabilizing telomere–telomere interactions. We noticed that telomere clusters are more visible by microscopy as well as by Hi-C in slow growing cells, possibly due to the time required for telomeres to encounter one another again after mitosis. This is also true for other interactions between Sir3-bound regions, such as the rDNA, or discrete internal or synthetic sites. We propose three nonexclusive explanations for this observation. First, some chromatin movements or constraints linked to genome activity could destabilize or prevent Sir3-mediated contacts. Indeed, rDNA-telomere contacts are limited when the rDNA is transcriptionally active, possibly owing to a stronger anchoring at the nuclear periphery when the rDNA is active. Alternatively, the high density of ongoing transcripts could prevent telomeres from contacting the rDNA through steric hindrance. Finally, long-range contacts may be limited by the time required for Sir3-bound loci to meet in the nuclear space.
Mechanisms of Sir3-mediated trans interactions
Recent work (Gibson et al. 2019) suggests that properties inherent to chromatin, including nucleosomal spacing and histone acetylation status, could promote phase separation phenomena within the nucleoplasm. Reconstituted chromatin undergoes liquid–liquid phase separation (LLPS) in physiologic salt, forming droplets that can be dissolved by acetylating histone tails. Given the presence of the histone deacetylase Sir2 in subtelomeric regions, telomere clustering could have been proposed to result from the LLPS of nonacetylated chromatin. However, we showed that cells expressing the sir3-A2Q allele can form telomere hyperclusters even in the absence of Sir2. Furthermore, the artificial tethering of Sir3 to two distant euchromatic loci is sufficient to bridge them independently of Sir2 activity. These results indicate that the clustering of Sir3-bound loci does not depend on their chromatin status but rather is mediated by direct Sir3–Sir3 interactions. Consistent with this hypothesis, Sir3 carries a very well-characterized dimerization domain on its C-terminal part, the wH domain (Oppikofer et al. 2013). In addition, the Sir3 C-terminal domain can interact with a more internal part of Sir3 (King et al. 2006). The importance of the dimerization properties of Sir3 has been mainly discussed for its contribution to Sir3 spreading in cis and its consequence on chromatin compaction and accessibility (Oppikofer et al. 2013; Swygert et al. 2014, 2018; Behrouzi et al. 2016). However, these specific Sir3–Sir3 interactions could also build up molecular bridges between Sir3-bound regions in trans. Sir3 can bridge mononucleosomes or nucleosomal arrays in vitro (Georgel et al. 2001; McBryant et al. 2008; Behrouzi et al. 2016), and the Sir3 ability to bridge mononucleosomes requires its wH dimerization domain. We thus propose that specific Sir3–Sir3 interactions between arrays of Sir3 molecules associated with subtelomeres, the rDNA, or internal regions would form a dynamic, gel-like network structure rather than a liquid droplet as previously proposed for other types of nuclear foci (Sawyer et al. 2019). Further work is needed to decipher the physical nature of these foci. Although heterochromatin foci in mouse cells were first described as liquid droplets (Strom et al. 2017), a recent study indicates that these foci rather resemble collapsed polymer globules (Erdel et al. 2020). Whether specific heterochromatin factors are responsible for this collapse is not known, but HP1, the functional homolog of Sir3, does not seem to be required (Peters et al. 2001; Mateos-Langerak et al. 2007; Erdel et al. 2020).
Independently of the exact mechanism by which Sir3 promotes long-range interactions, it provides a means to cluster SIR nucleation domains into a subvolume of the nucleus, thus increasing the local concentration of the SIR complex. This will in turn favor the spreading of the complex along the chromatin fiber, thus reinforcing the clustering of these regions. As proposed earlier (Meister and Taddei 2013), this positive feedback will lead to the sequestration of the SIR complex favoring the silencing in these subnuclear compartments while preventing its action elsewhere in the genome.
We show that Sir3 plays an important, direct role in the colocalization of multiple chromatin loci in budding yeast. How these long-range interactions are regulated in response to environmental cues, eventually reshaping chromosome folding to a larger extent and playing new roles in genetic regulation, for instance, has to be further investigated.
Methods
Media and growth conditions
Yeast cells were grown either in rich medium (YPD) or in enriched complete synthetic medium (2× final concentration of CSM, MP Biomedicals) supplemented with 2% glucose or 2% galactose (wt/vol). All the strains were grown at 30°C with shaking at 250 rpm. For galactose induction, cells were grown overnight in YPGal medium (yeast extract, peptone, 2% galactose wt/vol) for preculture and diluted the next day in the same medium for the experiment. For Sir2 inhibition, splitomycin was added directly to the culture at a final concentration of 62.5 µM for 4 h. G2/M synchronization was done as in Lazar-Stefanita et al. (2017). Synchronization at the G2/M transition was achieved by restarting G1 cells in YPD at 30°C for 1 h, followed by the addition of nocodazole (Calbiochem; 15 µg/mL) and incubation for another 2 h at 30°C.
Strains
The strains used in this study are listed in Supplemental Tables S1 and S3. They are all derivatives of W303 (Thomas and Rothstein 1989) except the ones used for the Hi-C experiment (BY4741, Euroscarf). Gene deletions, insertions of alternative promoters, and gene tagging were performed by PCR-based gene targeting.
Plasmids
pAT234 is an integrative plasmid expressing GFP-LacI-Sir3 under the HIS3 promoter. This plasmid is derived from a plasmid expressing GFP-LacI-NLS under the HIS3 promoter (pAFS135) (Straight et al. 1998) and was built in two steps. First, the STOP codon of GFP-LacI-NLS TAA was mutated to AGA by direct mutagenesis using primer pair am449 (5′ AAAGAAGAAGAGAAAGGTTGCCAGATCTAGAGCGGCCGCCACCGCGGTGG 3′) /am450 (5′ CCACCGCGGTGGCGGCCGCTCTAGATCTGGCAACCTTTCTCTTCTTCTTT 3′) yielding pAT233. The full length SIR3 gene was amplified by PCR using primer pair am446 (5′ ATAAGAATGCGGCCGCaATGGCTAAAACATTGAAAG 3′) /am448 (5′ AGTCGAGCTCTCAAATGCAGTCCATATTTTTG 3′). The resulting PCR product was digested with NotI/sacI and cloned into pAT233 digested with NotI/SacI to generate the plasmid encoding the GFP-LacI-Sir3 protein.
Chromatin immunoprecipitation experiments
A total of 20 OD600nm equivalent of cells were fixed in 20 mL with 0.9% formaldehyde for 15 min at 30°C, quenched with 0.125 M glycine for 5 min, and washed twice in cold TBS 1× pH 7.6. Pellets were suspended in 1 mL TBS 1×, centrifuged, and frozen in liquid nitrogen for −80°C storage. All following steps were done at 4°C unless indicated. Pellets were resuspended in 500 µL of lysis buffer (0.01% SDS, 1.1% Triton X-100, 1.2 mM EDTA pH 8, 16.7 mM Tris pH 8, 167 mM NaCl, 0.5% BSA, 0.02 g/L tRNA, and 2.5 µL of protease inhibitor from Sigma-Aldrich P1860) and mechanically lysed using a FastPrep instrument (MP Biomedicals) with 0.5 mm zirconium beads (Biospec Products): intensity 6, 3 cycles of 30 sec with 3 min incubation on ice in between cycles. The chromatin was fragmented to a mean size of 500 bp by sonication in the Bioruptor XL (Diagenode) for 14 min at high power with 30 sec on/30 sec off and centrifuged 5 min at 16,000g. Ten microliters were kept to be used as input DNA. Cleared lysate was incubated overnight with 1 µL of polyclonal antibody anti-Sir3 (Agro-bio) (Ruault et al. 2011). Fifty microliters of magnetic beads protein A (NEB) were added to the mixture and incubated for 4 h at 4°C on a rotating wheel. Magnetic beads were washed sequentially with lysis buffer, twice with RIPA buffer (0.1% SDS, 10 mM Tris pH 7.6, 1 mM EDTA pH 8, 0,1% sodium deoxycholate, and 1% Triton X-100), twice with RIPA buffer supplemented with 300 mM NaCl, twice in LiCl buffer (250 mM LiCl, 0.5% NP-40, 0.5% sodium deoxycholate), with TE 0.2% Triton X-100, and with TE. Inputs were diluted 1/10 with elution buffer (50 mM Tris, 10 mM EDTA pH 8, 1% SDS) and beads were resuspended in 100 µL of elution buffer. A reversal cross-linking was performed by heating samples overnight at 65°C. Proteins were digested with Proteinase K (0.4 mg/mL) in the presence of glycogen, and the remaining DNA was purified on QIAquick PCR purification columns. Finally, samples were treated with 29 µg/mL RNase A for 30 min at 37°C and used for quantitative PCR.
ChIP quantification by quantitative PCR
Quantitative PCR was performed on 1/40 of the immunoprecipitated DNA and 1/3200 of the input DNA. Sequences of interest were amplified using the SYBR Green PCR Master Mix (Applied Biosystems) and the primers listed in Supplemental Table S2. PCR reactions were conducted at 95°C for 10 min followed by 40 cycles at 95°C for 15 sec and 60°C for 30 sec on a real-time quantitative PCR system (Applied Biosystems 7900HT Fast Real-Time PCR). Each real-time PCR reaction was performed in triplicate. The signal from a given region was normalized to the one from the OLI1 control locus in immunoprecipitated and input DNA samples. Plots represent the mean value obtained for at least three independent experiments; error bars correspond to the SEM.
Microscopy
The set of images from any given figure panel was acquired using identical acquisition parameters. For all fluorescent images, the axial (z) step is 200 nm and images shown are a maximum intensity projection of z-stack images, with the exception of Figure 4C and Supplemental Figure S4D that present only a slice of the z-stack. Images were acquired on two different systems, either a wide-field microscopy system or a spinning disk system, both of them being driven by MetaMorph software (Molecular Devices). Images of panel 3D, S3D, and S6B were acquired using a wide-field microscopy system based on an inverted microscope (Nikon TE2000) equipped with a 100×/1.4 NA immersion objective, a charge-coupled device (CCD) camera (Photometrics Coolsnap HQ2). A xenon arc lamp (Sutter Instrument Co. Lambda LS) was used to illuminate the samples. A dual-view micro-imager device, described in Guidi et al. (2015), allows the simultaneous measurement of two-color information on the same sensor. Images of panels 1A, 4A, 4C, S4A, and S4D were acquired with the same microscope, using a C-mos camera and illumination system, Spectra X light engine lamp (Lumencor, Inc). This system allows the fast acquisition of dual-color images when used in combination with a double filter. Images of panels 3A, 4B, 6A, S4B, and S4C were acquired on a spinning-disk confocal microscope (ANDOR Revolution XD Confocal System) equipped with a spinning-disk unit (Yokogawa CSU-X1), a microscope (Nikon Ti 2000) with a 100×/1.4 NA oil immersion objective, and an EM CCD camera (ANDOR iXON DU-885). Images shown in panel 4C and S4D were deconvolved (see below). Images of panel 3A were acquired with an upgraded version of the system allowing the simultaneous acquisition of the GFP and RFP channels using a Tu-Cam module (Andor).
Microscopy data processing
Deconvolution of images acquired with the wide-field microscopy system was made using the Meinel algorithm in MetaMorph (eight iterations; sigma = 0.8; frequency 3; MDS Analytical Technologies).
Hi-C library generation
Hi-C libraries were generated as described in Lazar-Stefanita et al. (2017), using a DpnII four-cutter enzyme. The Hi-C libraries were sequenced on an Illumina NextSeq 500 apparatus (2 × 75 high-throughput kits). The PE reads were filtered and aligned according to the protocol described in Cournac et al. (2012).
Visualization of contact maps
Sparse matrices were binned at 5- or 50-kb pixel size as indicated in figure legends. Normalization, when applied, used the sequential normalization procedure (SCN) (Cournac et al. 2012) and rasterized using a color-scale.
Ratio maps and serpentine binning
The log2 ratio of pairs of raw contact maps, binned at 50 kb, was computed. The resulting matrix was normalized by subtracting the median of its values. The result was rasterized using a color-scale. Serpentine binning was applied for some of the ratio plots. Serpentine is an algorithm developed to overcome the limits of standard binning with respect to read coverage. When using Serpentine, the bin sizes are chosen nonuniformly on the maps as a function of the sequencing coverage (see Baudry et al. 2020 for validation and details on the program). Filtering was applied to remove speckles to avoid artifacts. The Serpentine ratio allows emphasizing contact patterns otherwise drowned in the sampling noise of Hi-C data. Serpentine binning was run independently on contact maps of single or a couple of chromosomes, binned at 1250-bp resolution, with the following parameters: threshold = 70 and minthreshold = 7, as discussed in Baudry et al. (2020). Resulting ratio maps were then normalized by subtracting the mean value of their pixels and rasterized using a color-scale centered in zero.
Contact probability as a function of genomic distance
To compute the intra-chromosomal P(s) plots, pairs of reads aligned in intra-chromosomal positions were partitioned by chromosome arms. Reads oriented toward different directions or separated by <1.5 kb were discarded to discard self-circularizing events. For each chromosome, read pairs were log-binned as a function of their genomic distance s (in kb), according to the following formula: bin = floor(log(s) / log(1.1)). The P(s) plot is the histogram computed on the sum of read pairs for each bin. This sum is weighted by the bin size 1.1(1+bin) (because of the log-binning), as well as the difference between the length of the chromosome and the genomic distance s. The difference acts as a proxy for the number of possible events.
Inter-subtelomere and inter-centromere cumulative contact maps
Inter-subtelomere/centromere contacts where obtained by selecting all the centromere/telomere centered trans interactions in normalized contact maps as a set of small submatrices. A mean interaction profile is then obtained by taking the mean of all these submatrices, reoriented consistently, excluding all the unmapped regions. Subtelomeric regions are aligned and centered around the X-core element.
Inter-subtelomere and inter-centromere contacts as a function of the distance to telomeres
The profile of co-alignment has been obtained by selecting the row/column as the basis of interaction between, respectively, the X-core element or the centromere, and the subtelomere/pericentromere region, in the respective cumulative contact map. The error bar reflects the standard error on the mean, as a consequence of the variability of interaction between each telomere–telomere or centromere–centromere intra couple.
Ratio of the inter-subtelomere contacts as a function of the distance to telomeres
The contacts are obtained by comparing the maps obtained in exponential phase of the wild type, the Sir3 overexpression, and the Sir3-A2Q overexpression strains, to the maps obtained in the sir3Δ mutant. For each telomere we obtained, in tabular form, the interactions between the bin containing the X-core element with the subtelomeres of all the other chromosomes. Chromosome III was excluded from the analysis. The table is then reordered as a function of the distance between subtelomere location and the X-core element of the corresponding telomere. The values obtained the aforementioned way in the considered condition are then divided by the values obtained the same way in the sir3Δ mutant. The mean ratio-table, obtained over all X-core elements and subtelomeres, was then computed, discarding infinite and NaN values. The resulting table represents the inter-subtelomere contacts as a function of the distance from telomeres for the considered condition.
Detection of ESD boundaries
Sir3 binds to telomeres and subtelomeres in the wild-type and overexpressed strains. We obtained the location of ESD boundaries from the z-transformed Sir3 ChIP binding profiles from Hocher et al. (2018). Those profiles show z-values higher than 1 at the extremities of chromosomes. We obtained z-values by binning the ChIP data in 5-kb bins, removing bins without data, then calculating the median and the median-absolute-deviation over all chromosomes. The z-value is then computed by the following formula: log [0.67599 (x − median)/ (MAD)]. For each chromosome, the two ESD boundaries were chosen as the coordinate of the lowest/higher bin in the chromosome with z-values less than 1 in ChIP signal of the overexpressed Sir3 mutant.
Cumulative ratio matrices at the ESD boundary
The cumulative ratio matrices were obtained starting from two raw contact maps binned at 5 kb, here named the biological condition and the control. For each couple of ESD boundaries belonging to different chromosomes, excluding Chromosome III, for each matrix, a submatrix was obtained such that the pixel accounting for the direct ESD–ESD interaction is placed at the center. Each of those submatrices is then reoriented such that the telomere–telomere boundary is placed toward the lower right corner; all interactions beyond the telomere–telomere boundary (such as interaction belonging to adjacent chromosomes) are discarded by setting the values to NaN. The ratio submatrices are obtained by dividing the biological condition over the control and taking its logarithm in base 2. Ratio-submatrices are then normalized by subtracting from all of them the same value, computed by the mean of all the pixels in all the ratio-submatrices, excluding NaNs and infinite values. Finally, the cumulative ratio matrix is the mean of all ratio-submatrices.
4C extraction profiles
From Hi-C matrices, is it possible to extract interactions between a single locus on the genome with the rest of the genome by slicing the full rows of the matrix corresponding to the bin containing the locus of interest. If the locus spans multiple bins, we sliced multiple rows and then reduced the signal to one dimension by taking the sum of the values for each column. In the case of the 4C of the rDNA locus, in order to obtain maximum specificity, the raw data was realigned on a genome with the rDNA removed from the sequence of Chromosome XII and placed on an individual chromosome, stripped of any repeated sequences. The 4C data was then obtained by slicing the full rows corresponding to that individual chromosome and summing the values of each column on the matrix obtained using this special genome as a reference. To detect significant interactions, we plot in green color the z-transform of the 4C signal. The procedure is computed on the logarithm in base 10 of the 4C signal. The median μ and the median absolute deviation (MAD) were computed. The standard deviation was estimated using the following formula: σ = MAD / 0.675. The z-transformed 4C is obtained by the following formula: z = (x − μ)/σ. Interactions are considered significant, and plotted in red color, according to the following condition: z > 2.5.
Data access
The Hi-C library data generated in this study have been submitted to the NCBI BioProject database (https://www.ncbi.nlm.nih.gov/bioproject/) under accession number PRJNA642827.
Competing interest statement
The authors declare no competing interests.
Acknowledgments
The authors thank Pr. Masayasu Nomura for sharing strains, Mickael Garnier for his help on image analysis, Julien Mozziconacci and the members of the Taddei laboratory for helpful discussions, and Susmita Sridhar for her help in editing the manuscript. The A.T. team was financially supported by funding from the Labex DEEP (ANR-11-LABEX-0044 DEEP and ANR-10-IDEX-0001-02 PSL), from the Agence Nationale de la Recherche DNA-Life (ANR-15-CE12-0007), Fondation pour la Recherche Médicale (DEP20151234398), and CNRS grant 80prime PhONeS. The authors also thank the PICT-IBiSA@Pasteur Imaging Facility of the Institut Curie, a member of the France Bioimaging National Infrastructure (ANR-10-INBS-04). V.F.S. is the recipient of a Roux-Cantarini Pasteur fellowship. L.L.-S. was supported in part by a fellowship from the Fondation pour la Recherche Médicale. This research was supported by funding to R.K. from the European Research Council under the Horizon 2020 Program (ERC grant agreement 771813).
Author contributions: M.R. and A.H. generated strains. M.R. performed and analyzed the microscopy experiments. L.L.-S. performed the Hi-C experiments. A.H. performed and analyzed ChIP on chip experiments. I.L. carried out FACs and ChIP-qPCR experiments. V.F.S. conducted the bioinformatics analyses. A.T., R.K., M.R., and A.H. contributed to the design of the experiments. A.T., R.K., V.F.S., and M.R. contributed to the interpretation of the data, the drafting of the figures, and the writing/revision of the manuscript.
Footnotes
-
[Supplemental material is available for this article.]
-
Article published online before print. Article, supplemental material, and publication date are at https://www.genome.org/cgi/doi/10.1101/gr.267872.120.
- Received June 29, 2020.
- Accepted December 30, 2020.
This article is distributed exclusively by Cold Spring Harbor Laboratory Press for the first six months after the full-issue publication date (see https://genome.cshlp.org/site/misc/terms.xhtml). After six months, it is available under a Creative Commons License (Attribution-NonCommercial 4.0 International), as described at http://creativecommons.org/licenses/by-nc/4.0/.