Targeted delivery of CRISPR interference system against Fabp4 to white adipocytes ameliorates obesity, inflammation, hepatic steatosis, and insulin resistance
- 1Department of Bioengineering, Institute for Bioengineering and Biopharamceutical Research Hanyang University, 04763 Seoul, South Korea;
- 2BK21 Plus Future Biopharmaceutical Human Resource Team, Hanyang University, 04763 Seoul, South Korea
-
↵3 These authors contributed equally to this work.
Abstract
Obesity is an increasing pathophysiological problem in developed societies. Despite all major progress in understanding molecular mechanisms of obesity, currently available anti-obesity drugs have shown limited efficacy with severe side effects. CRISPR interference (CRISPRi) mechanism based on catalytically dead Cas9 (dCas9) and single guide RNA (sgRNA) was combined with a targeted nonviral gene delivery system to treat obesity and obesity-induced type 2 diabetes. A fusion peptide targeting a vascular and cellular marker of adipose tissue, prohibitin, was developed by conjugation of adipocyte targeting sequence (CKGGRAKDC) to 9-mer arginine (ATS-9R). (dCas9/sgFabp4) + ATS-9R oligoplexes showed effective condensation and selective delivery into mature adipocytes. Targeted delivery of the CRISPRi system against Fabp4 to white adipocytes by ATS-9R induced effective silencing of Fabp4, resulting in reduction of body weight and inflammation and restoration of hepatic steatosis in obese mice. This RNA-guided DNA recognition platform provides a simple and safe approach to regress and treat obesity and obesity-induced metabolic syndromes.
Obesity is an increasing pathological and social problem worldwide (Lissner et al. 2010; Ouchi et al. 2011). The prevalence of childhood obesity has increased threefold during the past decades. The pathophysiology underlying obesity is not evident, but the interaction between genetics and obesogenic environmental factors related to weight gain with the increased availability of energy-dense food and reduced energy expenditure plays a major role, resulting in an excess amount of fat or lipid accumulation in both visceral and subcutaneous white adipose tissues (Smorlesi et al. 2012). White adipose tissues store energy in the form of triglycerides and secrete fatty acids and various kinds of pro-inflammatory cytokines, making it a primary therapeutic target for the treatment of obesity and obesity-induced metabolic syndromes. Anti-obesity drugs under development have been directed toward restriction of caloric intake by acting on the gastrointestinal tract or the central nervous system. However, most of these drugs have shown little efficacy, accompanied by severe side effects (Elangbam 2009). The low therapeutic effectiveness of anti-obesity drugs might result from their off-targeting effects toward other tissue types; an adipose tissue-targeted drug delivery system is necessary for highly effective anti-obesity drugs (Watanabe et al. 2003). One advantage of gene therapy in the treatment of various diseases is its ability to passively or actively target the organs-of-interest (Hrkach et al. 2012), and it has been considered a promising therapeutic approach for the treatment of obesity (Rehman et al. 2018; Tasyurek et al. 2018). However, this approach has been hampered by the inability to target adipose tissues (Stenkula et al. 2007).
Diverse methods regarding gene therapy have been studied. One common method, viral vector delivery, shows uncontrolled long-term expression of the therapeutic gene and immunogenicity, thereby resulting in severe side effects. The use of nonviral gene carriers provides a safer approach in terms of nonimmunogenic response and controlled gene expression (Song et al. 2016). The specific binding of a short targeting peptide, CKGGRAKDC, to prohibitin and its internalization into vascular endothelial cells located on the surface of mature adipocytes has been reported (Kolonin et al. 2004; Mishra et al. 2005). Prohibitin is expressed ubiquitously on adipose tissue vasculature and plays a vital role in cell cycle regulation and homeostatic mitochondrial function. It is mostly present in the inner mitochondrial membrane and cytoplasm but shifts its location to the cell membrane and nucleus in differentiated adipocyte cells.
The development of genome editing technology, including the clustered regularly interspaced short palindromic repeats system (CRISPR), has made it possible to perform precise genetic engineering. Genetic reprogramming of cells requires precise and predictable methods for expression or repression of target sets of genes. RNA interference (RNAi) and engineered DNA-binding proteins such as zinc finger nuclease or transcription activator-like effector (TALE) proteins have emerged as powerful genetic modification tools (Hannon 2002). RNAi has the potential for knockdown expression of targeted genes; however, it is limited to particular organisms and suffers from significant off-target effects, immunogenicity, and RNA stability. On the other hand, DNA-binding proteins such as zinc finger nucleases and TALE proteins have the potential for precise genome editing, although they remain somewhat difficult and are expensive to design and develop (Cox et al. 2015). Therefore, the simultaneous modification of multiple genes remains a primary challenge. The CRISPR system provides a platform for targeted genetic regulation in a variety of cell types and organisms (Mali et al. 2013). Diverse CRISPR systems are present in different microorganisms; the type II CRISPR system derived from Streptococcus pyogenes has been reprogrammed to engineer mammalian genomes to induce double-strand breaks, requiring a single gene encoding Cas9 protein and a single guide RNA (sgRNA) consisting of a hairpin mimicking the tracrRNA-crRNA complex (Shalem et al. 2014; Wang et al. 2014). Several types of genome editing using the CRISPR system and genome modification using a catalytically inactive version of Cas9 for RNA-guided transcription regulations, known as CRISPR interference (CRISPRi), have been studied (Qi et al. 2013). The nuclease-deactivated Cas9 (dCas9), when co-expressed with sgRNA, generates a DNA recognition complex that can specifically interfere with transcription elongation, RNA polymerase binding, or transcription factor binding without altering target sequences (Larson et al. 2013).
In our previous study, silencing of the lipid chaperone gene, fatty acid binding protein 4, adipocyte (Fabp4), was proven to be an effective strategy for metabolic recovery and body weight loss in high-fat diet-induced mice (HFD-induced mice) (Won et al. 2014). Recent clinical studies have shown the importance of obesity-related vaccination and genetic deletion of Fabp4 for the treatment of obesity-induced inflammation (Sheridan et al. 2012; Steen et al. 2017). Despite this clinical significance, few therapeutic drugs have been developed for obesity and obesity-induced inflammation, therefore requiring a new therapeutic approach to treatment of obesity and obesity-induced inflammation (Furuhashi et al. 2014). A short peptide, adipose targeting sequence combined with the D-form of 9-mer arginine (ATS-9R) showed white adipose tissue specificity in vivo, and ATS-9R gene complexes were highly distributed in stromal fractions (Won et al. 2014). Various studies have proven the successful delivery of the CRISPR/Cas9 system in vivo (Ablain et al. 2015; Jiang et al. 2017; Miller et al. 2017), and to this end, carrier systems that can deliver plasmid DNA encoding Cas9, Cas9 mRNA together with sgRNA, or direct delivery of Cas9 protein/sgRNA complexes have been investigated (Mout et al. 2017; Yue et al. 2018). Although they show effective genome editing, they require more work compared to delivery using plasmids and they are more favorable in terms of high stability and effective genome editing (Jinek et al. 2013). Nonviral delivery vehicles, such as peptides, have been exploited as alternative carriers to deliver Cas9/sgRNA plasmids. However, they could also work in nontarget cells, resulting in potential nonspecific gene modification in vivo. Given these reports, we hypothesize that the use of a nonviral prohibitin-targeting gene delivery system to direct the CRISPRi system consisting of dead Cas9 and Fabp4 sgRNA to adipose tissues can be an efficient and potential strategy for treating obesity and obesity-induced metabolic syndromes.
Results
Adipocyte-targeted nonviral CRISPR interference system
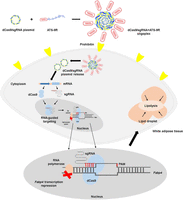
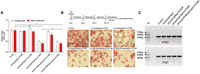
A CRISPR interference system based on catalytically dead Cas9 lacking endonuclease activity and single guide RNA was targeted to white adipocytes with a nonviral gene delivery carrier consisting of adipose tissue targeting sequence (ATS) peptide (CKGGRAKDC) conjugated to D-form of 9-mer arginine (9R). The peptide has been reported to bind specifically to white adipose tissue through membrane protein prohibitin (Fig. 1; Won et al. 2014). Four site-specific sgRNAs against Fabp4 were constructed (Supplemental Table S1), and sgRNA #4 was selected for further experiments as it showed enhanced Fabp4 repression as compared to sgRNA #1 and sgNegative RNA (Fig. 2A). Antilipogenesis effects of oligoplexes system were confirmed by Oil Red O staining, and the number of lipid droplets significantly decreased following treatment with the (dCas9/sgFabp4) + ATS-9R oligoplexes (Fig. 2B). To show the null activity of the dCas9/sgFabp4, dCas9-KRAB/sgFabp4 system, the T7E1 assay was performed (Fig. 2C, Supplemental Fig. S8) demonstrating the null activity of the dCas9/sgFabp4 system in mature adipocytes.
Schematic illustration representing nonviral CRISPR interference system delivery to white adipocytes. The dCas9 plasmid and sgRNA against Fabp4 gene formed complexes with ATS-9R peptide via electrostatic interaction. The highly cationic ATS-9R functions as a targeting and condensing peptide to deliver dCas9/sgRNA to white adipose tissues, and dCas9 plasmid contains nuclear localization signals (NLSs) on both the N-terminus and C-terminus for delivery to the nucleus.
Validation of (dCas9/sgFabp4) + ATS-9R and (dCas9-KRAB/sgFabp4) + ATS-9R oligoplexes in adipocytes. (A) dCas9/sgNeg, dCas9/sgFabp4 #1,#4, dCas9-KRAB/sgFabp4 #1,#4 were complexed with ATS-9R in pre-adipocytes and mature adipocytes and transfected under optimized conditions. The Fabp4 mRNA level was normalized and calculated as a value relative to the nontreated cells. (B) Mature adipocytes were treated as mentioned above, and the lipid droplets were assessed by Oil Red O staining to evaluate the antilipogenesis effect through CRISPRi-mediated delivery of dCas9 and sgFabp4. (C) Agarose gel image showing the CRISPR interference repression activity measured by T7E1 assay after transfection of dCas9/sgFabp4, (dCas9/sgNegative) + ATS-9R, (dCas9/sgFabp4) + ATS-9R, and (dCas9-KRAB/sgFabp4) + ATS-9R in mature adipocytes. (M) Molecular weight size marker.
Characterization of (dCas9/sgFabp4) + ATS-9R oligoplexes
Various studies have shown successful delivery of Cas9 protein, Cas9 mRNA, or direct delivery of Cas9 together with sgRNA (Mout et al. 2017; Yue et al. 2018), but they require a complicated process and show low gene editing. Therefore, plasmid delivery is more favorable in terms of high stability and efficient gene editing, and nonviral vectors have gained much attention for plasmid delivery. To evaluate the condensing ability of dCas9 and sgRNA plasmids with ATS-9R, gel retardation studies were performed and no DNA migration was observed at or above a weight ratio of 3, suggesting complete condensation (Supplemental Fig. S1A). The (dCas9/sgFabp4) + ATS-9R oligoplexes revealed round morphologies with a diameter of 160 nm in transmission electron microscopic images (Supplemental Fig. S1B). Dynamic light scattering analysis was performed to observe the sizes and zeta potentials of the oligoplexes. The mean diameter of the (dCas9/sgFabp4) + ATS-9R oligoplexes was ∼200 nm above weight ratio 3 (Supplemental Fig. S1C). The overall positive charge on the surface of the oligoplexes facilitates cellular uptake via interaction with the negatively charged cell membranes, and zeta potentials were also measured and showed an overall positive charge above weight ratio 1 (Supplemental Fig. S1D). To characterize the transfection efficiencies, the oligoplexes were analyzed by luciferase assay and flow cytometry analysis. ATS-9R oligopeptide complex with luciferase DNA plasmid was transfected into mature adipocytes. The transfection efficiency gradually increased with an increased amount of ATS-9R compared with naked DNA and nontransfected cells (Supplemental Fig. S2A). Expression levels observed by luciferase assay of ATS-9R oligoplexes showed the highest expression, confirming the delivery ability of ATS-9R to adipocytes (Supplemental Fig. S2B). Differentiated adipocytes presented over 80% cell viability up to a weight ratio of 5, indicating that the oligoplexes are nontoxic (Supplemental Fig. S2C). The percentage of cells transfected with (dCas9/sgFabp4) + ATS-9R oligoplexes was 99% as determined by flow cytometry (Supplemental Fig. S2D). These results further prove that ATS-9R shows the ability to target adipose tissues, and to further allow intracellular visualization of oligoplexes at appropriate weight ratios, 3T3-L1 pre-adipocytes were treated with Cy5.5-labeled DNA complexes and fluorescein isothiocyanate (FITC)-labeled ATS-9R in a time-dependent manner. The DNA and carrier were well distributed in the cytoplasm with strong fluorescence intensity after 24 h (Supplemental Fig. S3A). Mature adipocytes were also treated with FITC-conjugated ATS-9R, and CellMask Deep Red was used to stain the plasma membrane. In mature adipocytes, earlier and higher FITC fluorescence was observed as compared to pre-adipocytes, indicating the early binding of ATS-9R with enhanced prohibitin expression in mature adipocytes (Supplemental Fig. S3B).
Fabp4 silencing mediated by (dCas9/sgFabp4) + ATS-9R oligoplex in adipocytes
To test gene silencing and antilipogenesis effect of the (dCas9/sgFabp4) + ATS-9R, 3T3-L1 pre-adipocytes and mature adipocytes were treated with oligoplexes. While the (dCas9/sgFabp4) + ATS-9R oligoplexes-treated groups silenced Fabp4 mRNA expression by 30% in pre-adipocytes, the silencing percentage increased up to 60% in mature adipocytes (Supplemental Fig. S4A). These results further confirmed the relationship between prohibitin expression and ATS-9R binding. The antilipogenesis effects of oligoplexes system were confirmed by Oil Red O staining, and the number of stained lipid droplets significantly decreased following treatment with the (dCas9/sgFabp4) + ATS-9R oligoplexes (Supplemental Fig. S4B). Recent studies have shown that a catalytic mutant dCas9 can mediate gene editing by modifying the transcription of genes without altering the target sequence (Gilbert et al. 2014; Zuris et al. 2015). To show the null activity of the dCas9/sgFabp4 system, the T7E1 assay was performed (Fig. 5A; Supplemental Figs. S4C, S10) demonstrating the null activity of the dCas9/sgFabp4 system in the treated cells. A recent study showed an enhanced CRISPR repressor system by addition of a repressor domain KRAB to nuclease-dead Cas9, dCas9-KRAB proving enhanced repression (Yeo et al. 2018). To compare the gene silencing and antilipogenesis effect of (dCas9/sgFabp4) and (dCas9-KRAB/sgFabp4) with ATS-9R, the oligoplexes were treated in pre- and mature adipocytes and Fabp4 mRNA levels were detected (Fig. 2A). However, from the results, (dCas-KRAB/sgFabp4) + ATS-9R-treated groups did not achieve a stronger silencing effect of Fabp4 mRNA as compared to (dCas9/sgFabp4) + ATS-9R-treated groups. Oil Red O staining showed no difference of antilipogenesis effect between (dCas9/sgFabp4) + ATS-9R and (dCas9-KRAB/sgFabp4) + ATS-9R oligoplexes; therefore, for further experiments (dCas9/sgFabp4) was selected.
Fabp4 silencing by (dCas9/sgFabp4) + ATS-9R oligoplex in epididymal adipose tissues
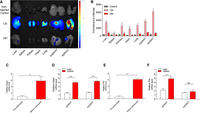
To confirm the biodistribution of the CRISPRi system in a high-fat diet-induced type 2 diabetic obese tissues, (dCas9/sgFabp4) + ATS-9R oligoplexes were intraperitoneally injected into HFD-induced mice. At 12 and 24 h postinjection, the oligoplexes were found to be distributed mostly in epididymal white adipose tissue (epiWAT), followed by subcutaneous white adipose tissue (subWAT) and liver (Fig. 3A). Fluorescence was also detected in the subWAT with much lower intensity. Analysis of relative total fluorescence intensities showed the highest localization at 12 h in epiWAT, proving specific targeting to white adipose tissue via prohibitin binding (Fig. 3B). The uptake by the liver is due to the clearance of the injected nanoparticle (Tsoi et al. 2016). To further confirm Fabp4 and Phb (prohibitin) expression, the mRNA levels in 3T3-L1 pre-adipocyte, mature adipocytes, and lean and obese mice were verified (Fig. 3C,D). Significant increases were observed in both Fabp4 and Phb levels in mature adipocytes by ∼5× and ∼5.8×, respectively, compared to those in the pre-adipocytes. The Fabp4 mRNA levels in the epiWAT and subWAT of the obese mice were approximately 4× to 5× higher than those of nonobese mice (Fig. 3E). A higher level of Phb gene expression was also confirmed in epiWAT of obese mice (Fig. 3F), while there was no difference in subcutaneous white adipose tissue between normal and obese mice. These results represent that high fatty acid binding protein 4, adipocyte expression is a crucial therapeutic biomarker of high fat-induced obesity and metabolic syndrome, and high prohibitin expression in adipocytes and adipose tissues is a key factor necessary for targeted gene delivery.
Epididymal adipose tissue targeting of (dCas9/sgFabp4) + ATS-9R oligoplexes. (A) Biodistribution image of intraperitoneally injected (dCas9/sgFabp4) + ATS-9R oligoplexes at 12 and 24 h postinjection. (B) Mean fluorescence intensity in various organs. Adipocytes were differentiated into mature adipocytes for in vitro analysis. For in vivo studies, 6-wk-old male C57BL/6 mice were fed on standard chow or a high-fat diet for 11 wk. Total RNAs were extracted and relative mRNA levels were determined by real-time PCR. Fabp4 mRNA was increased in the mature adipocytes (C) as well as both in the epidydimal white adipose tissue (epiWAT) and subcutaneous white adipose tissue (subWAT) of obese mice (D). Phb mRNA levels also increased in (E) mature adipocytes and (F) obese epiWAT. Data are presented as mean ± SEM. (n = 4–5). (ns) nonsignificant, (**) P < 0.01, (***) P < 0.001 was considered significant.
(dCas/sgFabp4) + ATS-9R oligoplexes alleviate obesity and insulin resistance in an obesity-induced diabetic model
Therapeutic gene editing has a broad potential for treating diverse diseases through correction of genetic defects. The in vivo anti-obesity therapeutic effect by the CRISPR interference system was determined in a HFD-induced obesity model (Supplemental Fig. S5). The mice were treated through intraperitoneal injection twice a week for up to 6 wk, and the mean body weights decreased by ∼20%. The 9R groups showed less decrease in body weight (Fig. 4A,B); the slight decrease was due to the transduction ability of 9R (Kim et al. 2006). Excessive fat accumulation, lipolysis, and adipokines attract immune cells such as monocytes, resulting in accumulation of macrophages, triggering cytokine release (Weisberg et al. 2003; Berg and Scherer 2005). These cytokines stimulate AKT signaling, leading to insulin resistance and type II diabetes. In addition to the confirmation of decrease in body weight by CRISPR interference, the insulin tolerance test revealed lowered blood glucose levels following repeated treatment of (dCas9/sgFabp4) + ATS-9R oligoplexes (Fig. 4C). No significant changes in food intake were recorded during the treatment period, indicating that the loss of body weight is not due to eating less (Fig. 4D). It was concluded that the CRISPR interference system selectively targeting white adipocyte and silencing overexpressed Fabp4 levels is effective and safe in decreasing body weight and recovering insulin resistance in obese mice. To further confirm the specific gene delivery and silencing effect to epiWAT, (dCas9/sgFabp4) + 9R and (dCas9/sgFabp4) + ATS-9R oligoplexes were intraperitoneally injected into mice and total RNA was isolated from epiWAT and subWAT for expression analysis. Fabp4 mRNA was significantly down-regulated in epiWAT (Fig. 5A; Supplemental Fig. S9A) as compared to subWAT (Fig. 5B; Supplemental Fig. S9B), demonstrating specific gene silencing in epiWAT, and the T7E1 assay demonstrated CRISPRi mediated Fabp4 silencing. These results demonstrate that silencing of Fabp4 alleviates obesity and insulin resistance in the high-fat diet-induced type 2 diabetic model.
In vivo therapeutic effects of the CRISPRi system in a high-fat diet-induced mouse model. (A) Weekly body weight. (B) Fat masses for epiWAT, subWAT, and total white adipose tissue. (C) Insulin tolerance test (ITT) after 6 wk of treatment. Insulin was injected intraperitoneally and blood glucose was detected at each time point, and AUC was determined. (D) Weekly food intake. Data are presented as mean ± SEM. (n = 5). (ns) nonsignificant, (*) P < 0.05, (**) P < 0.01, (***) P < 0.001 was considered significant.
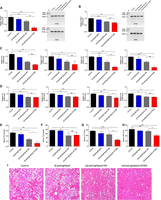
The (dCas9/sgFabp4) + ATS-9R oligoplexes ameliorate inflammation and hepatic steatosis in obese mice. Data are presented as relative mRNA levels in (A) epiWAT Fabp4, (B) subWAT Fabp4, (C) epiWAT cytokines, and (D) subWAT cytokines. Serum levels of (E) FFA, (F) TG, (G) ALT, and (H) AST. (I) Representative hematoxylin and eosin sections of liver tissue. Error bars indicate SEM (n = 5) and scale bar = 100 µm. Significance was computed by analysis of variance and Bonferroni's post hoc test in comparison to control groups. (*) P < 0.05, (**) P < 0.001, (***) P < 0.001 was considered significant.
Amelioration of obesity-induced inflammation and hepatic steatosis in obesity-induced diabetic mouse model
To confirm amelioration of inflammation, inflammatory cytokines in both epiWAT and subWAT were evaluated. The inflammatory cytokine levels were significantly reduced in the (dCas9/sgFabp4) + ATS-9R-treated groups (Fig. 5C), while cytokine levels in the subWAT were not decreased as much (Fig. 5D), demonstrating effective targeting to epiWAT suppressing obesity-induced inflammation. The major mechanism that has been linked to obesity-induced type 2 diabetes involves attraction of cytokines in the bloodstream released from excessive fat accumulation and lipolysis from adipose tissues. To confirm that Fabp4 silencing in adipose tissues suppressed inflammation, total RNA was isolated from epiWAT and subWAT to analyze the cytokines Tnf, Ccl2, Il6, and Il1b. The cytokines were down-regulated in epiWAT as compared to subWAT. This data is consistent with the targeting ability of (dCas9/sgFabp4) + ATS-9R oligoplexes as shown in the biodistribution studies (Fig. 3A). Free fatty acids are secreted by hydrolysis of stored triglycerides within white adipocytes, released in plasma, and stored in sites where they can participate in metabolic reactions. High levels of plasma free fatty acids and fatty acid binding proteins are observed in patients with obesity, and high amounts of lipids are also found in microvesicles (Heymsfield et al. 2015). Due to excess adiposity, the size of microvesicles in hepatocytes increases. This condition, known as steatosis, leads to other hepatic morbidities like nonalcoholic fatty liver disease, cirrhosis, and steatohepatitis (Adams et al. 2005). Therefore, the levels of plasma free fatty acids, triglycerides, ALT, and AST levels in the mice were determined (Fig. 5E–H). The free fatty acid, triglyceride, ALT, and AST levels in serum were reduced by as much as ∼50%, restoring hepatic metabolism. Furthermore, histological analysis of liver tissue sections from the obese mice revealed severe steatosis, while the groups treated with (dCas9/sgFabp4) + ATS-9R showed a slight reversal, showing significantly reduced lipid droplets (Fig. 5I).
CRISPRi-mediated repression of Fabp4 alleviates obesity
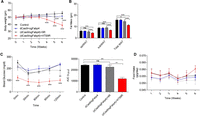
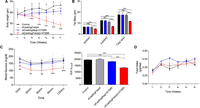
In the above results, targeting the epididymal white adipose tissue alleviated obesity and obesity-induced type 2 diabetes by suppression of Fabp4. However, the main concern with weight loss could be argued that it is due not only to CRISPRi-mediated repression of Fabp4 but to the nonspecific toxicity and activity of the CRISPRi system or the ATS-9R itself. Therefore, to prove the alleviation of obesity by CRISPRi-mediated repression of Fabp4, in vivo anti-obesity therapeutic effects including the (dCas9/sgNegative) + ATS-9R groups were determined in a HFD-induced obesity model. The mice were treated using the same protocol as mentioned above. Weekly body weight was measured and compared to (dCas9/sgFabp4) and (dCas9/sgNeg) + ATS-9R; the (dCas9/sgFabp4) + ATS-9R-treated groups showed a decrease in body weight and fat mass (Fig. 6A,B) In addition, the insulin tolerance test revealed lowered blood glucose levels following repeated treatment of (dCas9/sgFabp4) + ATS-9R oligoplexes (Fig. 6C). The loss of body weight only in the (dCas9/sgFabp4) + ATS-9R-treated groups proved that the weight loss is due to the CRISPRi repression of Fabp4 and not due to the ATS-9R reagent or the activity of the CRISPRi system. Also no significant changes in food intake were recorded during the treatment period (Fig. 6D). To further prove CRISPRi repression of Fabp4, reduction of inflammation, and restoration of hepatic metabolism, epiWAT, subWAT, and serum were analyzed. Fabp4 mRNA levels were decreased in the (dCas9/sgFabp4) + ATS-9R-treated groups as compared to (dCas9/sgNeg) + ATS-9R groups in epiWAT, while subWAT showed less suppression (Supplemental Figs. S6A,B, S11). (dCas9/sgFabp4) + ATS-9R-treated groups also showed suppression of inflammation and restoration of hepatic metabolism as compared to (dCas9/sgNeg) + ATS-9R (Supplemental Fig. S7A–F).
CRISPRi repression of Fabp4 in a high-fat diet-induced mouse model. (A) Weekly body weight. (B) Fat masses for epiWAT, subWAT, and total white adipose tissue. (C) Insulin tolerance test after 6 wk of treatment. Insulin was injected intraperitoneally and blood glucose was detected at each time point, and AUC was determined. (D) Weekly food intake. Data are presented as mean ± SEM (n = 3–5). (ns) nonsignificant, (*) P < 0.05, (**) P < 0.01, (***) P < 0.001 was considered significant.
Discussion
RNA-guided genome editing has been transforming research fields in biology, biotechnology, and medicine (Kim et al. 2014). Compared to ZFNs and TALENs, RNA-guided endonuclease (RGENs) are more scalable as they are easier to synthesize. However, they are limited by off-target effects, and efficient in vivo Cas9/sgRNA delivery can also be a challenge. In contrast to protein delivery, plasmid delivery of Cas9, dCas9, and sgRNA is more favorable in terms of high stability and effective genome editing (Jinek et al. 2013). Nonviral carriers such as lipids and peptides have been exploited as an alternative approach to deliver Cas9/sgRNA plasmids resulting in potential gene modification in vivo. In the present study, a fusion peptide consisted of adipocyte-targeting peptide and protein transduction domain, which was previously reported, was used to deliver the CRISPRi system to adipose tissues for the treatment of obesity and obesity-induced metabolic syndrome.
Studies have proven that in obese individuals the body mass is composed of white adipose tissue (WAT). WAT stores triglycerides and cholesterol ester in the intracellular lipid droplets and secretes various kinds of pro-inflammatory cytokines, making it a primary therapeutic target for the treatment of obesity and obesity-induced metabolic syndrome (Trayhurn and Wood 2004). Targeted delivery to WAT for the amelioration of obesity was driven by delivery to prohibitin of mature adipocytes and adipose tissues using the CRISPR interference mechanism. Prohibitin is expressed on adipose tissue vasculature of mature adipocytes (Kolonin et al. 2004), and our previous study exhibited prohibitin-specific targeted gene delivery in adipose tissue in vivo by use of a short peptide, ATS-9R (Yong et al. 2017). Free fatty acids play a pivotal role in obesity and insulin resistance, and FABP4 is an important therapeutic target for the treatment of obesity and induced metabolic disorders. Strategies to treat metabolic disease by targeting FABP4 have been studied by Burak et al. (2015). In this study, the delivery of dCas9/sgFabp4 with ATS-9R showed increased gene complexation and selective internalization to adipose tissues. ATS-9R-mediated adipose tissue targeting was observed in obese mice at 12 and 24 h postinjection, and intraperitoneally injected (dCas9/sgFabp4) + ATS-9R oligoplex accumulation was retained in adipose tissues and mostly cleared by the liver due to peptidase-mediated degradation. In vivo studies in a HFD-induced obesity mouse model showed that silencing of Fabp4 mediated by the CRISPRi system reduces inflammation in adipose tissues, ameliorates obesity, and restores hepatic steatosis. Obesity is associated with macrophage and immune cell infiltration leading to secretion of pro-inflammatory cytokines, TNF, CCL2, IL6, and IL1B, contributing to metabolic syndrome and insulin resistance (Sartipy and Loskutoff 2003). Thus, the nonviral gene delivery-mediated CRISPR interference system to adipose tissues possesses a strong potential as a gene therapeutic system for obesity and obesity-related metabolic syndromes. In a recent study, Matharu et al. provided a novel strategy for the treatment of obesity using CRISPR activation mediated by a recombinant adeno-associated virus to rescue a haploinsufficient phenotype in vivo (Matharu et al. 2019). This study showed that the CRISPR-mediated activation system could be a potential tool to treat obesity.
In conclusion, prohibitin and free fatty acid binding protein 4, adipocyte are potential delivery targets for obesity and obesity-induced metabolic syndromes. We have proved and confirmed the overexpression of Phb and Fabp4 in adipocytes and obese mice, making it a therapeutic target to regress obesity and obesity-related metabolic syndrome. In addition, the effective targeting to white adipose tissues using the nonviral gene delivery carrier ATS-9R together with dCas9/sgFabp4 by binding to prohibitin provided a robust CRISPR-mediated interference tool in vitro as well as in vivo. Despite its therapeutic potential, the translational research from a mouse model to patients is yet a hurdle to overcome. In the in vivo experiments, the formulations were administered twice a week for up to 6 wk. For humans, we cannot be sure whether a regimen of twice a week for up to 6 wk could be applied. Further studies regarding the translational research from a mouse model to human patients need to be carried out before clinical use. The translational research for clinical application should include dose, frequency, and interval based on PK/PD studies, maximum tolerable dose, toxicity, dose-response curve, and off-target effects. Our convenient method of the nonviral-mediated targetable CRISPR interference system can be applied for future therapeutic uses to treat obesity and obesity-induced metabolic disorders and gene dosage-sensitive disorders.
Methods
Validation of dCas9, dCas9-KRAB, and sgRNA function
Four candidate sequences to target Mus musculus Fabp4 (NM_024406.2) were designed, and their sgRNA expression vectors were constructed at ToolGen. The Fabp4 knockdown was evaluated in adipocytes and the best candidate, sgFabp4, was selected based on the highest Fabp4 knockdown efficiency as evaluated by T7E1 nuclease assay and quantitative real-time polymerase chain reaction. pcDNA-dCas9 was a gift from Charles Gersbach (Addgene plasmid # 47106); dCas9-KRAB was a gift from Alejandro Chavez and George Church (Addgene plasmid # 110820).
Peptide synthesis
The peptides ATS-9R (CKGGRAKD-RRRRRRRRRC) and 9R (CRRRRRRRRRC), with or without FITC conjugation, were purchased from Peptron, Inc. The molecular weights of peptides were 9R: 1628 Da, ATS: 937 Da, ATS-9R: 2341 Da, and FITC-ATS-9R: 2844 Da.
Synthesis and characterization of (dCas9/sgFabp4) + ATS-9R oligoplexes
The morphological characteristics of the oligoplexes were confirmed by energy filtered transmission microscopy (EM9120, Carl Zeiss Vision GmbH) in the Korea Basic Science Institute (Chuncheon, Korea). To prepare each sample, 10 µg of dCas9/sgFabp4 were condensed with 9R and ATS-9R in deionized water for 30 min. Samples were then dried at 50°C on 10 × 10 transmission electron microscopy grids. Electrophoretic mobility shift assays were performed to confirm the ATS-9R condensation properties. Oligoplexes with different amounts of ATS-9R were prepared with a constant amount of dCas9/sgFabp4 in deionized water. After 30 min of incubation, samples were electrophoresed on 0.8% (w/v) agarose gels (Lonza) in TBE buffer solution at 100 V for 20 min and observed using an imaging station. The particle size distributions and surface charges of oligoplexes at different weight ratios were determined using dynamic light scattering with a Zetasizer Nano ZS (Malvern Instruments) after 30 min of incubation at room temperature.
Cell culture and 3T3-L1 adipocyte differentiation
3T3-L1 pre-adipocytes were cultured and maintained in high glucose Dulbecco's Modified Eagle's Medium supplemented with 10% fetal bovine serum (FBS), 1% penicillin and streptomycin at 37°C in 5% CO2. For differentiation, 72 h after seeding, pre-adipocytes were treated for an additional 72 h in differentiation medium containing complete medium, 10 µg/mL insulin, 1 µM dexamethasone, and 0.5 mM isobutyl methylxanthine. The differentiation medium was then replaced with maintenance medium containing complete medium and 10 µg/mL insulin every other day for a total of 9 d.
Competition assay
3T3-L1 cells were seeded and incubated on 12-well plates for 24 h. Oligopeptide complexes were prepared by mixing a constant amount of luciferase plasmid DNA with ATS-9R at weight ratios of 3, 4, and 5. Cells were transfected 24 h after seeding. Additionally, cells were transfected with DNA/9R, DNA/ATS-9R, or DNA/Lipofectamine oligoplexes and incubated for an additional 48 h. The 9R and Lipofectamine oligoplexes were used as positive controls. After 48 h of incubation and washing with phosphate buffered saline (PBS), cells were treated with 150 µL of 1× cell lysis buffer for 10–15 min. Then, the cell lysates were scraped, collected, and transferred to microtubes for centrifugation. Transfection efficiency was assessed by measuring the luciferase expression of the luciferase plasmid DNA. The relative luminescence units (RLU) emitted after adding luciferase substrate solution were recorded on a 96-well plate luminometer (Berthold Detection Systems) with 20-sec integration and expressed as RLU/mg of cell protein. The protein concentrations of the cell lysates were determined by the DC protein assay kit using a bovine serum albumin standard. Results were expressed as RLU/mg of cell lysate.
Flow cytometry analysis
Cy5.5 fluorescent-labeled dCas9/sgFabp4 was used to assess the mean fluorescence intensity of 3T3-L1 cellular uptake by flow cytometry. After 24 h of transfection, cells were collected following trypsinization, and a single-cell suspension was prepared in FACS buffer (2% FBS and 0.02% sodium azide in PBS). Internalization of oligoplexes was assessed by a FACS Canto II Caliber flow cytometer (BD Biosciences), and data were quantified by CellQuest Pro. Adipocytes were properly gated by forward and side scatter, and 10,000 events were perceived per sample.
Confocal microscopy
Further assessment of (dCas9/sgFabp4) + ATS-9R oligoplexes was performed by confocal microscopy. Both 3T3-L1 pre-adipocytes and mature adipocytes were grown on glass coverslips in six-well plates. Cells were then transfected with fluorescein-conjugated ATS-9R and incubated at various times up to 24 h. One day after transfection, the cells were extensively washed and fixed with 4% formaldehyde. Plasma membranes were stained with CellMask Deep Red (Thermo Fisher Scientific). Cells were washed with PBS and mounted using DAPI Fluoromount-G (Southern Biotech) to stain nuclei. Images were visualized with a Carl Zeiss confocal laser scanning microscope (LSM510 META NLO, Carl Zeiss Jena GmBH) at the Korea Basic Science Institute (Chuncheon, Korea).
Cytotoxicity assay
3T3-L1 pre-adipocytes were seeded on 24-well plates at a density of 5 × 104 cells/well. After 24 h of incubation, culture media were replaced with media containing the (dCas9/sgFabp4) + ATS-9R oligoplexes. Oligoplexes were prepared by mixing with 9R and ATS-9R at weight ratios of 4 and 5. After 24 h, 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) solution was added to the media at 10% v/v and incubated for 4 h. Then, the medium was replaced with dimethyl sulfoxide and incubated for an additional 15 min. The absorbance of each sample was measured at 540 nm using a UV/Vis spectrophotometer (Tecan). The viability was measured three times for each of four replicates.
In vitro Fabp4 knockdown
For RT-PCR analysis, cells were seeded on 12-well plates for 24 h. Cells were transfected with (dCas9/sgFabp4) + ATS-9R oligoplexes and incubated for an additional 24 h. Total RNA were isolated from pre-adipocytes and mature adipocytes using the RNeasy Lipid Tissue Mini kit (Qiagen GmbH) according to the manufacturer's protocol, and cDNA was synthesized using an iScript cDNA synthesis kit (Bio-Rad). The Fabp4 mRNAs were measured by quantitative real-time PCR (qRT-PCR) using a SensiFAST SYBR Lo-ROX kit (Bioline). Mouse Gapdh was used as an endogenous control. The ΔΔCt method was used to calculate the Fabp4 mRNA level relative to that of Gapdh.
Oil Red O staining
Oil Red O staining test was performed to confirm adipocyte differentiation and the knockdown effect of the therapeutic gene. For Oil Red O solution preparation, 0.7 g of Oil Red O powder was dissolved in 200 mL isopropanol, stirred overnight, and filtered through a 0.22-µm syringe filter. Oil Red O solution was mixed with deionized water at a 6:4 ratio to make Oil Red O working solution and filtered through a 0.22-µm syringe filter. For staining, cells were washed with PBS and fixed with 4% formaldehyde at room temperature for 1 h. Then, cells were rinsed with 60% isopropanol and dried on a clean bench. Cells were next stained with Oil Red O solution for 15 min in an incubator. Before microscopic observation, the cells were washed with deionized water to remove excess stain.
T7E1 assay
Mature adipocytes were transfected with 3 µg of dCas9 plasmid and sgFabp4 plasmid with ATS-9R. Cells were incubated at 37°C for 48 h posttransfection prior to genomic DNA isolation. Genomic DNA was isolated using a DNA isolation kit (DNeasy Blood & Tissue Kit, Qiagen) following the manufacturer's protocol. The genomic region flanking the CRISPRi target site for the Fabp4 gene was amplified by PCR (Fabp4 primer) using the HiPi DNA polymerase (Elpis Bio) following the manufacturer's protocol. Then, 100 ng of PCR products were subjected to a re-annealing process to enable heteroduplex formation: 95°C for 10 min, 95°C to 85°C ramping at −2°C/sec, and 85°C to 25°C at −0.1°C/sec. Re-annealing products were further reacted with SURVEYOR nuclease (NEB) following the manufacturer's protocol. After 20 min of incubation at 37°C, the annealed products were analyzed on a 2% polyacrylamide gel.
High-fat diet-induced obese mouse model
Six-wk-old male C57BL/6J mice were obtained from Orient Bio. All experimental procedures were reviewed and approved by the Institutional Animal Care and Use Committee at Hanyang University. Mice were randomly assigned to one of four treatment groups (n = 5 per group). For obesity and metabolic syndrome induction, the C57BL/6J mice were fed normal chow (Orient Bio) for the first 2 wk; after that, the food was mixed with 10% of a high-fat diet that provided 60% of calories from fat (Central Lab Animal, Inc.). The proportion of the HFD of the total rodent diet was gradually increased until 6 wk, after which the mice were fed only the HFD for eight more weeks. All mice were housed under specific pathogen-free conditions and were given free access to food and water. Mice became obese and insulin resistant after week 14, with approximate body weight of 45–60 g and glucose level >250 mg/dL.
Biodistribution studies
ATS-9R was conjugated with Cy5.5 according to the manufacturer's instructions. Oligoplexes were prepared by mixing dCas9/sgFabp4 with Cy5.5-ATS-9R for 30 min at room temperature. The Cy5.5-conjugated oligoplexes were injected intraperitoneally into obese mice. Mice were sacrificed after 12 and 24 h of injections, and the liver, spleen, kidneys, lungs, heart, visceral adipose tissues, and subcutaneous adipose tissues were harvested. Ex vivo Cy5.5 fluorescence in each organ was visualized using the VISQUE In Vivo Smart Fluorescence device at the Korea Basic Science Institute (Chuncheon, Korea).
In vivo therapeutic experiments
The dCas9/sgFabp4 formed complexes with 9R and ATS-9R peptides in PBS at the optimum weight ratios and incubated at room temperature for 30 min. The dCas9/sgNegative also formed complexes with ATS-9R and was incubated at room temperature for 30 min. Mice were injected with oligoplexes intraperitoneally twice a week for 6 wk with a dose of 0.35 mg/kg. The body weights and food intake of the mice were measured every week after the first treatment.
Ex vivo cytokine analysis and T7E1 analysis in adipose tissues
Epididymal white adipose tissue and subcutaneous white adipose tissue samples were mechanically crushed and then incubated in 1 mg/mL of collagenase solution for 1 h at 37°C. The homogenates were centrifuged at 400g for 10 min to isolate the stromal vascular fraction from the adipose tissue, after which total RNA was extracted from the cells. After preparing equal amounts of cDNA from the extracted RNA, the relative expression levels of Tnf, Ccl2, Il6, Il1b, Fabp4, and Phb in the adipose tissue were quantified by real-time PCR compared to the Gapdh control calculated using the ΔΔCt method. Genomic DNA (Qiagen, GmbH) was obtained from epididymal white adipose tissue and subcutaneous white adipose tissue samples according to the manufacturer's protocol, and T7E1 assays were carried out as previously described.
In vivo insulin tolerance test (ITT)
Initial blood glucose level was measured after 6 h of fasting using an Accu-Chek Active model GC kit (Roche Diagnostics GmbH). Insulin (0.75 U/kg body weight) was injected intraperitoneally after measuring the initial blood glucose level. Blood samples were collected from the tail vein at 0, 30, 60, 90, and 120 min postinjection.
Statistical analysis
For all in vitro studies, a minimum of three experimental replicates was used. To achieve statistical significance in the in vivo studies, samples were five per group. All statistical analyses were conducted using GraphPad Prism version 5 for Windows (GraphPad Software). The statistical significance of comparisons between two groups was calculated by a two-tailed Student's t-test, and P-values less than 0.05 were considered significant. All other experiments were analyzed using two-way analysis of variance with Bonferroni's correction and two-sided tests for multiple comparisons between subgroups. The significance level was set as P < 0.001.
Acknowledgments
This research was partially supported by grants from the National Research Foundation of Korea (NRF-2015R1A2A1A09003019, NRF-2019R1A2C3008992) and Bio & Medical technology development program (NRF-2017M3A9F5029655), the research fund of Hanyang University (HY-201800000000282), the Brain Korea 21 plus program (22A20130011095), and the Korean Health Technology R&D project through the Ministry of Health and Welfare (HI17C0888).
Author contributions: J.Y.C., Q.U.A., and Y.-H.K. designed the experimental protocols. J.Y.C. carried out the in vitro therapeutic assays. Q.U.A. performed the characterization experiments and in vitro transfection studies of oligoplexes. J.Y.C., Q.U.A., Y.S., and S.-B.Y. performed the in vivo studies. J.Y.C., Q.U.A., and Y.-H.K. discussed the results and cowrote the manuscript.
Footnotes
-
[Supplemental material is available for this article.]
-
Article published online before print. Article, supplemental material, and publication date are at http://www.genome.org/cgi/doi/10.1101/gr.246900.118.
- Received November 28, 2018.
- Accepted July 23, 2019.
This article is distributed exclusively by Cold Spring Harbor Laboratory Press for the first six months after the full-issue publication date (see http://genome.cshlp.org/site/misc/terms.xhtml). After six months, it is available under a Creative Commons License (Attribution-NonCommercial 4.0 International), as described at http://creativecommons.org/licenses/by-nc/4.0/.