Different Evolutionary Histories in Two Subgenomic Regions of the Major Histocompatibility Complex
Abstract
Two subgenomic regions within the major histocompatibility complex, the alpha and beta blocks, contain members of the multicopy gene families HLA class I, human endogenous retroviral sequence (HERV-16; previously known as P5 and PERB3), hemochromatosis candidate genes (HCG) (II, IV, VIII, IX), 3.8-1, and MIC (PERB11). In this study we show that the two blocks consist of imperfect duplicated segments, which contain linked members of the different gene families. The duplication and truncation sites of the segments are associated with retroelements. The retroelement sites appear to generate the imperfect duplications, insertions/deletions, and rearrangements, most likely via homologous recombination. Although the two blocks share several characteristics, they differ in the number and orientation of the duplicated segments. On the 62.1 haplotype, the alpha block consists of at least 10 duplicated segments that predominantly contain pseudogenes and gene fragments of the HLA class I and MIC (PERB11) gene families. In contrast, the beta block has two major duplications containing the genes HLA-B and HLA-C, and MICA(PERB11.1) and MICB(PERB11.2). Given the common origin between the blocks, we reconstructed the duplication history of the segments to understand the processes involved in producing the different organization in the two blocks. We then found that the beta block contains four distinct duplications from two separate events, whereas the alpha block is characterized by multisegment duplications. We will discuss these results in relation to the genetic content of the two blocks.
The major histocompatibility complex (MHC) has been described as a composite of conserved polymorphic blocks (Marshall et al. 1993). These blocks are ∼200–300 kb in length, and their combination is observed in a population as MHC haplotypes (Degli-Esposti et al. 1992). The central and telomeric region of the human MHC spans ∼2 Mb on the short arm of chromosome six and contains at least two blocks, the alpha and beta blocks (Marshall et al. 1993). The blocks contain members of the multicopy gene families HLA (human leukocyte antigen) class I (Geraghty et al. 1992); MIC (MHC classI-related chain) [alternatively termed PERB11(Perth beta block transcript11)] (Bahram et al. 1994; Leelayuwat et al. 1994); P5 (Vernet et al. 1993); PERB3 (Marshall et al. 1993); 3.8-1 (Pichon et al. 1997); hemochromatosis candidate genes (HCGs) II, IV, VIII, and IX (Pichon et al. 1996b); and 1AD3 (Totaro et al. 1995).
At least two of the multicopy gene families, HLA class I and MIC (PERB11), have an immunological role. The HLA class I family includes genes that encode cell surface glycoproteins that bind peptides and present them to cytotoxic T cells (Parham 1988). The function of MIC (PERB11) may involve γδ T cells and the protection of epithelial surfaces (Groh et al. 1998). Of the other multicopy gene families, P5 and PERB3 have been shown to form a genomic structure related to a human endogenous retroviral sequence-16 (HERV-16) sequence (Kulski and Dawkins 1999), HCGIX has similarity to transcription factors (Pichon et al. 1996c), and 1AD3 has no known transcripts.
The organization of the multicopy gene families in the alpha and beta blocks reflects a common origin and duplicative history (Leelayuwat et al. 1995; Pichon et al. 1996a). Within the beta block, we have recently shown that there are two large segmental duplications containing the loci HLA-B and HLA-C, and MICA(PERB11.1) and MICB(PERB11.2) (Gaudieri et al. 1997; Kulski et al. 1997). Based on L1 and Alu dating, the two duplications occurred as separate events resulting in four distinct segments (Kulski et al. 1997; Yamazaki et al. 1999). These duplications contain segments that share ∼30 kb of sequence, including coding and intergenic sequences. The major differences between the duplications are large insertions/deletions (indels) (>100 bp) containing retroelements [Alu, L1, HERV, MaLR (mammalianapparent LTR retrotransposon)], and processed pseudogenes and other repetitive sequences [mammalian wide interspersed repeat (MIR)/medium reiterated frequency repeat (MER)] (Kulski et al. 1997). The duplication and insertion sites of sequences within the duplications of the beta block are closely associated with retroelement sequences (Gaudieri et al. 1997; Kulski et al. 1997).
In contrast to the beta block, the alpha block contains a greater number of members of the multicopy gene families, particularly more pseudogenes and gene fragments of the HLA class I and MIC (PERB11) families (Geraghty et al. 1992; Leelayuwat et al. 1994). Within the alpha block, tandem gene duplications, gene conversion, and “block” duplications have been proposed to explain the complex arrangement of HLA class I loci resulting in the separation of phylogenetically related loci by more distantly related loci (Geraghty et al. 1992;Hughes 1995).
The purpose of this study was (1) to determine whether segmental duplications, as in the beta block, are involved in the organization of the multicopy gene families in the alpha block; (2) to determine the role of retroelements in the processes of duplication, truncation, and indels; and (3) to reconstruct the history of the duplications to understand the differing number of multicopy gene family members and complexity between the two blocks. To achieve these purposes we used recently released sequences spanning the alpha and beta blocks to compare the structural organization of the two blocks. We show that both blocks contain duplicated segments composed of linked multicopy gene families. The duplication and indel sites of the segments are associated with retroelements and other fragile sequences within the region and generate the dispersion and diversity of the multicopy gene families. However, the blocks differ in segment number, complexity, and orientation. The beta block contains four distinct segments from two separate duplication events, whereas the alpha block contains multisegment duplications. The multisegment duplications contain at least two tandem segments, one each from from the two major segment types in the alpha block. We will discuss the implications of the different duplication modes in relation to the genetic contents of each block.
RESULTS
Different Members of Multicopy Gene Families Are Linked Together as a Single Unit Within Duplicated Segments in Both Alpha and Beta Blocks
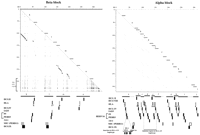
Figure 1 shows two dot plots of duplicated genomic segments within the alpha and beta blocks and the locations of the multicopy gene families HLA class I, 1AD3, P5 + PERB3 (HERV-16), 3.8-1, MIC (PERB11), and HCG (II, IV, VIII, and IX). The arrangement of the multicopy gene families and the lines within the dot plots indicate they are linked and form duplicated segments in both blocks, designated a–n. In this regard, segmental groups a–n depicted as homologous sequences by dot plots are defined by the genes and retroelements contained within the segment, as shown in Figure 2.
Segmental duplication of linked multicopy gene families. Dot-plot analysis of the intracomparison of the alpha (right) and beta (left) blocks. Diagonal lines within the dot plots indicate regions of duplication, with interruptions corresponding to indels. The position of the multicopy gene families HLA class I, HCG (II, IV, VIII, and IX), 1AD3, P5 + PERB3 (HERV-16), 3.8-1, and MIC (PERB11) are shown below the x-axis. Based on the duplication end points in the dot plot and the ordering of the multicopy gene families, duplicated segments have been designated a–n. In the alpha block, there is minimal distance between duplicated segments indicating that the duplication of the MIC (PERB11) members may have involved, though less likely, a neighboring segment. For the well-characterized HLA class I and MIC (PERB11) members, transcribed genes are indicated by shaded boxes, and pseudogenes and gene fragments are shown by open boxes. The HCG, 1AD3, and HERV-16 families are shown by black boxes. The endpoints of the human haplotype indels are indicated by horizontal dashed lines. Segment j is truncated at the 3′ end following anAlu Ya5/8 insert coinciding with the end of the alpha block sequence (accession no. AF055066). An Alu-rich region in the beta block is indicated.
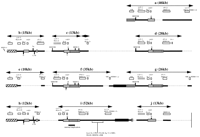
Truncations and indels of the alpha block segments associated with HERV-16 and other retroelement sequences. Schematic diagram of the retroelement content of the alpha block segments. The multicopy gene families are indicated above the bar by open boxes. Transcribed genes of the HLA class I and MIC (PERB11) family are indicated by shaded boxes. Retroelements are indicated by black vertical bars positioned above or below the horizontal boxes and bars to indicate orientation. The Alu retroelements are shown as their subtypes. Common clusters of retroelements are represented as follows: (hatched box) THE1C, L1PA, MER, and L1ME3A; (vertically lined bar) L1MB, MIR, MER20,Alu FRAM, LIM, and L2; (diagonally lined box) L1M4, MER21B, MSTB, and MSTA; (solid box) L1MB, L1M4, L1MB, FAM,Sx, Jb, MLT2D, L2, L1MB, and MER20; (dotted bar) L2, MLT1E, MLT, MLT1F, and LTR16B. (Dotted lines) Predicted indels. The duplicated segments are indicated by double arrows. The internal duplication within segment i, containing the retroelements from MLT to MER, is shown by a horizontal line. The end of the AF055066 sequence is indicated by a vertical line in segment j. For simplicity, the HCGIX sequences have not been included as they overlap the MIC (PERB11) members (Pichon et al. 1996c). HCG families II, IV, and VIII are represented by their respective roman numerals.
Comparison of the beta block against itself reveals two distinct duplications that are well separated and likely to be the result of single segment duplications. Each duplication set shares ∼30 kb of sequence. The two sets of duplications are in the same orientation but share limited sequence (Fig. 1). The segments k and l together contain HCGII, HLA class I, 1AD3, HERV-16, 3.8-1, MIC (PERB11), HCGIX, whereas segments m and n contain HCGII, HLA class I, and HCGIV.
Comparison of the alpha block against itself reveals a complex array of segment duplications (Fig. 1). All duplications are in the same orientation. There are no clear distinctions between the segments indicating little intervening sequence. The length of the segments varies between ∼10 and 52 kb. The alpha block segments a–j share HLA class I, HCGIV, 1AD3, and HERV-16. The duplications are imperfect and are interrupted with HCGII, HCGVIII, HCGIX, 3.8-1, and MIC (PERB11). The large insertion corresponding to the HLA-A31 haplotype contains a similar organization to the segments within the alpha block of the 62.1 haplotype (Fig. 1A). Although the segments within both blocks contain a similar order of the multicopy gene families, there are clear differences between the blocks that will be described in a later section.
Duplication, Truncation, and Indel Sites Associated with HERV-16 and Other Retroelement Sequences Within Duplicated Segments
The endpoints on several segments in both blocks, including the large indels corresponding to the HLA-A31 and HLA-A9 haplotypes, coincide with HERV-16 and other retroelement sequences (Figs. 1 and 2) (Kulski et al. 1999). Indel sites within the beta block have been shown to be associated with retroelement sequences (Gaudieri et al. 1997). Similarly, deletion sites in at least two segments in the alpha block are also associated with HERV-16 (Fig. 2). The insertion of retroelements THE1C, L1PA, MER, and L1ME3A is shared by several segments in the alpha block (Fig. 2). This insertion has occurred between an MLT and LTR16B (5′ end of HERV-16), and in segment i some of these retroelements form another internal duplication within the initial insertion (Fig. 2).
Other sites of truncations and rearrangements occur within the HLA class I and MIC (PERB11) family. Within the HLA class I family, the following loci contain only part of the full-length gene: HLA-80 and HLA-90 exon 3-8, HLA-16 exon 4-8, HLA-75 exon 1-3, HLA-17 exon 6-8, HLA-21 exon 3, and HLA-X intron C (Fig. 2) (Geraghty et al. 1992). These observations indicate a potential hot spot(s) between exons 2 and 4.
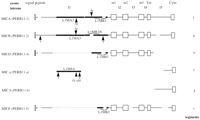
Figure 3 shows the MIC (PERB11) gene region in 6 of 14 segments from the alpha and beta blocks. The members differ from each other by large indels corresponding to L1 and Alusequences. MICD (PERB11.4) is truncated at the 3′ end in intron 5, which directly correlates with the start of MIC.b (PERB11.b). MIC.a (PERB11.a) commences further downstream of the MICD (PERB11.4) truncation in intron 5. In addition, the L1 sequence, L1MA4A, in MIC.a (PERB11.a) corresponds to a similar L1MA sequence within intron 1 of MICA (PERB11.1) and MICB(PERB11.2) (L1MA3). These L1 sequences are not present in MICD (PERB11.4) or MICE (PERB11.5). However, L1MB3 sequence is present in MICA (PERB11.1), MICD (PERB11.4), and MICE (PERB11.5), possibly in MICB(PERB11.2), but not in MIC.a (PERB11.a) and MIC.b (PERB11.b). The organization of MIC.a (PERB11.a) and MIC.b (PERB11.b) indicates introns 1 and 5 are fragile sites susceptible to change.
Indels/rearrangements within MIC (PERB11) members. A schematic diagram of the MIC (PERB11) gene family. The intron and exon structure is depicted on MICA (PERB11.1). Exons are shown as open boxes and labeled as signal peptide, α1, α2, α3, Tm (transmembrane region), and cyto (cytoplasmic region). The introns are shown as lines and labeled in order as I1–I5. The positions of gaps in MIC (PERB11) members are indicated by a dashed line. L1 sequences are depicted by solid bars. The positions of Alu sequences are shown by arrowheads. The signal peptide of MICD (PERB11.4) is shown by a grey box, as the level of identity with other members is low in this region and the corresponding open reading frame does not encode hydrophobic amino acids as found in the signal peptide of other members. MICA (PERB11.1) andMICB (PERB11.2) are full-length genes transcribed in the beta block. MICD (PERB11.4) and MICE (PERB11.5) are pseudogenes, and MIC.a (PERB11.a) and MIC.b (PERB11.b) are fragments containing the 3′ end of the gene.
The association of retroelement sequences with duplication, truncation, and indel sites is a common feature of both blocks. However, significant differences between the blocks exist, which will be described in the next section.
Different Orientation and Complexity of Segmental Duplications in the Alpha and Beta Blocks
Comparison of the alpha and beta blocks shows the following differences:
- 1.
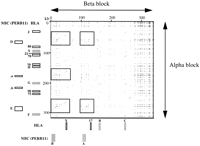
- There is opposite orientation of duplicated segments corresponding to segments containing both MIC (PERB11) and HLA class I members. The sequence surrounding HLA-B and HLA-C in the beta block does not form large segment duplications with the class I region of the alpha block; only the HLA class I gene or pseudogene and limited surrounding sequence is retained (∼5–10 kb) (Fig.4).
- 2.
- The alpha block, on the 62.1 haplotype, contains at least 10 segments compared with the four segments within the beta block.
- 3.
- The percentage of paralogous retroelements, which is correlated to the number of duplicated segments, is greater in the alpha block (Table1). The overall percentage of retroelements within the beta block is greater than the alpha block (Table 1), although this is predominantly from an increase in Alu sequences near the 3′ end of the beta block (Fig. 1).
- 4.
- The ratio of pseudogene and gene fragments to functional genes of the HLA class I and MIC (PERB11) families is greater in the alpha block.
- 5.
- Large indels occur in some human haplotypes within the alpha block, indicating recent genomic changes.
Different orientation and complexity of segments in the alpha and beta blocks. Intercomparison of the alpha and beta blocks. The orientation of the duplications shown by the small diagonal lines indicates that the segments from the two blocks are in opposite orientation to each other. Large segment duplications corresponding to segments containing MIC (PERB11) and HLA class I members are shown by boxes. For the well-characterized HLA class I and MIC (PERB11) members, transcribed genes are indicated by grey boxes, and pseudogenes and gene fragments are shown by open boxes along the x- and y-axes.
Different Properties of the Alpha and Beta Blocks
Table 1 lists the properties of the alpha and beta blocks. Points 2, 3, and 4 above indicate the complexity of the segment duplications is a major difference between the two blocks. This is particularly interesting given the common origin of the two blocks. Within the beta block, individual segment duplications have been described (Gaudieri et al. 1997; Kulski et al. 1997); however, the mode of duplication within the alpha block appears to be more complex.
Reconstruction of the Duplication History Shows Multisegment Duplications and Translocations Occur Within the Alpha Block
To determine the complex arrangement of duplications in the alpha block, we examined the relationship between the segments a–j, using phylogenetic analysis (Fig. 5) and the level of identity between paralogous L1 retroelement sequences (Table 2). The results show that sequences spanning HCGII to HERV-16 on the duplicated segments (Fig. 2) share a common phylogenetic organization (Fig. 5), supporting the duplication of linked multicopy gene families.
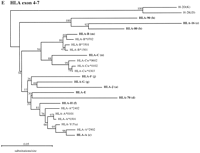
The alpha block segments can be separated into two main groups. The phylogenetic relationships of segments a–n were examined by analyzing the multicopy gene families of 1AD3 (A), HCGIV (B), HCGII (C), HERV-16 (D), and HLA class I (E). A scale bar for the number of substitutions per site is included for each tree and the bootstrap percentage from 1000 replications is given for each node. All trees except the HLA class I tree are unrooted. In the HLA class I tree (E), mouse H2 sequences were used as an outgroup and segments corresponding to HLA class I loci are indicated by brackets.
Nucleotide Identity (%) Between Paralogous L1 Sequences Within the Alpha and Beta Block Segments
Figure 5 shows that segments b, h, and e (containing HLA-80, HLA-90, and HLA-16, respectively), separate from the other segments within the alpha block, with high bootstrap values and form a separate lineage in agreement with previous results for HLA-80, HLA-90, and HLA-16 (Hughes 1995). Nucleotide comparison of paralogous L1 sequences between the alpha block segments shows segments b and h share 90.3% identity but only 76.3% with the other alpha block segments (Table 2). In addition, the segments b, e, and h share a distinct set of retroelements at the 5′ end corresponding to the region containing HCGII, which separates them from the other alpha block segments, which lack both HCGII and the retroelements (Fig. 2). There are two main groups of segments in the alpha block that must have diverged early in the evolution of the segments.
The segments a, d, g, and i, which harbor HLA-J, HLA-70,HLA-G, and HLA-75, respectively, do not have distinct clustering patterns in the different phylogenetic trees (Fig. 5A,B,D,E) indicating similar levels of identity, supported by the comparison of paralogous L1 sequences (88%–91%) (Table 2). Similarly, the segments b, h, and e share 90.3% identity between paralogous L1 sequences. The position of the two major groups of segments in the alpha block and equivalent levels of identity between paralogous L1 sequences suggest at least two instances of bisegmental duplications involving a member from each group (i.e., a bisegmental duplication of segments h and i, and segments e and f) (Fig. 2). In addition, the sequence content between segments f and g is similar to that between segments i and j, suggesting bi- or trisegmental duplications within this block (Fig. 2). A more detailed analysis of the evolution of the segments in the alpha and beta blocks based on retroelement dating will be presented elsewhere (Kulski et al. 1999). The complex history of the segments in the alpha block differs from the single segment duplications present in the beta block.
Furthermore, the segments f (containing HLA-H) and c (containingHLA-A) form a cluster (Fig. 5A,B,D,E) with high reliability. The level of identity between paralogous L1 sequences in segments f and c is ∼94.1%, greater than any other comparison, supporting a recent common ancestor (Table 2). Previous phylogenetic results ofHLA-A and HLA-H showed they had diverged recently (Hughes 1995). The segments c and f can be further separated from the other alpha block segments by the insertion of a MER9 and Alu Yinsert (Fig. 2). To account for the large distance between the two segments, a likely translocation event has occurred subsequent to duplication. The alpha block segment history includes single and multisegment duplications as well as translocations. In contrast, individual segment duplications have been described in the beta block (Gaudieri et al. 1997; Kulski et al. 1997).
DISCUSSION
Common Features of the Alpha and Beta Blocks
The alpha and beta blocks are characterized by the duplication of linked multicopy gene families. We have shown that both blocks contain imperfect segmental duplications resulting in the dispersion and diversity of the families and associated retroelements. An important consequence of the duplications is the increased possibility of recombination between paralogous retroelements and other repetitive sequences resulting in genomic changes (Mazzarella and Schlessinger 1997).
Ancestral Segment Contained Linked Members of the HLA Class I, HERV-16, and MIC (PERB11) Gene Families
The composition of the segments in the alpha and beta blocks suggests that the ancestral HLA class I gene was linked to other multicopy gene families through retroelements such as HERV-16. The ancestral segment of the alpha and beta blocks contained linked members of the HLA class I, HCGIV, HERV-16, 3.8-1, and MIC (PERB11)/HCGIX gene families and the associated retroelements L2, MLT1E, MLT, MLT1F, and LTR16B (Figs. 1 and 2). Although the functions of HCGIV, HERV-16, and 3.8-1 are unknown, the HLA class I and MIC (PERB11) genes are involved in antigen presentation to T cells and their linkage may be of functional importance.
The distinct phylogenetic cluster of the segments b, h, and e away from the other segments within both blocks (Fig. 5E) suggests that the ancestral segment of b, h, and e was the result of an imperfect duplication at HERV-16, or the ancestral segment did not extend beyond HERV-16. In the HLA class I exon 4–7 tree (Fig. 5E), segments m and n containing HLA-B and HLA-C, respectively, clustered between the two major groups of alpha block segments, suggesting that the ancestral segment of m and n diverged from the alpha block segments after the separation of the two main groups of segments. Therefore, the ancestral segment of m and n was imperfect, resulting in limited common sequence with other segments.
Homologous Recombination Between Retroelements Is an Important Process Within the MHC
The segment duplications, truncations, indels, and internal duplications coincide with regions associated with retroelements, particularly HERV-16. These results indicate that homologous recombination between retroelements is an important mechanism of such changes.
There is increasing evidence that retroelements (including HERVs) contribute to the recombination and duplications observed in the MHC (Kulski et al. 1997; Andersson et al. 1998) and in other gene clusters in the genome (Erickson et al. 1992; Schwartz et al. 1998). Furthermore, instability of operon structures in bacteria has been shown to be associated with transposon insertion sequences (Itoh et al. 1999).
Differences Between Alpha and Beta Blocks
Although the two blocks in this study shared a common origin, the duplicative history is different in their overall genomic organization. The blocks differ in the number of duplicated segments and in the proportion of pseudogenes and gene fragments. The alpha block contains a greater proportion of paralogous retroelements (Table 1) resulting in a greater level of identity between the sequences, and therefore, an increase in the possibility of exchange and indel events within the block. The process of imperfect duplications and the subsequent indels and rearrangements results in a large number of nonfunctional sequences. The alpha block appears to have accumulated nonfunctional sequences. The functional HLA-A, HLA-G, andHLA-F sequences in the alpha block have not been disrupted, however, the importance of these three loci has been questioned (Klein et al. 1998).
The differences between the two blocks are a result of the mode of segment duplications, which in turn have been generated by retroelements. The blocks differ either through an intrinsic property of the genomic sequence within the blocks, or more likely, through chance events whereby duplicated segments have provided a greater opportunity for homologous recombination between retroelements, with the result of additional gene copies. The formation of noncoding sequences of the HLA class I and MIC (PERB11) gene families within the alpha block (particularly early in the duplicative history through the formation of the ancestral segment of b, h, and e) may have allowed the propagation of the segment duplications in the alpha block, resulting in the present complex organization. The arrangement of segments in the alpha block appears to have occurred by several duplications of more than one segment, whereas two separate duplication events have occurred in the beta block. Therefore, the complex segment history in the alpha block is different from the history of the beta block segments that have arisen via single segment duplications.
METHODS
DNA Sequences
A 319-kb sequence within the alpha block spanning 32 kb centromeric of HLA-J to 6 kb telomeric of HLA-F was obtained from the DDBJ/EMBL/GenBank accession no. AF055066. A 337-kb sequence within the beta block spanning MICB (PERB11.2) to 87 kb telomeric of HLA-C was obtained from DDBJ/EMBL/GenBank accession nos. AB000878, AB000879, AB000880, and D84394. AF055066,AB000878, AB000879, and AB000880 are from the cell line BOLETH and contain the HLA alleles HLA-A2, HLA-B62, HLA-Cw10, and HLA-DR4 (62.1 haplotype). D84394 is from the heterozygous cell line CGM1 with the HLA allelesHLA-A3,29; HLA-B8,14; HLA-Cw-,-; and HLA-DR3,7.
The HLA-A, HLA-B, and HLA-C alleles used for the phylogenetic tree in Figure 5E were obtained from the World Wide Web (http://square.umin.ac.jp/JSHI/hla_data/data.html). The sequences for HLA-Y(YU) (accession no. AB012685), HLA-E(accession no. M20022), H-2D (K) (accession no. U47327), and H-2K (D) (accession no. U47329) were obtained from the DDBJ/EMBL/GenBank database.
Location of Multicopy Gene Families, Retroelements, and Duplications
The locations of the HLA class I, HERV-16 (P5 + PERB3), MIC (PERB11), 1AD3, 3.8-1, and HCGII, HCGIV, HCGVIII, and HCGIX family members on the sequences were determined using the program BLASTN (National Center for Biotechnology Information, Bethesda, MD) and the annotation information of the database entry. The position and type of retroelement within each sequence was determined using the RepeatMasker program (http://ftp.genome.washington.edu/cgi-bin/RepeatMasker). The inter- and intracomparison of the sequences was performed using the dot matrix program Dotter (Sonnhammer and Durbin 1996).
Phylogenetic Analysis
The members of the HLA class I, HERV-16 (P5 + PERB3), 1AD3, HCGII, and HCGIV families were each aligned using the CLUSTALW program (GCG, Madison, WI). The resultant alignments were used in the construction of the phylogenetic trees by the neighbor-joining method (Saitou and Nei 1987) based on Kimura’s two-parameter model (Kimura 1981). To assess the reliability of the clusters, 1000 bootstrap replications were performed (Felsenstein 1985).
Acknowledgments
We thank Dr. Naoko Takezaki, Professor Yoshio Tateno, Dr. Kazuho Ikeo, Dr. Annalise Martin, Sonia Cattley, and two anonymous reviewers for helpful comments regarding the manuscript. S.G. is supported by a Japanese Society for the Promotion of Science fellowship (97119). J.K.K. and R.L.D. are supported by the National Health and Medical Research Council.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
Footnotes
-
↵3 Corresponding author.
-
E-MAIL sgaudier{at}lab.nig.ac.jp; FAX 81-559-81-6848.
-
- Received December 15, 1998.
- Accepted May 10, 1999.
- Cold Spring Harbor Laboratory Press