Abstract
The CRISPR/Cas9 system has revolutionized mammalian somatic cell genetics. Genome-wide functional screens using CRISPR/Cas9-mediated knockout or dCas9 fusion-mediated inhibition/activation (CRISPRi/a) are powerful techniques for discovering phenotype-associated gene function. We systematically assessed the DNA sequence features that contribute to single guide RNA (sgRNA) efficiency in CRISPR-based screens. Leveraging the information from multiple designs, we derived a new sequence model for predicting sgRNA efficiency in CRISPR/Cas9 knockout experiments. Our model confirmed known features and suggested new features including a preference for cytosine at the cleavage site. The model was experimentally validated for sgRNA-mediated mutation rate and protein knockout efficiency. Tested on independent data sets, the model achieved significant results in both positive and negative selection conditions and outperformed existing models. We also found that the sequence preference for CRISPRi/a is substantially different from that for CRISPR/Cas9 knockout and propose a new model for predicting sgRNA efficiency in CRISPRi/a experiments. These results facilitate the genome-wide design of improved sgRNA for both knockout and CRISPRi/a studies.
The clustered regularly interspaced short palindromic repeats (CRISPR) system allows efficient perturbation of gene regulation in somatic cells and is increasingly being used for genome-wide functional screens. The CRISPR-based screens fall into two categories: Cas9-mediated knockout and dCas9 fusion-mediated inhibition (CRISPRi) or activation (CRISPRa). In a CRISPR/Cas9 knockout system, the Cas9 nucleases induce double-strand breaks on targeted genomic regions directed by single-guide RNAs (sgRNAs). Indels produced at these breakpoints, as a result of an error-prone DNA repair mechanism, cause loss-of-function gene knockouts at the DNA level (Jinek et al. 2012; Mali et al. 2013; Yang et al. 2013). Alternatively, in the CRISPRi/a systems, catalytically inactive Cas9 (dCas9) proteins guided by sgRNAs enable the localization of effector domains to repress or activate gene transcription without modifying DNA (Gilbert et al. 2013; Qi et al. 2013). CRISPR/Cas9 knockout screens are better than previous functional shRNA knockdown screens (Berns et al. 2004; Luo et al. 2008) for the study of phenotypes that require complete inactivation of genes (Shalem et al. 2014). CRISPRi/a are more flexible than shRNA in defining gene functions across a wider dynamic range (Gilbert et al. 2014). In light of this, several sgRNA libraries with diverse configurations have been developed for CRISPR/Cas9 knockout or CRISPRi/a screens in human and mouse (Gilbert et al. 2014; Koike-Yusa et al. 2014; Shalem et al. 2014; Wang et al. 2014; Zhou et al. 2014; Konermann et al. 2015).
The design of sgRNA is critical to the CRISPR-based screens. In a CRISPR system, an sgRNA contains a spacer sequence complementary to the targeted DNA sequence to guide the Cas9 or dCas9 proteins to genomic targets. Several studies have investigated sgRNA off-target effects based on the alignment of spacer sequences to the genome (Hsu et al. 2013), dCas9 ChIP-seq (Wu et al. 2014) and GUIDE-seq (Tsai et al. 2015). On the other hand, DNA sequence also influences the efficiency of sgRNA at the on-target genomic loci. It is well established that the Protospacer Adjacent Motif (PAM) containing NGG consensus is required for Cas9-DNA binding and cleavage in a CRISPR/Cas9 system (Sternberg et al. 2014). A recent study showed that the sgRNA efficiency depends on the purine/pyrimidine composition near the 3′ end of the spacer sequence (Wang et al. 2014). Moreover, Doench et al. (2014) found the nucleotide composition of the DNA downstream from the spacer target also contributes to the sgRNA efficiency and proposed a predictive sequence model for the design of CRISPR/Cas9 knockout experiments.
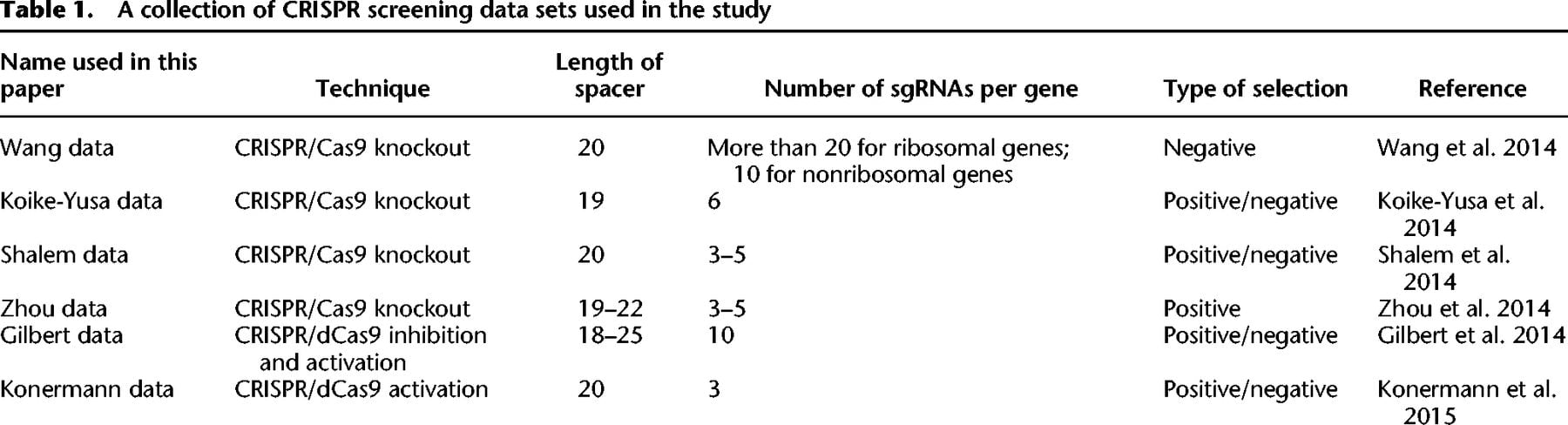
Despite these advances, it is unclear whether the sequence features reported in previous studies are reproducible across different sgRNA library designs and genomes. Importantly, these features were only tested on negative selections using CRISPR/Cas9 knockout systems, in which loss-of-function of a gene results in growth disadvantage to the cells. For model justification and broader applications, it is necessary to investigate their relevance to positive selections in which gene perturbation promotes cell growth and proliferation. Moreover, no sequence model has been developed for predicting sgRNA efficiency in CRISPRi/a screens. To fill these gaps, we systematically assessed the effect of sequence context on sgRNA efficiency based on six published data sets (Table 1). Our analysis led to the development of two computational models for predicting sgRNA efficiency, corresponding to the CRISPR knockout and CRISPRi/a studies, respectively. To this end, our findings and models facilitate the genome-wide design of improved sgRNA for CRISPR-based screens.
A collection of CRISPR screening data sets used in the study
Results
Selection of the efficient and inefficient sgRNAs in CRISPR/Cas9 knockout screens
To construct a predictive model, we took two steps to identify efficient and inefficient sgRNAs in published data sets. First, for each data set, we identified a list of essential genes whose deletion resulted in a growth disadvantage in genome-wide CRISPR/Cas9 knockout screens. Second, the sgRNAs targeting these essential genes were grouped into “efficient” and “inefficient” categories based on their decline in abundance in the screens. We applied our method to Wang and Koike-Yusa data (see Table 1). These data sets have large numbers of sgRNAs per gene, which help to identify reliable sets of essential genes.
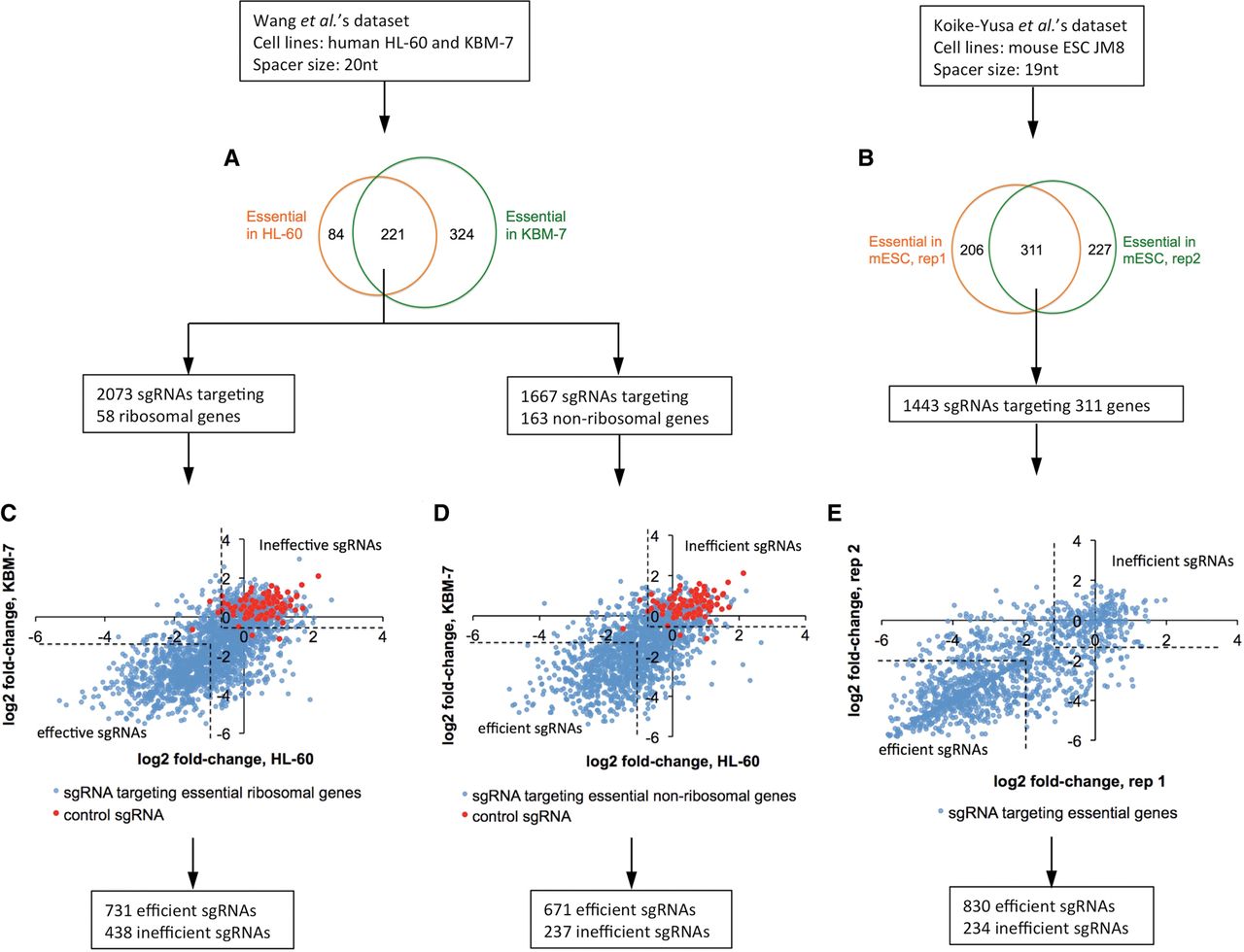
The procedures of sgRNA selection and categorization are depicted in Figure 1. In Wang data, we identified 305 and 545 essential genes in the leukemia cell lines HL-60 and KBM-7, respectively (Fig. 1A). Among them, 58 ribosomal and 163 nonribosomal genes were essential in both cell lines. The essentiality of ribosomal genes was also reported in other studies (Koike-Yusa et al. 2014; Shalem et al. 2014), which established them as good positive controls in a CRISPR/Cas9 knockout screen. In the Wang library, each ribosomal gene was targeted by more than 20 sgRNAs, and each nonribosomal gene was targeted by 10 sgRNAs. Therefore, we split the sgRNAs targeting common essential genes into “ribosomal” and “nonribosomal” sets in order to take the difference of sgRNA sample space into consideration. We examined signals of sgRNAs by comparing their abundance before and after selection. For the “ribosomal” set, 731 (35.3%) sgRNAs showed more than twofold depletion in both cell lines, indicating that they reproducibly knocked out their target genes in the experiments and thus were defined to be efficient. Four hundred thirty-eight (21.1%) failed to show significant differences from control sgRNAs that do not target any genomic locus and thus were defined to be inefficient. The remaining sgRNAs showed relatively weak effects in both cell lines, or had significant effects in only one cell line, and were excluded from our analysis (Fig. 1B). For the “nonribosomal” set, we selected 671 efficient and 237 inefficient sgRNAs using the same criteria (Fig. 1C). In Koike-Yusa data, we identified 311 genes that showed essentiality in two biological replicates of mouse embryonic stem cells (mESCs) (Fig. 1D). Upon examining the signals of sgRNAs targeting these genes, we found a bimodal distribution of the relative sgRNA abundance in cell growth (Fig. 1E; Supplemental Fig. 1). Categorization based on the bimodality resulted in 830 efficient and 234 inefficient sgRNAs.
A schematic view of procedures for sgRNA selection and categorization. (A,B) Venn diagrams showing the overlap of essential genes between human HL-60 and KBM-7 cells (A) and two biological replicates in mouse ESC JM8 cells (B). (C–E) Scatter plots showing the log2 fold-change of sgRNA abundance in negative selection upon cell growth. (C) sgRNAs targeting essential ribosomal genes in Wang data. (D) sgRNAs targeting essential nonribosomal genes in Wang data. (E) sgRNAs targeting essential genes in Koike-Yusa data. The dashed lines represent the threshold chosen to determine efficient and inefficient sgRNAs.
To this end, we selected three training sets of sgRNAs, namely “ribosomal,” “nonribosomal,” and “mESC,” for the construction of sequence models (Supplemental Table 1). The differences among these sets reflect variations of sgRNA design, species, and the experimental protocols.
Sequence features associated with sgRNA efficiency in CRISPR/Cas9 knocking out
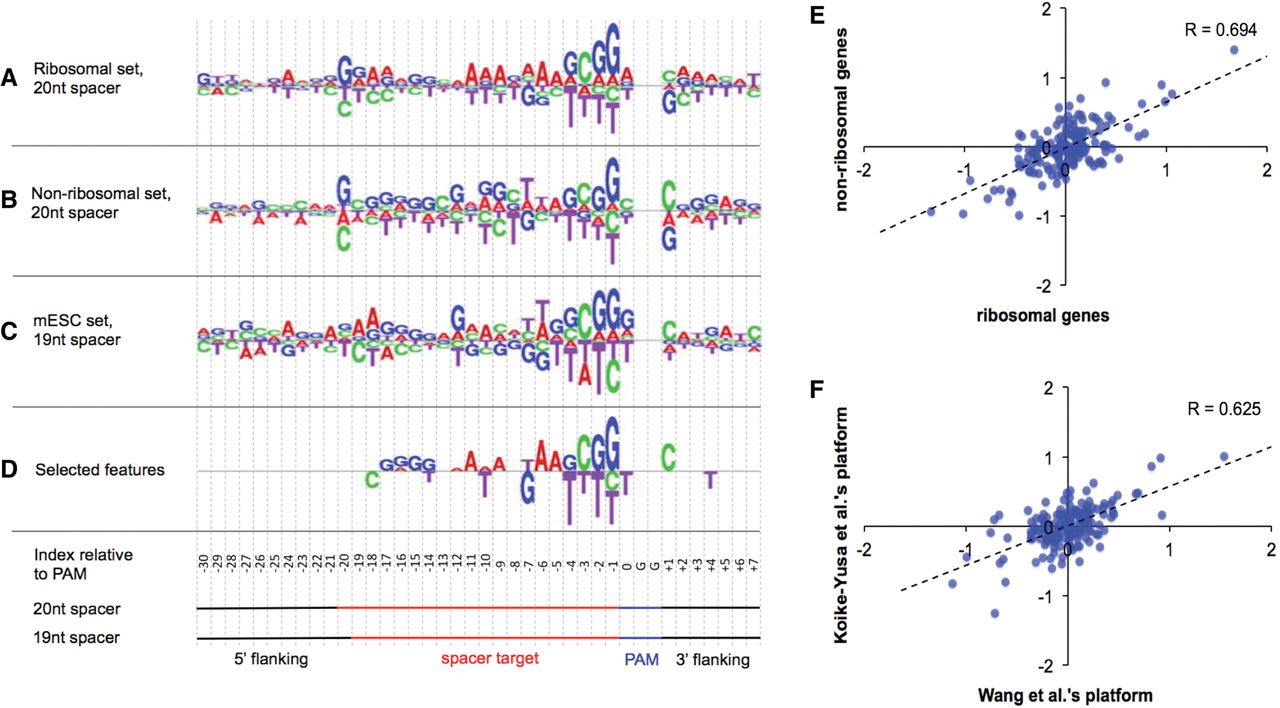
To explore sequence features that contribute to sgRNA efficiency, we computed the log odds ratio of nucleotide frequency between DNA sequences targeted by efficient and inefficient sgRNAs (Fig. 2A–C). These sequences are 40 bp in length, including the 19-bp or 20-bp spacer targets as well as their 3′ and 5′ flanking DNAs. All sequences were aligned at the PAM. The log odds ratios are correlated between the ribosomal and nonribosomal sgRNA sets (Fig. 2E) as well as between the two libraries developed independently (Fig. 2F). The correlations indicate that many sequence features are robust against the variation of sgRNA designs, species, and the spacer length. Meanwhile, some features are discordant among the three different sgRNA training sets. For example, guanine is preferred at the 5′ end of the spacer in the “ribosomal” and “nonribosomal” sets, but not in the “mESC” set. This might be ascribed to subtle differences in sgRNA design. If the 5′ end of the spacer is not “G,” Wang et al. prepended a “G” for the expression of sgRNAs from a U6 promoter, in the form of G(A/C/T)X19NGG; otherwise, a form of GX19NGG was taken without any prepended “G” (Wang et al. 2014). Alternatively, Koike-Yusa et al. chose the form of GX19NGG regardless of nucleotide composition at the 5′ end (Koike-Yusa et al. 2014). These different designs of sgRNAs can result in the variation of the nucleotide pairings and 5′ overhang in sgRNA-DNA binding structure, which could further impact sgRNA efficiency.
Preference of nucleotide sequences that impact sgRNA efficiency. (A–C) Logos showing the sequence preference of the three sgRNA sets defined in Figure 1. The height of the nucleotides represents the log odds ratio of nucleotide frequency between efficient and inefficient sgRNAs. (D) A logo showing the selected features that reproducibly impact sgRNA efficiency in the three sgRNA sets. The height of the nucleotides represents the coefficients computed from the Elastic-Net. (E,F) Scatter plots showing the correlation of sequence preference for sgRNAs targeting ribosomal versus nonribosomal genes in Wang data (E) and sgRNAs in Wang data versus Koike-Yusa data (F). Each dot represents a nucleotide in a 40-bp region centered by the spacer. The sequence preference is measured as the log2 odds ratio of nucleotide frequency between efficient and inefficient sgRNAs.
To determine the reproducible features, we first selected nucleotides that satisfy the following criteria across the three training sets: (1) Signs of the odds ratios are concordant; and (2) magnitudes of the odds ratios are above a threshold in all three sgRNA sets, where the threshold was computed from a statistical significance analysis (see Methods). To identify the dominating nucleotide features, we next applied Elastic-Net, a penalized linear regression model with proven utility in feature selection (Zou and Hastie 2005). In our analysis, the predictor variable of Elastic-Net was a binary vector representing the presence or absence of the nucleotides selected based on the odds ratio, and the response variable was 1 or −1 corresponding to the efficient and inefficient sgRNAs, respectively. The union of the three sgRNA training sets was used for Elastic-Net feature selection and parameter estimation.
With this two-stage feature selection approach, we identified 28 sequence features, as shown in Figure 2D. Most features are located within the spacer region. Our result confirmed several features reported previously. First, guanines are strongly preferred at the −1 and the −2 positions proximal to the PAM sequence, which are associated with the sequence preference in Cas9 loading (Wang et al. 2014). Second, thymines are disfavored at the four positions closest to the PAM, consistent with the fact that multiple uracils in the spacer cause low sgRNA expression (Wu et al. 2014). Third, in line with recent findings, the nucleotides downstream from the PAM contribute to sgRNA efficiency, whereas the sequences upstream of the spacer have no significant effect (Doench et al. 2014). We also identified novel features that reproducibly impact the sgRNA efficiency. For example, cytosine is preferred at the −3 position, which is the DNA cleavage site by the CRISPR/Cas9 complex (Cong et al. 2013). This feature might contribute to the efficiency of cleavage or the introduction of mutation upon DNA repair. Moreover, adenines are preferred from position −5 to −12, and guanines are preferred at positions −14 to −17.
Experimental validation of the sequence model in predicting mutation rates and protein knockout efficiency
To experimentally test the model in Figure 2D, we first assessed the mutation rates mediated by a small set of sgRNAs targeting the AAVS1 safe harbor genomic locus. We selected 10 targets with low efficiency scores and 10 with high scores in the AAVS1 region (Supplemental Table 3) and designed CRISPR/Cas9 lentivirus based on the selected targets. Upon examining the transduced 293T cells, we found that sgRNAs with higher scores showed much higher indel mutation rates than those with lower scores (P = 1.9 × 10−5; t-test) (Fig. 3A). This confirmed that the sequence features we selected have significant effect on the rate of on-target mutation in a CRISPR/Cas9 system.
Experimental validation of the sequence model in predicting sgRNA efficiency. (A) A SURVEYOR gel picture (top) and a bar chart (bottom) showing the indel rates of the sgRNAs predicted to be inefficient (low sequence score) or efficient (high sequence score). The sgRNAs were selected to target the AAVS1 locus. The experiment was conducted in 293T cells. (B) A scatter plot showing the correlation of the predicted sequence scores and the protein knockout efficiency for sgRNAs targeting AR and FOXA1 in LNCaP-abl cells. The knockout efficiency is measured as the percentage of reduction in protein level upon sgRNA infection.
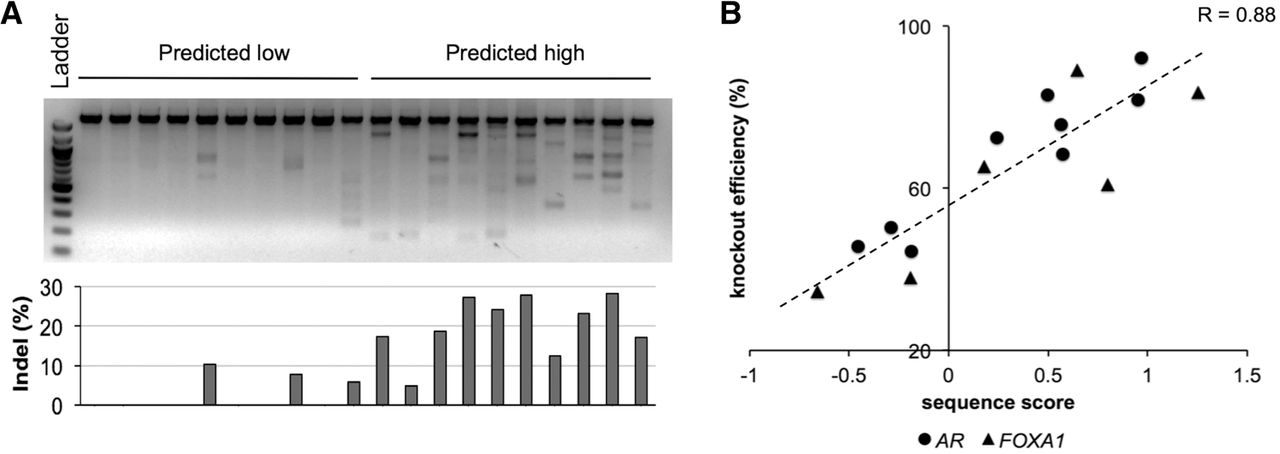
We next tested if the sequence model is informative to predict knockout efficiency on the protein expression level. We designed sgRNAs against AR and FOXA1 and tested them in prostate cancer LNCaP-abl cells using Western blot to assay protein expression (Supplemental Fig. 2; Supplemental Table 4). The predicted sgRNA efficiency score showed strong positive correlation with the protein knockout efficiency (P = 9.0 × 10−6; Pearson's correlation) (Fig. 3B), where the efficient and inefficient sgRNAs can be clearly separated. Our result suggested a cutoff of zero as a reasonable threshold for sgRNA classification, in line with the configurations in model training, in which the values of 1 and −1 were assigned to efficient and inefficient sgRNAs, respectively.
Predictive power of the sequence model in negative selection screens
With the satisfactory results in the low-throughput validation mentioned above, we next assessed the predictive power of the sequence model using published high-throughput CRISPR/Cas9 knockout screening data. We designed four schemes for this in silico validation. First, randomized threefold cross validation was carried out on the categorized sgRNAs in the ribosomal set to assess the predictive power under identical experimental settings; second, the Elastic-Net model was trained on the ribosomal set and tested on the nonribosomal set, of which the result was expected to reflect the within-library performance under different configurations of sgRNA sampling; third, to evaluate the inter-library and inter-species performance, the model was trained on the union of ribosomal and nonribosomal sets in human and tested on the mESC set; last, the model was trained on the union of all three sets and tested on an independent library of sgRNAs (Shalem et al. 2014). The sgRNA list used in the fourth validation was compiled by Doench et al. (2014) to assess their model in predicting sgRNA “activity,” a measure analogous to the “efficiency” defined in this paper. We achieved reasonable predictive power in all four validations, in which the Area Under Curve (AUC) scores were above 0.7 in Receiver Operating Characteristic (ROC) (Fig. 4A,B). Our model also outperformed the model proposed by Doench et al. (2014) in predicting sgRNA activity (Fig. 4B). With a cutoff threshold of zero, 50%–60% of the inefficient sgRNAs are predictable, at the cost of 10%–20% of efficient sgRNAs misclassified.
Predicting sgRNA efficiency from sequence context in CRISPR/Cas9 knockout screens. (A) ROC curves showing the predictive power of the proposed model. (Red) Threefold cross-validation on the sgRNAs targeting ribosomal genes in Wang data; (blue) trained on ribosomal genes, and tested on nonribosomal genes in Wang data; (green) trained on Wang data, and tested on Koike-Yusa data. The black error bars on the red curve represent standard deviations computed from 10 iterations of random sampling in cross-validation. (B) ROC curves comparing the performance of the proposed model and the Doench et al. (2014) model in predicting sgRNA efficiency in Shalem data. (C) Scatter plot showing the correlation between the predicted sequence score and the relative sgRNA abundance for ABL1 and BCR in KBM-7 cells. The P-values were computed based on the Pearson correlation test. (D) Box plot showing the distributions of correlations between sequence scores and relative sgRNA abundances for essential and nonessential genes in KBM-7. The distribution of random background was computed by permuting the sequence scores within each gene in the data set. (E) Distributions of relative sgRNA abundances in KBM-7 cells, where the sgRNAs were categorized based on the predicted efficiency and the essentiality of their targeted genes.
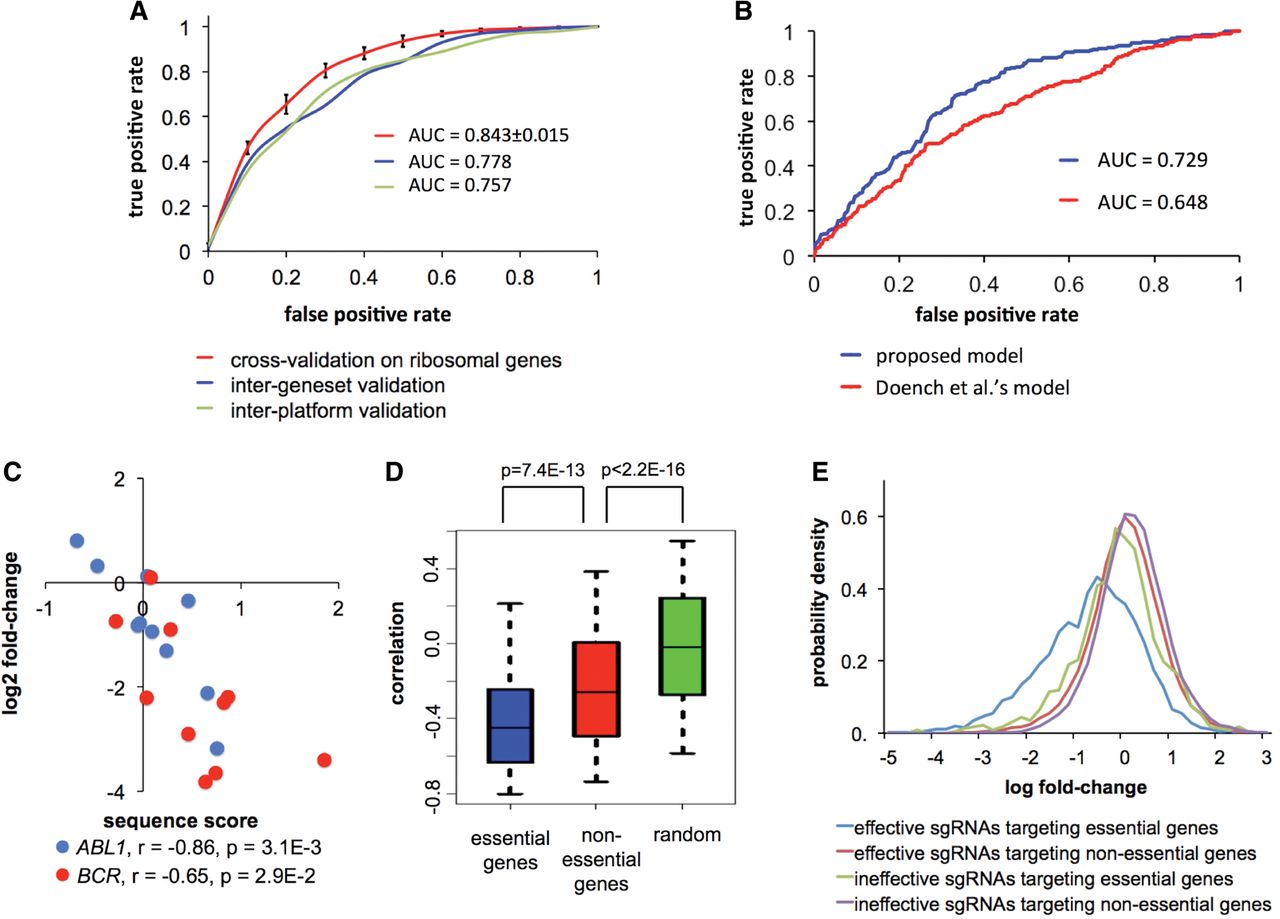
Since the sequence model was trained based on the sgRNAs targeting common essential genes, we further tested if the model is applicable to the sgRNAs targeting cell-type-specific essential genes. We examined the correlations between sequence scores and the relative sgRNA abundances for ABL1 and BCR, two genes coding for an oncogenic fusion protein in KBM-7 cells (Andersson et al. 1995). For both genes, significant correlations were observed between predicted sgRNA efficiency and the changes of sgRNA abundance in the screen (Fig. 4C). To evaluate whether sequence-specific sgRNA efficiency also influences sgRNAs targeting nonessential genes, we compared distributions of the correlations for essential and nonessential genes identified in KBM-7 (Fig. 4D) or HL-60 (Supplemental Fig. 3). As expected, sequence scores and relative sgRNA abundances are in strong negative correlation for essential genes. Meanwhile, there are weak but statistically significant negative correlations for nonessential genes, which might be attributed to (1) the false negatives in identifying essential genes; (2) decreased cell growth due to intrinsic toxicity of DNA cleavage, similar to the toxicity reported on Cre recombinase (Silver and Livingston 2001); and/or (3) potential off-target effects of sgRNAs. Notably, no correlation was observed for those control sgRNAs lacking genomic targets (Supplemental Fig. 4), consistent with the requirement of chromatin binding for an sgRNA to exert its cell functions.
We next checked if a proper selection of sgRNAs based on the prediction result could refine the sgRNA pool. As shown in Figure 4E, the signals of essential and nonessential genes are not well separable with the predicted inefficient sgRNAs. In contrast, the efficient sgRNAs produced highly informative signals that distinguish essential genes from others. This indicates that the removal of inefficient sgRNAs can reduce the number of required sgRNAs without compromising the sensitivity of the CRISPR/Cas9 knockout screens.
Generality of the sequence model in positive selection screens
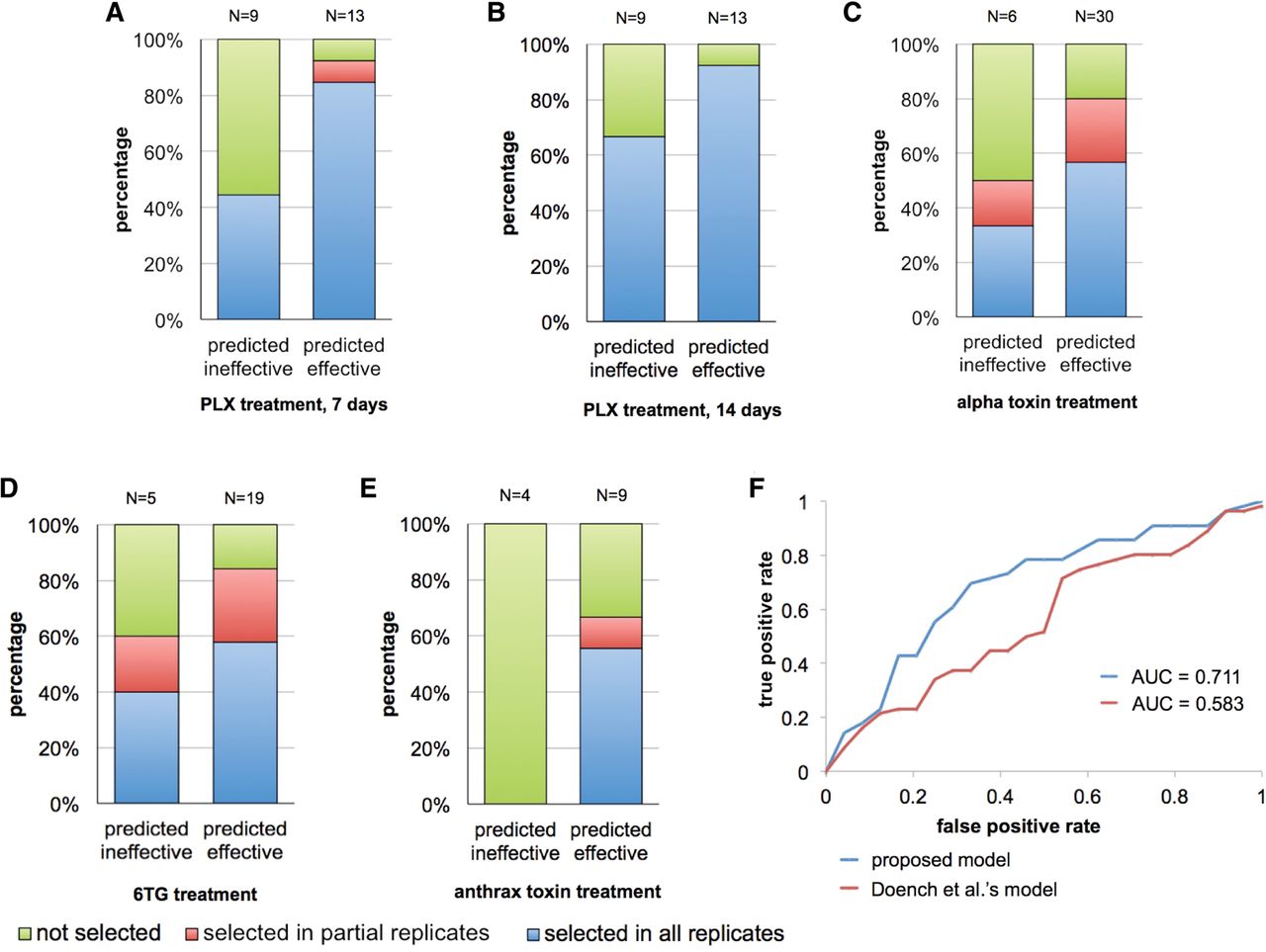
Our model was trained based on the sgRNAs targeting essential genes that were negatively selected in the experiments. We next asked if the model is applicable to positive selection screens. To answer this, we curated four lists of genes involved in the resistance to drug or toxin treatments, including the BRAF inhibitor Vemurafenib (PLX) in melanoma A375 cells (Shalem et al. 2014), alpha-toxin and 6-thioguanine (6TG) in mouse embryonic stem cells (Koike-Yusa et al. 2014), and anthrax toxin in HeLa cells (Supplemental Table 5; Zhou et al. 2014). The sgRNAs targeting these genes were categorized based on the reproducibility of positive selection, i.e., the increase of the sgRNA abundance in multiple biological replicates upon treatments. As shown in Figure 5A–E, the predicted efficient sgRNAs expressed higher reproducibility of positive selection in all the experiments, indicating our model's applicability to positive selection screens. Next, we assessed the predictive power of our model and the model by Doench et al. (2014) on the curated sgRNA sets, in which the sgRNAs “selected in all replicates” were defined as positives and the sgRNAs “not selected in any replicates” were defined as negatives. Our model achieved an AUC score of 0.711 and significantly outperformed the Doench et al. (2014) model (Fig. 5F). Notably, 42.3% of the efficient sgRNAs contain a cytosine at the cleavage site, in comparison with 25.9% for inefficient sgRNAs, suggesting this novel feature also contributes to sgRNA efficiency in positive selections.
Assessment of the sequence models in predicting sgRNA efficiency in positive selection experiments. (A–E) Bar charts showing the capability of selection and the experimental reproducibility for predicted efficient and inefficient sgRNAs. The tested sgRNAs target the genes known to be involved in the resistance to different drug treatment or external stimulus. (F) ROC curves comparing the performance of the proposed model and the Doench et al. (2014) model in predicting sgRNA efficiency in positive selection experiments. In the evaluation, the positive test set consists of the sgRNAs selected in all replicates in B–E; and the negative test set consists of those not selected in B–E.
Our prediction is therefore consistent with the phenotype in both positive and negative selections, where the abundance of efficient sgRNAs is expected to change in opposite directions. This line of evidence, together with our validation results on mutation rate and protein knockout efficiency, confirmed that the selected sequence features impact cell phenotype mainly through the loss-of-function of the target genes, but not through their intrinsic toxicity on cell growth.
Sequence-specific sgRNA efficiency in CRISPRi/a screens
Recently, Gilbert et al. (2014) proposed to use a CRISPRi/a system for genome-wide functional screens. To explore the impact of sequence context on the efficiency of sgRNAs in their study, we selected 6624 sgRNAs targeting the 500 most essential genes in the published CRISPRi negative selection data in K562 cells (Supplemental Table 6). These sgRNAs were grouped into “efficient” and “inefficient” based on the phenotype scores that measure the relative sgRNA abundance (Fig. 6A).
Preference of the length and sequence context of spacers in CRISPR/dCas9 inhibition (CRISPRi) and activation (CRISPRa) screens. (A) Distribution of phenotype scores (Gilbert et al. 2014) for sgRNAs targeting the top 500 essential genes and the control sgRNAs in CRISPRi experiments. The dashed line represents the threshold chosen to determine efficient and inefficient sgRNAs. (B) A bar chart showing the effect of spacer length on sgRNA efficiency. (C) Logos showing the sequence preference of spacers. The height of the nucleotides represents the log odds ratio of nucleotide frequency between efficient and inefficient sgRNAs. The nucleotide at the 5′ end of the spacers is fixed to be guanines in the library design and is excluded from the logos. (D) Bar charts comparing the performance of CRISPRi model and CRISPR/Cas9 KO model in predicting sgRNA efficiency in CRISPRi negative selection, CRISPRi positive selection upon CTx-DTA treatment, and CRISPRa negative selections in Gilbert data and Konermann data. The length of spacers is 20 nt. Cross-validation was used to assess the performance of the CRISPRi model in the CRISPRi negative selection experiment. The error bars represent the standard deviations in 10 iterations of threefold cross validation. The P-value was computed using a paired t-test.
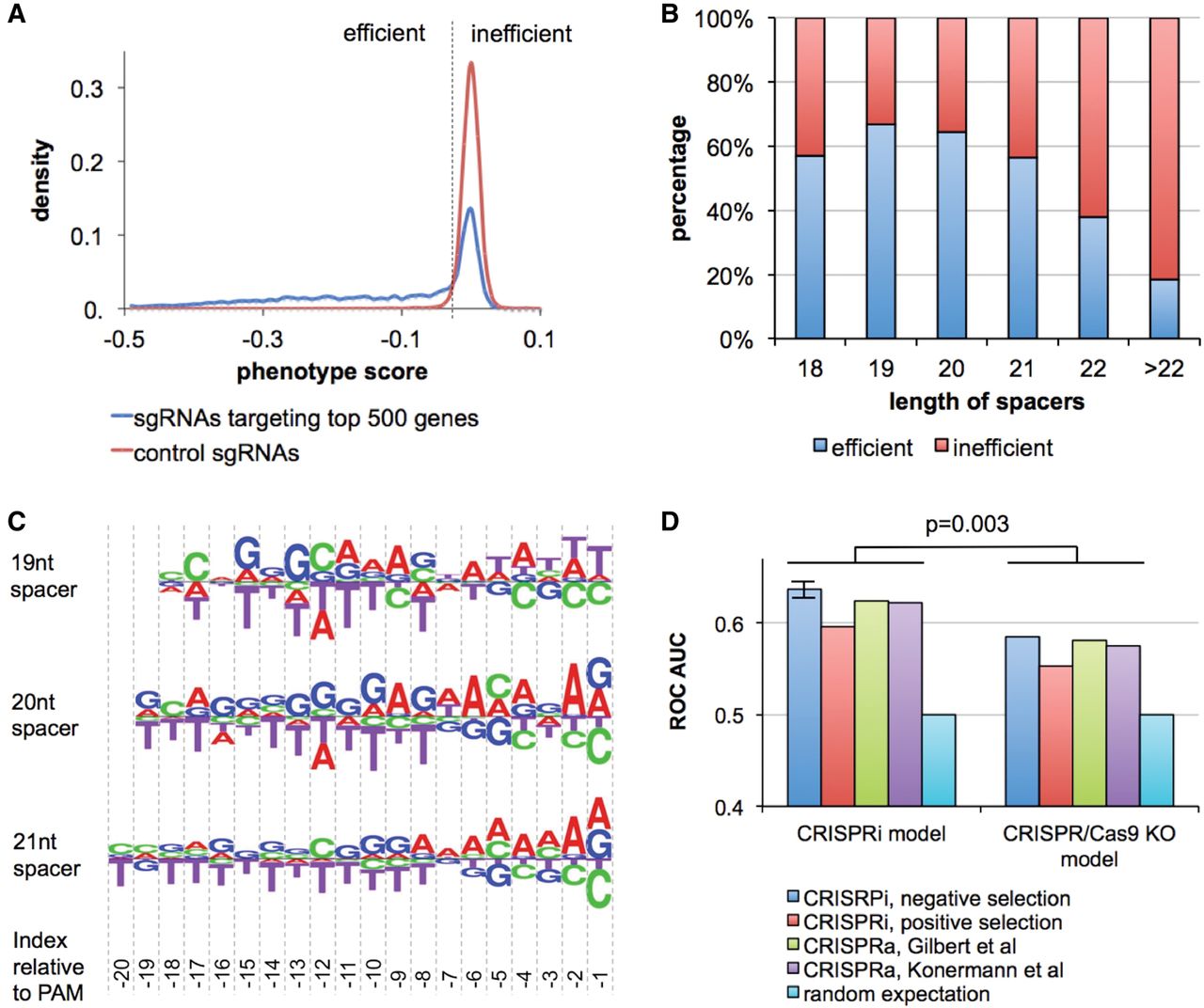
Since the spacers have variable lengths in this data set, we first examined the percentage of efficient sgRNAs with different spacer lengths. We found sgRNAs with a spacer length of 19 nt have the highest efficiency, slightly better than those with 20 nt spacers (Fig. 6B). Despite the previous report that truncated sgRNAs with 17- to 18-nt spacers have less off-target effects (Fu et al. 2014), shorter spacers showed less on-target effect in comparison with 19-nt or 20-nt spacers. To remove the confounding effect of spacer length, we computed the log odds ratio of nucleotide frequency for the efficient and inefficient sgRNAs based on different spacer lengths (Fig. 6C). Similar to CRISPR/Cas9 knockout, the CRISPRi/a system prefers purines at most of the nucleotides in the spacer. However, there are also significant differences. In CRISPR/Cas9, a small number of nucleotides, mostly adjacent to the PAM, dominate the sequence preference (Fig. 2A–D). In contrast, many nucleotides throughout the spacer region collectively contribute to sgRNA efficiency in the CRISPR/dCas9 system. The disparity can be ascribed to several factors. First, unlike CRISPR/Cas9 knockout, CRISPRi/a does not introduce DNA cleavage, which explains the observation of no cytosine enrichment at the −3 position. Second, the structural and functional differences between Cas9 and dCas9 can lead to diverse sequence preference for protein loading and the initiation of gRNA-DNA pairing. Third, the effector domains that play crucial roles for gene perturbation in CRISPRi/a do not exist in a CRISPR/Cas9 system. Fourth, the U6 promoter and the sgRNA constant region are different between the Gilbert et al. (2014) design and other platforms (Chen et al. 2013). Last but not least, CRISPRi/a mainly targets gene promoters, where the sequence contexts are substantially different from those in coding regions.
To evaluate the sequence-specific sgRNA efficiency in CRISPRi/a, we designed linear classifiers using the Elastic-Net and validated the classifiers with four settings: (1) threefold cross-validation on the sgRNAs targeting negatively selected essential genes in the CRISPRi experiment; (2) training on the essential genes and testing on the positively selected genes upon CTx-DTA treatment; (3) training on the essential genes in the CRISPRi experiment and testing on the growth-inhibiting genes in CRISPRa data (Gilbert et al. 2014); and (4) repeating the training process in (3) and testing in Konermann CRISPRa data (Konermann et al. 2015). We achieved ROC-AUC scores of 0.6–0.65 in the four validations, which indicated that the sequence model is applicable to positive selections, CRISPRa experiments, as well as to the CRISPRi/a systems of independent designs (Fig. 6D). There is less predictive power when applying the CRISPR/Cas9 knockout model on CRISPRi/a data (ROC-AUC < 0.6). Notably, comparing with the higher predictive power in CRISPR/Cas9 knockout screens (ROC-AUC > 0.7), we found the sgRNA efficiency in CRISPRi/a to be less dependent on the sequence context. It is possible that transcription factor binding and chromatin states at the target sites of CRISPRi/a influence knockdown efficiency, which cannot be explained by the sequence context alone.
Discussion
We have systematically assessed the contributions of sequence features to sgRNA efficiencies, which are critical to the performance of genome-wide CRISPR based functional screens. To achieve sufficient statistical power, at least hundreds of sgRNA samples are needed for the analysis (see Supplemental Material). Therefore, we sought to collect samples from published high-throughput screening data sets. In the study of CRISPR/Cas9 knockout screens, we used an integrative method to select sequence features that are reproducible among different data sets. Such a method is helpful to improve the robustness of the sequence model for the prediction in various applications. In addition to known features reported previously, our model suggested novel features to better understand the molecular mechanism of the CRISPR/Cas9 genome-editing systems. We demonstrated the predictive power of our model through multiple validations—including tests on independent designs, various cell types, and different growth selections—which support the robustness of the model for customized sgRNAs designs.
Despite these findings, ∼40% of inefficient sgRNAs are not predictable with our sequence model, which can be ascribed to sgRNA secondary structure, chromatin structure, exon-skipping during transcription, strand and location (e.g., 5′ versus 3′ end) of sgRNA target relative to the gene, possibly nonlinear interactive effects among different positions, as well as the sgRNA-specific likelihood of a frame shift (Doench et al. 2014; Wang et al. 2014). The performance of the model might be further improved by including these other features in the future. Another limitation of our study is the relatively small number of cell lines (five for CRISPR/Cas9 KO and two for CRISRPi/a) and library designs for the training and testing of our computational model. It is possible that some cell-specific sequence preferences were missed in our study. With the expected growth of CRISPR data sets, more comprehensive and systematic assessment could be helpful to refine the predictive model.
It is worth noting that selection against sgRNAs targeting nonessential genes is also dependent on the sgRNA efficiency in a moderate but statistically significant manner. This can be partially explained by false negatives in identifying essential genes and the potential off-target effects of sgRNAs. On the other hand, it has been reported that the DNA cleavage induced by Cre recombinase results in growth retardation of cells that is independent of the targeted genomic loci (Silver and Livingston 2001). Therefore, it is possible that the on-target DNA cleavage introduced by CRISPR/Cas9 system might have similar toxicity on cell growth. Future work is needed to validate this hypothesis, and to normalize the CRISPR/Cas9 screen data against the sgRNA toxicity and efficiency.
In this study, we found that the sequence preference in CRISPRi/a is significantly different from that in CRISPR/Cas9 knockout. This is expected since the CRISPRi/a system uses a distinct mechanism to perturb gene function without introducing DNA cleavage. We note that our analysis is based on published data sets in which coding regions were chosen for CRISPR/Cas9 targeting and promoters were chosen for CRISPRi/a targeting. The difference of nucleotide composition between the coding regions and the promoters, such as codon structure and GC enrichment, might introduce bias to our analysis. It would be interesting to compare these two systems with an unbiased design in future study. Despite this, our validation result on AAVS1 locus suggests that the CRISPR/Cas9 knockout model is also applicable to noncoding regions.
The models proposed in this study focus on sgRNA sensitivity. Meanwhile, the specificity of sgRNAs should also be considered when designing an sgRNA pool. Wu et al. (2014) observed numerous off-target binding sites at the genomic loci with sufficient sequence similarity to the sgRNA spacer and reported that thymine-rich sgRNA spacers have fewer off-target sites than others due to the low level of sgRNA expression and the scarcity of complementary sites in accessible chromatin. This finding, however, is not contradictory to our model showing disfavored thymine near the 3′ end of sgRNA target, but rather reflects the requirement of an appropriate tradeoff between specificity and sensitivity in sgRNA design. Surprisingly, a recent study showed that the majority of off-target cleavage sites are undetectable using existing computational methods or dCas9 ChIP-seq (Tsai et al. 2015). Further studies are needed to address both sgRNA efficiency and off-target effect for optimal sgRNA designs.
Methods
Preprocessing
Suppose there are N experiments performed on a set of M sgRNAs, and the read count of sgRNA i on experiment j is xij, . To normalize against the sequencing depth and the global trend of cell growth, we assume that a majority of the sgRNA-targeted genes have no effect on cell growth. Therefore, we adjusted read count , where sj is the median read count of all sgRNAs in the jth experiment. This median normalization approach was applied to Wang, Koike-Yusa, Shalem, and Konermann data. For Zhou data, the median read count was zero in the experiments after positive selection since the strength of selection was very strong so that a majority of the sgRNAs were completely depleted except a few that contribute to drug resistance. In this case, we set the normalization factor sj to be the average read count of all sgRNAs. Normalization in such a way might introduce bias to the estimation of relative sgRNA abundance, but it will not affect the conclusion drawn from Figure 5E because the bias is a constant added to all sgRNAs.
Next, we compute the relative sgRNA abundance between two experiments. Suppose and are the adjusted read counts of the ith sgRNA in the jth and kth experiments, the relative sgRNA abundance is computed as the log2-transformed ratio
For negative selection in Wang data, the relative sgRNA abundance was computed by comparing the cell population after 12 cell doublings against the initial seeding states in HL-60 and KBM-7 cells, respectively. For negative selection in Koike-Yusa's data set, the transfected mESCs cultured for 6–7 d were compared against plasmid control. For positive selection in Koike-Yusa data, the mESCs treated with toxin were compared against plasmid control. For positive selection in Shalem data, A375 cells with PLX treatment were compared against the population without PLX treatment at day 7 and day 14. For positive selection in Zhou data, HeLa cells after three rounds of toxin treatment were compared against plasmid control. For negative selection in Konermann CRISPRa data, A375 cells with DMSO treatment were compared between day 21 and day 3. The negative selection data by Shalem et al. (2014) and the CRISPRi/a data by Gilbert et al. (2014) were not preprocessed, as the phenotypic measure of the sgRNAs were retrieved directly from their studies.
Selecting phenotype-associated genes
For the CRISPR/Cas9 knockout data sets, we ran the beta-0.1 version of RIGER (http://www.broadinstitute.org/cancer/software/rnai/riger/) to call the negatively selected genes (Luo et al. 2008). In the configurations of RIGER, the measures of relative sgRNA abundance were used to rank the sgRNAs, and a Kolmogorov-Smirnov test was chosen to determine the significant genes. The threshold of P-values was set to be 0.001. The positively selected genes in Figure 5 were collected based on experimental validations in the original studies (Koike-Yusa et al. 2014; Shalem et al. 2014; Zhou et al. 2014), and a cutoff threshold of two folds in relative sgRNA abundance were used to determine efficient and inefficient sgRNAs. The phenotype-associated genes in the CRISPRi/a data set were retrieved from the original paper (Gilbert et al. 2013).
Thresholds for identifying efficient and inefficient sgRNAs
As shown in Figure 1, we used different thresholds for identifying efficient and inefficient sgRNAs in Wang data and in Koike-Yusa data. This is because the sgRNAs targeting essential genes in Wang data and Koike-Yusa data have different distributions of log ratios due to the variance of cell growth rate, Cas9 activity, and the depth of selection in the experiments. In Wang data, the sgRNAs with a log ratio smaller than −1.0 in both HL-60 and KBM-7 were identified to be efficient, and the thresholds for inefficient sgRNAs are in the fifth percentile of the log ratios of control sgRNAs in HL-60 and KBM-7. The thresholds in Koike-Yusa data were determined based on the bimodal distribution of signals (Supplemental Fig. 1). Notably, when the thresholds in Koike-Yusa data were set to be identical to those used in Wang data, we observed little variation of nucleotide frequency (r = 0.96). Therefore, our sequence model is insensitive to the threshold chosen.
Extracting sequence features
For Wang, Koike-Yusa, and Shalem data, sequences of spacer targets and their flanking regions were extracted from hg19 or mm9 genome assembly. Since the length of sgRNA spacers varies from 18 bps to 22 bps in the Zhou et al. (2014) platform, we mapped the spacer sequences to hg19 assembly to find the genomic loci of the targets and extracted the 40-bp sequence (spacer + 3′ + 5′) based on the aligned PAM loci. The sequences in CRISPRi/a data were retrieved from the original paper (Gilbert et al. 2014; Konermann et al. 2015).
Determining reproducible features in CRISPR/Cas9 knockout screens
To measure the sequence preference, we computed the log2 odds ratio of nucleotide frequency between efficient and inefficient sgRNAs, for the “ribosomal,” “nonribosomal,” and “mESC” sets. Let , , and represent the log2 odds ratios for the ith nucleotide computed from the 3 sgRNA sets, we define a score si as
To determine the background distribution of si, we randomly permuted the sgRNAs in the efficient and inefficient categories within each set and computed si based on the permuted data. This results in an empirical null distribution of si. A threshold T was then set to be at the 95% confidence interval of the null distribution, and the nucleotides with were selected as reproducible features.
The Elastic-Net model
Suppose is the set of encoded sequence vectors, and is the set of outputs representing the efficiency of sgRNAs, where N is the number of sgRNA samples for training. Let M be the length of the input vectors, the Elastic-Net model computes the parameters that minimize an objective function E
When training the Elastic-Nets for sequence feature selection, we found the optimal α to be 1.0 for CRISPR/Cas9 knockouts and 0.0 for CRISPRi/a (Supplemental Figs. 5, 6). We note that the Elastic-Net is equivalent to the Least Absolute Shrinkage and Selection Operator (LASSO) when α = 1.0 (Tibshirani 1996) and is equivalent to the Ridge regression model when α = 0.0 (Tikhonov and Arsenin 1977). In practice, the LASSO model is preferred to select a small number of features from a large amount of candidates, whereas the Ridge regression is more relevant in situations when many features cumulatively contribute to the response. Therefore, our result on the selection of optimal α is explainable since the sgRNA efficiency depends on fewer sequence features in Cas9 knockout in comparison with that in CRISPRi/a.
Application of previous model
To apply the Doench et al. (2014) model, we downloaded the python script at http://www.broadinstitute.org/rnai/public/analysis-tools/sgrna-design. For fair comparison, the tested sgRNA targets and our regression model (Fig. 2D) were truncated into a 30-bp sequence harboring the spacer target, following the requirement in the Doench et al. (2014) model.
Cell lines and cell culture
The LNCaP-abl (abl) cell line was provided by Zoran Culig (Innsbruck Medical University, Austria). The abl cells were cultured in the RPMI 1640 phenol red-free medium supplemented with 10% charcoal/dextran- treated fetal bovine serum, 2 mM glutamine, 100 µg/mL penicillin, and 100 units/mL streptomycin for the experiments. The 293T cells obtained from the American Type Culture Collection were maintained in DMEM media supplemented with 10% fetal bovine serum, glutamine, and penicillin-streptomycin.
Plasmid construction, lentivirus production, and transduction into human cells
Twenty sgRNAs targeting the AAVS1 locus, nine sgRNAs targeting the AR gene, and six sgRNAs targeting the FOXA1 gene were designed and selected according to our prediction of sgRNA efficient scores. The sgRNA oligos were cloned into lentiCRISPRv2 plasmid (Addgene) as previously described (Shalem et al. 2014). Each plasmid containing inserted sgRNA sequence was verified using Sanger sequencing. To make lentivirus, the constructed lentiCRISPRv2 plasmids were cotransfected into 293T cells with packaging plasmids pMD2.G and psPAX2 (Addgene) using X-tremeGENE HP DNA Transfection Reagent (Roche) in 12-well plates according to the manufacturer's instructions. These 35 types of packaged lentivirus were then transduced into 293T and LNCaP-abl cells using 24-well plates, followed by puromycin selection for 3 d. The surviving cells were maintained for another 3 d before isolation of the DNA and proteins.
SURVEYOR nuclease assay and Western blot
The DNA fragments of the AASV1 locus were amplified from extracted genomic DNA by PCR using the Q5 high-fidelity DNA polymerases (NEB). SURVEYOR nuclease assays were subsequently performed according to the manufacturer's instructions (Trans genomic). The indel mutation rates were calculated as previously described (Ran et al. 2013). Western Blot for AR and FOXA1 knockout in LNCaP-abl cells by different sgRNAs was carried out using whole cell lysates as described previously (Xu et al. 2012), The antibody used for immunoprecipitation was anti-AR (N-20, Santa Cruz Biotechnology) and anti-FOXA1 (ab23738, Abcam), or anti-GAPDH (sc-25778, Santa Cruz Biotechnology). The images were analyzed using ImageJ (Schneider et al. 2012).
Software availability
A software package named Spacer Scoring for CRISPR (SSC) was developed for predicting sgRNA efficiency from genomic sequences. The SSC source codes are available in the Supplemental Material and at http://sourceforge.net/projects/spacerscoringcrispr/. A web-based application is available at http://crispr.dfci.harvard.edu/SSC.
Acknowledgments
We thank Drs. Carl Novina, Tim Wang, David Sabatini, Eric Lander, Ophir Shalem, and Neville Sanjana for providing the data sets and helpful discussions. The project was supported by the National Institute for Health Research (NIHR) grants U01 CA180980 (to X.S.L.), R01 GM113242-01 (to J.S.L.), National Science Foundation (NSF) grant DMS-1120368 (to J.S.L.), and the Claudia Adams Barr Award in Innovative Basic Cancer Research from the Dana-Farber Cancer Institute.
Author contributions: H.X. and X.S.L. designed the study. C.A.M., J.S.L., and M.B. provided conceptual advice. H.X., C.H.C., and D.W. developed the algorithm and performed the analyses. T.X. conducted experimental validations. W.L., T.X., L.C., F.Z., and M.B. helped with technical clarifications and result interpretations. H.X., T.X., and X.S.L. wrote the manuscript. All the authors participated in discussions and manuscript revision.
Notes
[1] Supplementary material [Supplemental material is available for this article.]
[2] Article published online before print. Article, supplemental material, and publication date are at http://www.genome.org/cgi/doi/10.1101/gr.191452.115.
References
- ↵Andersson BS, Collins VP, Kurzrock R, Larkin DW, Childs C, Ost A, Cork A, Trujillo JM, Freireich EJ, Siciliano MJ. 1995. KBM-7, a human myeloid leukemia cell line with double Philadelphia chromosomes lacking normal c-ABL and BCR transcripts. Leukemia 9: 2100–2108.
- ↵Berns K, Hijmans EM, Mullenders J, Brummelkamp TR, Velds A, Heimerikx M, Kerkhoven RM, Madiredjo M, Nijkamp W, Weigelt B, 2004. A large-scale RNAi screen in human cells identifies new components of the p53 pathway. Nature 428: 431–437.
- ↵Chen B, Gilbert LA, Cimini BA, Schnitzbauer J, Zhang W, Li GW, Park J, Blackburn EH, Weissman JS, Qi LS, 2013. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas9 system. Cell 155: 1479–1491.
- ↵Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, 2013. Multiplex genome engineering using CRISPR/Cas systems. Science 339: 819–823.
- ↵Doench JG, Hartenian E, Graham DB, Tothova Z, Hegde M, Smith I, Sullender M, Ebert BL, Xavier RJ, Root DE. 2014. Rational design of highly active sgRNAs for CRISPR-Cas9-mediated gene inactivation. Nat Biotechnol 32: 1262–1267.
- ↵Fu Y, Sander JD, Reyon D, Cascio VM, Joung JK. 2014. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat Biotechnol 32: 279–284.
- ↵Gilbert LA, Larson MH, Morsut L, Liu Z, Brar GA, Torres SE, Stern-Ginossar N, Brandman O, Whitehead EH, Doudna JA, 2013. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154: 442–451.
- ↵Gilbert LA, Horlbeck MA, Adamson B, Villalta JE, Chen Y, Whitehead EH, Guimaraes C, Panning B, Ploegh HL, Bassik MC, 2014. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159: 647–661.
- ↵Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V, Li Y, Fine EJ, Wu X, Shalem O, 2013. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31: 827–832.
- ↵Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. 2012. A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 337: 816–821.
- ↵Koike-Yusa H, Li Y, Tan EP, Velasco-Herrera MDC, Yusa K. 2014. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat Biotechnol 32: 267–273.
- ↵Konermann S, Brigham MD, Trevino AE, Joung J, Abudayyeh OO, Barcena C, Hsu PD, Habib N, Gootenberg JS, Nishimasu H, 2015. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517: 583–588.
- ↵Luo B, Cheung HW, Subramanian A, Sharifnia T, Okamoto M, Yang X, Hinkle G, Boehm JS, Beroukhim R, Weir BA, 2008. Highly parallel identification of essential genes in cancer cells. Proc Natl Acad Sci 105: 20380–20385.
- ↵Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM. 2013. RNA-guided human genome engineering via Cas9. Science 339: 823–826.
- ↵Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, Lim WA. 2013. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152: 1173–1183.
- ↵Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. 2013. Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8: 2281–2308.
- ↵Schneider CA, Rasband WS, Eliceiri KW. 2012. NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9: 671–675.
- ↵Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS, Heckl D, Ebert BL, Root DE, Doench JG, 2014. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343: 84–87.
- ↵Silver DP, Livingston DM. 2001. Self-excising retroviral vectors encoding the Cre recombinase overcome Cre-mediated cellular toxicity. Mol Cell 8: 233–243.
- ↵Sternberg SH, Redding S, Jinek M, Greene EC, Doudna JA. 2014. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature 507: 62–67.
- ↵Tibshirani R. 1996. Regression shrinkage and selection via the lasso. J R Stat Soc Series B 58: 267–288.
- ↵Tikhonov AN, Arsenin VY. 1977. Solutions of ill-posed problems. Winston, New York.
- ↵Tsai SQ, Zheng Z, Nguyen NT, Liebers M, Topkar VV, Thapar V, Wyvekens N, Khayter C, Iafrate AJ, Le LP, 2015. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat Biotechnol 33: 187–197.
- ↵Wang T, Wei JJ, Sabatini DM, Lander ES. 2014. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343: 80–84.
- ↵Wu X, Scott DA, Kriz AJ, Chiu AC, Hsu PD, Dadon DB, Cheng AW, Trevino AE, Konermann S, Chen S, 2014. Genome-wide binding of the CRISPR endonuclease Cas9 in mammalian cells. Nat Biotechnol 32: 670–676.
- ↵Xu K, Wu ZJ, Groner AC, He HH, Cai C, Lis RT, Wu X, Stack EC, Loda M, Liu T, 2012. EZH2 oncogenic activity in castration-resistant prostate cancer cells is Polycomb-independent. Science 338: 1465–1469.
- ↵Yang H, Wang H, Shivalila CS, Cheng AW, Shi L, Jaenisch R. 2013. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154: 1370–1379.
- ↵Zhou Y, Zhu S, Cai C, Yuan P, Li C, Huang Y, Wei W. 2014. High-throughput screening of a CRISPR/Cas9 library for functional genomics in human cells. Nature 509: 487–491.
- ↵Zou H, Hastie T. 2005. Regularization and variable selection via the elastic net. J R Stat Soc B 67: 301–320.

