A long-term demasculinization of X-linked intergenic noncoding RNAs in Drosophila melanogaster
- Ge Gao1,8,
- Maria D. Vibranovski2,3,8,
- Li Zhang1,2,
- Zheng Li4,
- Ming Liu4,
- Yong E. Zhang2,5,
- Xinmin Li6,9,
- Wenxia Zhang4,
- Qichang Fan4,
- Nicholas W. VanKuren2,7,
- Manyuan Long2,10 and
- Liping Wei1,10
- 1State Key Laboratory of Protein and Plant Gene Research, College of Life Sciences and Center for Bioinformatics, Peking University, Beijing 100871, China;
- 2Department of Ecology and Evolution, University of Chicago, Chicago, Illinois 60637, USA;
- 3Departamento de Genética e Biologia Evolutiva, Instituto de Biociências, Universidade de São Paulo, São Paulo, Brazil 05508;
- 4Center of Developmental Biology and Genetics, College of Life Sciences, Peking University, Beijing 100871, China;
- 5Key Laboratory of the Zoological Systematics and Evolution, Institute of Zoology, Chinese Academy of Sciences, Beijing 100101, China;
- 6Genome Facility, University of Chicago, Chicago, Illinois 60637, USA;
- 7Committee on Genetics, Genomics, and Systems Biology, University of Chicago, Chicago, Illinois 60637, USA
-
↵8 These authors contributed equally to this work.
Abstract
Recent studies have revealed key roles of noncoding RNAs in sex-related pathways, but little is known about the evolutionary forces acting on these noncoding RNAs. Profiling the transcriptome of Drosophila melanogaster with whole-genome tiling arrays found that 15% of male-biased transcribed fragments are intergenic noncoding RNAs (incRNAs), suggesting a potentially important role for incRNAs in sex-related biological processes. Statistical analysis revealed a paucity of male-biased incRNAs and coding genes on the X chromosome, suggesting that similar evolutionary forces could be affecting the genomic organization of both coding and noncoding genes. Expression profiling across germline and somatic tissues further suggested that both male meiotic sex chromosome inactivation (MSCI) and sexual antagonism could contribute to the chromosomal distribution of male-biased incRNAs. Comparative sequence analysis showed that the evolutionary age of male-biased incRNAs is a significant predictor of their chromosomal locations. In addition to identifying abundant sex-biased incRNAs in the fly genome, our work unveils a global picture of the complex interplay between noncoding RNAs and sexual chromosome evolution.
Sex chromosomes are major targets for sex-related selection, and many genome-wide studies have revealed differences between sex chromosomes and autosomes with respect to divergence rate, gene content, and gene expression patterns (Vicoso and Charlesworth 2006; Ellegren and Parsch 2007; Mank 2009; Qvarnstrom and Bailey 2009). Male-biased coding genes, i.e., genes that are more highly expressed in males than in females, are unevenly distributed between the sex chromosomes and autosomes in Drosophila, mammals, and worms (Betrán et al. 2002; Parisi et al. 2003; Ranz et al. 2003; Khil et al. 2004; Reinke et al. 2004; Wang et al. 2005). In mammals, coding genes expressed in male meiotic or post-meiotic cells are underrepresented on the X chromosome, whereas coding genes expressed in premeiotic stem cells are overrepresented on the X chromosome (Khil et al. 2004). Similar results were obtained from transcriptional profiling of samples enriched with D. melanogaster meiotic cells (Vibranovski et al. 2009a). In D. melanogaster and in Caenorhabditis elegans gonads, male-biased coding genes are found to be underrepresented on the X chromosome (Betrán et al. 2002; Parisi et al. 2003; Ranz et al. 2003; Reinke et al. 2004). In Drosophila this underrepresentation is observed only for old X-linked coding genes. Young male-biased coding genes, those that emerged after the split of the melanogaster subgroup (<3–6 million yr ago) (Russo et al. 1995), are found to be overrepresented on the X chromosome (Zhang et al. 2010a).
Directional movement of male-biased genes out of the X chromosome is one evolutionary process that could contribute to such an uneven chromosomal distribution of male-biased genes in these taxa. New Drosophila retrogenes tend to escape from the X chromosome and are more likely to be expressed in testis (Betrán et al. 2002), and excessive male-biased retrogene traffic has been observed on the mammalian X chromosome (Emerson et al. 2004). Further studies showed that new DNA-based duplicate coding genes exhibit a similar chromosomal distribution pattern to retrogenes (Betrán et al. 2002; Emerson et al. 2004; Meisel et al. 2009; Vibranovski et al. 2009b), suggesting that the uneven chromosomal distribution of male-biased genes might not depend on a specific molecular mechanism but rather is the product of natural selection acting on genes with male-related functions (Meisel et al. 2009; Vibranovski et al. 2009b). This hypothesis is supported by independent evidence from population genomic analysis of copy number variation of Drosophila retrogenes (Schrider et al. 2011).
Several different hypotheses invoking natural selection could explain the paucity of X-linked male-biased genes. First, inactivation of X-linked genes during male meiosis (meiotic sex chromosome inactivation [MSCI]) (Lifschytz and Lindsley 1972) may favor the accumulation of testis-expressed genes in autosomes (Betrán et al. 2002; Emerson et al. 2004). Second, the sexual antagonism hypothesis predicts that the chromosomal distribution of sex-biased genes is driven by sexually antagonistic forces (Rice 1984), such that dominant alleles with beneficial fitness effects in males, but detrimental effects in females, have a higher probability of being fixed on the autosomes (Charlesworth et al. 1987). Another dominance-independent hypothesis of sexual antagonism driving germline X-inactivation predicted the demasculinization of the X chromosome based on different sojourning times of X, 1/3 of the time in males and 2/3 of the time in females, which would result in an excess of male-biased genes out of the X chromosome and enrichment of X-linked female-biased genes (Wu and Xu 2003). Third, the dosage compensation hypothesis predicts that hypertranscription of the Drosophila X chromosome in males prevents further up-regulation of X-linked male-biased genes, thus favoring their relocation to an autosome (Vicoso and Charlesworth 2009; Bachtrog et al. 2010). Although evidence for all three hypotheses has been demonstrated using coding genes in Drosophila (Parisi et al. 2003; Hense et al. 2007; Vibranovski et al. 2009a, Bachtrog et al. 2010; Kemkemer et al. 2011, 2013), no systematic experimental study on noncoding RNAs has been done so far to test these hypotheses. Noncoding RNAs (ncRNAs) play important roles in many reproductive processes (Mattick and Makunin 2005; Prasanth and Spector 2007). If selection governs the chromosomal distribution of sex-biased genes, we expect male-biased ncRNAs to exhibit a chromosomal distribution similar to that observed for coding genes.
In this report, we tested these hypotheses by experimentally identifying male-biased ncRNAs in D. melanogaster and analyzing their chromosomal distribution. Whole-transcriptome profiling revealed a large number of intergenic noncoding RNAs (incRNAs) with male-biased expression in both whole body and reproductive organs, which we confirmed with RT-PCR. We demonstrate that these incRNAs are unevenly distributed between the autosomes and X chromosome. Comparisons of germline and somatic tissue transcriptional profiles suggest that sexual antagonism and male germline MSCI both could be contributing to the peculiar chromosomal distributions of male-biased incRNAs. In concordance with previous studies on coding genes, comparative genomics analyses revealed that male-biased incRNAs that originated during different evolutionary periods have different chromosomal distribution patterns, indicating that evolutionary time has a significant effect on their chromosomal locations (Zhang et al. 2010a). As for coding genes (Zhang et al. 2010a), we found that old male-biased incRNAs (>6 my old) are enriched on autosomes, whereas new male-biased incRNAs are enriched on the X chromosome.
In addition, our analyses shed some light in the current debate about the demasculinization of the X chromosome in Drosophila. More specifically, our analyses clarify why recent studies have shown that Drosophila testis-specific genes are not underrepresented in the X chromosome, but male-biased genes are (Meiklejohn and Presgraves 2012; Meisel et al. 2012). Gene age is positively correlated with expression breadth (Zhang et al. 2012). Therefore testis-specific genes (narrowly expressed genes) are enriched with very young genes, which were previously shown to be overrepresented in the X chromosome (Zhang et al. 2010a). In order to evaluate the chromosomal distribution of testis-specific genes in an unbiased way, we analyzed the old testis-specific coding genes and found that they are underrepresented in the X chromosome. These results corroborate our previous findings that the process of desmasculinization is an evolutionary process that appears only over evolutionary time in both Drosophila and mammals (Zhang et al. 2010a,b).
Results and Discussion
Transcriptome profiling reveals abundant incRNAs with sex-biased expression
We first profiled the transcriptomes of whole male and female D. melanogaster adults using Affymetrix whole-genome tiling arrays. The arrays used 3,116,816 25-nt probe pairs to assay transcription of 109,088,560 bp of repeat-free euchromatic genome. We detected a total of 35,884,625 bp (∼32.89%) in male and 32,921,857 bp (∼30.18%) in female as being transcribed. Of those nucleotides transcribed, 7,738,215 bp (∼21.56%) exhibited male-biased expression and 3,649,022 bp (∼11.08%) exhibited female-biased expression with a greater than twofold increase. In addition to whole body samples, we profiled the transcriptome of adult reproductive tracts (gonads: testis, ovaries; and accessory glands) and nonreproductive tracts (gut and thorax). We identified 29,188,400 bp (∼26.76%), 25,316,156 bp (∼23.21%), and 25,843,450 bp (∼23.69%) as being transcribed in testis, ovaries, and accessory glands, respectively. Of those nucleotides transcribed, 10,066,819 bp (∼34.49%) in testis and 7,633,727 bp (∼29.54%) in accessory glands were identified as male-biased with a greater than twofold increase over expression in ovaries. As expected, a relatively low proportion of male-biased transfrags (transcribed fragments; see Methods for further details) was found in gut (7.36%; 2,605,069 bp of 35,390,445 bp) and thorax (9.51%; 3,043,360 bp of 32,001,727 bp) (detailed transfrag counting and proportions could be found in Supplemental Fig. S1 and Supplemental Table S1).
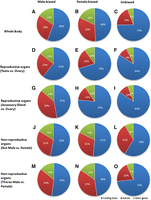
According to FlyBase annotation release 5.46, 10%–20% of male-biased and female-biased transfrags belong to intergenic regions (Fig. 1). Analysis of coding potential with Coding Potential Calculator (CPC) (Kong et al. 2007) suggested that >98% of these intergenic transfrags are truly noncoding transcripts. We now focus on these intergenic noncoding RNAs, or incRNAs. Intronic noncoding transfrags were excluded from most of our analyses because current annotations might not distinguish true intronic noncoding RNAs from novel exons of protein-coding genes or nondegraded intronic transcripts (we provide the chromosomal distribution of intronic ncRNAs in Supplemental Fig. S2).
Genomic distribution of sex-biased transfrags. Expression profiling was done with Affymetrix whole-genome tiling arrays. Exon/intron/intergenic annotations were retrieved from FlyBase (version 5.46). Rows represent comparisons of male and female whole body RNA (A–C), testis versus ovary (D–F), accessory gland versus ovary (G–I), male versus female gut (J–L), and male versus female thorax (M–O). Columns represent male-biased (A,D,G,J,M), female-biased (B,E,H,K,N), and non-sex-biased expression (C,F,I,L,O).
In all three comparisons performed using male/female adult transcriptome profiles derived from either whole body or reproductive tracts organs (Fig. 1A–I), the fraction of incRNAs is significantly higher in sex-biased comparisons than in non-sex-biased comparisons (P < 2.2 × 10−16, χ2 test) (Fig. 1A,B versus Fig. 1C; Fig. 1D,E versus Fig. 1F; Fig. 1G,H versus Fig. 1I), suggesting as expected that incRNA expression differences between males and females tend to be associated with sex-related biological processes. As expected, comparisons between reproductive organs (testis and accessory glands versus ovaries) identified significantly more sex-biased incRNAs than in the nonreproductive organs comparisons (32%–35% versus 12%–16%; χ2 test, P < 2.2 × 10−16) (Supplemental Fig. S3), suggesting that incRNAs, like any other transcription unit, are more prone to be involved with sex-related functions in the reproductive organs. Two examples of sex-biased intergenic noncoding RNAs are shown in Supplemental Figure S4.
Sex-biased incRNAs are nonrandomly distributed between the autosomes and X chromosome
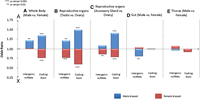
We tested the hypothesis that selection governs the chromosomal distribution of sex-biased genes by comparing the distributions of sex-biased coding and noncoding RNAs. If male-biased ncRNAs are randomly distributed along the chromosomes, there is probably no selection forces acting on noncoding regions, and this is a unique property of the coding genes. As predicted by our hypothesis, an excess of autosomal male-biased incRNAs were identified in whole body (21% excess) and testis (21%) (Fig. 2) (odds ratios [ORs] = 1.21, P ≤ 0.01, Fisher's exact test, for each comparison). We applied the Fisher's exact test to assess the uneven chromosomal distribution using the estimated odds ratio (OR) as an intuitive measurement. Odds ratio is the ratio between male-biased genes (autosomal/X-linked) and non-male-biased genes (autosomal/X-linked). Thus, an odds ratio >1 indicates male-biased genes are enriched on autosomes, and <1 indicates X enrichment. (See Methods for more details). A similar trend was found for coding transfrags, consistent with previous studies (Fig. 2; Parisi et al. 2003; Ranz et al. 2003; Emerson et al. 2004; Sturgill et al. 2007). In contrast, we found that female-biased incRNAs are significantly overrepresented on the X chromosome in comparisons of ovary versus testis and ovary versus accessory gland (OR range = 0.77–0.86, P < 6.53 × 10−3) (Fig. 2B,C). No significant departure from random chromosomal distribution, however, was observed for female-biased incRNAs derived from whole body female versus male comparison (OR = 1.01, P = 0.44) (Fig. 2A).
X chromosome demasculinization was observed for incRNAs as well as coding transfrags, based on comparison of whole body of males versus females (A), testes versus ovaries (B), and accessory glands versus ovaries (C). Comparison of the gut and thorax of males versus females is shown in D and E, respectively. Odds ratios >1 indicate enrichment on autosomes, and <1 indicates enrichment on X chromosome. Significant deviations are indicated: (***) P < 0.001; (**) P < 0.01, Fisher's exact test. Blue and red bars represent male-biased and female-biased transfrags, respectively.
We verified sex-biased expression of incRNAs using tissue-specific RT-PCR. We previously detected 528 incRNAs in adult flies in a random sample of D. melanogaster intergenic regions using the RT-PCR approach (Li et al. 2009). Using the same primers, we detected about half of these incRNAs (261 of 528) in testes, ovaries, and heads in Drosophila adults (RT-PCR primers are listed in Supplemental Table S2). We considered testis-biased incRNAs those that were detected in testis but not in ovaries. The reverse logic was applied to ovary-biased incRNAs. We considered non-sex-biased incRNAs those that were detected in both testis and ovary and those that were detected only in head. The chromosomal distribution of testis-biased and ovary-biased incRNAs differed significantly from the distribution of non-sex-biased incRNAs (P = 0.0287, Fisher's exact test) (Supplemental Table S3), with a significant deficiency of X-linked testis-biased incRNAs (OR = 0, P = 0.0223 for testis versus non-sex-biased incRNAs) and a marginal enrichment of X-linked female-biased incRNAs (OR = 1.35, n.s., probably due to small sample size). Thus the PCR-based independent test revealed the same robust chromosomal distribution patterns for Drosophila male-biased incRNAs, verifying that the observed underrepresentation of male-biased incRNAs in the X chromosome is not a methodological artifact.
Disentangling the contribution of sexual antagonism and MSCI to the demasculinization of the X chromosome
X chromosome demasculinization is the evolutionary process by which selective forces drive male-biased genes off the X chromosome, either relocating them to the autosomes or eliminating them from the genome entirely. We investigated the contribution of MSCI and sexual antagonism to the observed X chromosome demasculinization for incRNAs in Drosophila. It is not trivial to separate the effects of sexual antagonism and MSCI, as MSCI only occurs in male meiosis, but sexual antagonism may occur in any tissue or cell. However, most sex-biased expression is found in testes and ovaries, especially in the meiotic phase (Parisi et al. 2003; Vibranovski et al. 2009a). MSCI could therefore be assessed by analyzing testis-expressed genes with biased expression in meiosis but not in mitosis, thus including the effect of inactivation of X-linked genes in meiotic cells but ignoring sexual antagonistic effects present in mitotic cells (Vibranovski et al. 2009a). However, meiotic cells could also be under the effect of sexual antagonistic forces preventing the complete separation of those two processes. We thus identified male-biased RNAs involved in meiosis as those that are testis-biased but not accessory-gland biased, as accessory glands only contain mitotic cells. Accessory glands produce proteins and compounds that comprise seminal fluid and affect the reproductive capacity of both sexes (Ravi Ram and Wolfner 2007). Accessory gland-biased genes are therefore potential sexually antagonistic genes. We observed a statistically significant over-representation of strictly testis-biased incRNAs on the autosomes (OR = 1.20; P = 0.0069), suggesting that MSCI contributes to the desmasculinization of the X chromosome despite the effect of accessory gland expressed genes with sexual antagonistic effects. It is possible that there are sexually antagonistic genes expressed in testis mitotic cells that are not expressed in accessory gland mitotic cells. Therefore our data only suggest the role of X-inactivation in producing the paucity of X-linked male-biased incRNAs. Nevertheless, the role of MSCI observed for incRNAs was also observed for coding exons (OR = 1.50, P = 1.07 × 10−31) and is consistent with previous observations derived from protein-coding genes expressed in meiosis (Vibranovski et al. 2009a).
Conversely, we assessed the effects of sexual antagonism by comparing incRNAs with biased expression in accessory gland (somatic) but unbiased in testis (spermatogenesis). Statistical tests showed no significant X demasculinization for these incRNAs (OR = 1.01, P = 0.464). Although we found no X demasculinization of accessory gland-biased incRNAs, it should be noted that accessory gland-biased coding exons are significantly underrepresented on the X chromosome even after removing genes that are also testis-biased (OR = 1.41, P = 2.82 × 10−12). Moreover, sex-biased genes in other male-specific tissues may be sexually antagonistic; accessory gland is not the sole male-specific somatic tissue in Drosophila (e.g., male genitalia) (Liu et al. 1996).
To further investigate the effects of sexual antagonism in other somatic tissues, we performed additional transcriptome profiling on nonreproductive organs: the gut and thorax of male and female adults. No significant chromosomal distribution imbalance for either male-biased or female-biased coding exons was found (Fig. 2D,E). Female-biased incRNAs expressed in the thorax and gut do not deviate from the random chromosomal distribution in both tissues. Male-biased incRNAs are also randomly distributed on chromosomes, except those expressed in the gut, which are significantly overrepresented on the X chromosome (Fig. 2D), the opposite pattern expected for demasculinization. One possible explanation for the excess of X-linked male-biased genes found only in the gut is a higher proportion of young genes, which are known to be found in excess in the X chromosome (Zhang et al. 2010a). Indeed, 10% of male-biased genes expressed in the gut originated <3 million yr ago in comparison to 7% of those expressed in the thorax (Fisher's exact test, P = 0.066). The generally random chromosomal distribution of sex-biased genes in the gut and thorax, except for the excess of male-biased incRNAs in the X, suggests that the demasculinization of genes is more often associated with reproductive organs.
Moreover, our entire data, which combine reproductive and nonreproductive organs, suggest that the X demasculinization effects of sexual antagonism are limited to accessory gland-biased coding genes. These results support the hypothesis that sexual antagonism probably contributes less than MSCI to the nonrandom chromosomal distribution of male-biased genes in Drosophila.
Our results from nonreproductive organs do not support the involvement of dosage compensation in generating a paucity of male-biased protein-coding genes observed in the Drosophila X chromosome (Vicoso and Charlesworth 2009; Bachtrog et al. 2010). That is, although the thorax and gut also experience dosage compensation, we observed no paucity of X-linked male-biased protein-coding genes in those tissues.
X chromosome demasculinization
Two recent studies have questioned both the demasculinization of the X chromosome in Drosophila and the contribution of MSCI to the phenomenon (Meiklejohn and Presgraves 2012; Meisel et al. 2012). In both papers, the authors claimed that a deficit of male-biased genes on the X chromosome is attributable solely to lower average expression of genes on the X relative to the autosomes in Drosophila testes, most likely due to an absence of dosage compensation in the germline (Meiklejohn and Presgraves 2012; Meisel et al. 2012). In both studies, this argument is mainly based on the random chromosomal distribution of testis-specific genes (i.e., those that are expressed in testis but not in any other tissue) as opposed to the deficit of testis-biased genes (i.e., those that are expressed more in testis than ovary) on the X chromosome. The interpretations of those results are the following: (1) MSCI is not a factor that contributes to the nonrandom chromosomal distribution testis-biased genes because the X chromosome shows no deficit of testis-specific genes that, in theory, should be under meiotic inactivation; (2) deficit of testis-biased genes should be attributed to lack of dosage compensation in the male germline as comparisons between testis and ovaries are the only analysis that present the deficit; and (3) direct comparisons between testis and ovary expression do not control for correlation between expression breadth and sex-biased expression. Therefore, by comparing those sex-specific tissues, one might obtain evidence for the paucity of X-linked testis-expressed genes.
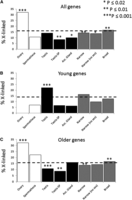
However, those studies did not take into account the importance of gene age when looking to the random chromosomal distribution of testis-specific genes (Meiklejohn and Presgraves 2012; Meisel et al. 2012). It is now well established that young male-biased genes both in Drosophila and mammals tend to be more frequently found in the X chromosome, whereas the opposite pattern is found for older male-biased genes (Zhang et al. 2010a,b). Therefore, the process of desmasculinization is an evolutionary process that appears over evolutionary time. In order to evaluate the chromosomal distribution of testis-specific genes in an unbiased way, we should take into account only older genes. Moreover, neither study accounted for the relationship between gene age and expression breadth. Testis-specific genes are genes narrowly expressed in testis. Young genes tend to be more narrowly expressed than older genes (Zhang et al. 2012). We tested the hypothesis that the random chromosomal distribution of testis-specific, but not of testis-biased, genes is caused by a large number of testis-specific genes being newly evolved genes. We classified coding genes according to their expression breadth following Meisel et al. (2012) and separated older and young genes according to ages as defined in Zhang et al. (2010a), branch 0 and branches 1–6, respectively. First, we confirmed that narrowly expressed coding genes are enriched with new genes (Supplemental Fig. S6; Zhang et al. 2012). This pattern is also true for testis-specific coding genes (Supplemental Fig. S6). Second, young testis-specific coding genes are enriched on the X chromosome, whereas older testis-specific coding genes are deficient from the X chromosome (Fig. 3B,C, respectively). We therefore conclude that the result of random distribution of testis-specific coding genes (Meiklejohn and Presgraves 2012; Meisel et al. 2012) is a consequence of the enrichment of testis-specific coding genes with recently evolved coding genes in a short evolutionary period. Therefore, neither demasculinization nor MSCI can be ruled out as important players in determining the chromosomal distribution of male-biased coding genes in Drosophila as older testis-specific coding genes are underrepresented in the X chromosome (Fig. 3C).
Percentage of D. melanogaster X-linked genes that are broadly and narrowly expressed. Following the methods in Meisel et al. (2012), genes narrowly expressed are also called specific genes (τ > 0.7) in each of four sex-limited tissues, with testis-specific expression and detectable in the sperm proteome (testis-SP), narrowly expressed in any of 14 tissues (narrow), narrowly expressed in one of 10 non-sex-limited tissues (no sex), or broadly expressed genes (τ ≤ 0.4). Percentage of X-linked in the genome are shown by dashed lines. Significant deviations (Fisher's exact test) are indicated: (***) P < 0.001; (**) P < 0.01; (*) P < 0.02. All X-linked genes (A) were separated according to their evolutionary age. New (B) and old (C) genes were defined according to Zhang et al. (2010a), in which old genes are at least as old as the split between Sophophora and Drosophila subgenera. At first (A), there is no paucity of testis-specific genes in the X chromosome. However, opposite patterns are found for old and new genes: Although old testis-specific genes are underrepresented (C), new testis-specific genes are found in excess in the X chromosome (B).
Although there is no argument over the presence of X chromosome reduced expression in male germline cells, there are different opinions for the period of this expression reduction. MSCI studies presented evidence of expression reduction in the meiotic stage (e.g., Vibranovski et al. 2009a), whereas one study believed that the reduction is also extended to the mitotic stage of spermatogenesis (Meiklejohn et al. 2011). However, recent analyses reported three independent lines of evidence in favor of MSCI analyzing the expression of testis-specific promoter reporters, testis from larval stages, and from meiotic arrest mutants (Deng et al. 2011; Vibranovski et al. 2012; Kemkemer et al. 2013). Nevertheless, the work presented here does not provide evidence in favor or against MSCI, but the patterns found are consistent with the phenomenon.
Male-biased incRNA gene linkage depends on gene age
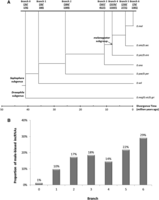
Both male-biased incRNAs and male-biased coding transfrags are significantly deficient from the X chromosome, but this trend is stronger for coding than for noncoding transfrags (Fig. 2). The comparison of testis-biased coding vs. noncoding transfrags shows OR = 1.21 versus 1.49 (P = 6.63 × 10−4), and the entire gene unit vs. noncoding transfrags shows OR = 1.21 versus 1.39 (P = 0.026) (see Supplemental Table S4 for details). Do different evolutionary ages of male-biased coding and noncoding genes play any role in determining the evolutionary dynamics? We inferred the evolutionary age of incRNAs through comparative sequence analysis of the 12 sequenced Drosophila genomes (Drosophila 12 Genomes Consortium 2007; Stark et al. 2007). Given the relatively fast evolutionary rate of incRNAs (Pang et al. 2006), we took a conservative dating strategy (see Methods). Following the parsimony principle, 23,165 incRNAs were assigned to a unique phylogenetic branch compared to out-group species. Among those, 2660 (or 11.48% of 23,165) were identified as “male-biased” in D. melanogaster during at least one comparison (Fig. 4A). By comparing the chromosomal distribution of incRNAs across two age groups, i.e., before and after the split of the melanogaster subgroup (3–6 million yr ago) (Fig. 4), we found that older male-biased incRNAs, those that originated before the split, are significantly enriched on autosomes (OR = 1.18, P = 0.037 for whole body; and OR = 1.38, P = 3.096 × 10−4 for gonad). In contrast, young male-biased incRNAs show the opposite pattern in both whole body (OR = 0.72, P = 0.05) and gonad (OR = 0.71, P = 0.03) comparisons. Furthermore, we found that a significantly larger percentage (10.86%, or 289 of 2660) of male-biased incRNAs emerged very recently (<3 million yr, on branches 5 and 6 in Fig. 4A) compared to male-biased coding genes that emerged during the same period (4.03%, or 400 of 9931) (Fisher's exact test, OR = 2.90, P < 2.2 × 10−16). A significantly negative correlation (Spearman correlation rho = 0.95, P = 0.0008) between the age of the lineages and the proportion of X-linked male-biased incRNAs was observed (Fig. 4B).
Analysis of incRNAs with different ages. (A) Numbers of newly originated incRNAs in each age branch inferred by comparative genomics analysis. For each branch, the counts of male-biased and all incRNAs are given as the underlined numbers in parentheses, separated by a slash (“/”). (B) Proportions of male-biased incRNAs among all identified male-biased transfrags in each age branch. Significant correlation between the age of the lineages and the proportion of male-biased ncRNAs was observed (Spearman correlation rho = 0.89, P = 0.01). The phylogeny and divergence times are from Stark et al. (2007).
The age analyses implemented in this study work on the DNA sequence level. That means, for sex-biased and unbiased transfrags, we can infer the age of the corresponding DNA locus based on the presence or absence of information. However, although the DNA sequence can be old, the transcription pattern may have only recently evolved. In this sense, our strategy provides an upper age estimate for the expression pattern. Since male-biased expression has a higher turnover rate, such an approximation may be too conservative, and the age of male-biased transfrags could have been overestimated. Therefore, there could be an even larger proportion of younger male-biased incRNAs, further strengthening our conclusions.
Zhang et al. (2010a) reported an excess of X-linked new protein coding genes in Drosophila that had been recently generated from DNA-level duplication or de novo gene origination, and the proportion of male-biased genes among the X-linked new genes diminishes with gene age. The incRNA genes in this study show a similar pattern to the new protein-coding genes. However, it appears that turnover is more recent in the incRNA genes, consistent with the known rapid evolution of ncRNA genes (Pang et al. 2006) and higher turnover rate of microRNA genes (Lu et al. 2008). These data indicate that different evolutionary forces, e.g., MSCI and sexual antagonism, might play roles at different evolutionary timescales (Zhang et al. 2010a).
In summary, we experimentally identified male-biased noncoding RNAs in D. melanogaster and analyzed their chromosomal distribution. The identification of a large number of incRNAs that showed male-biased expression patterns may explain the signals of natural selection previously detected in the noncoding genomic regions (Andolfatto 2005). By systematically profiling the whole transcriptome of D. melanogaster male and female adult whole bodies as well as reproductive tract organs, we revealed a long-term removal of male-biased incRNA genes from the X chromosome resulting in an uneven distribution of male-biased incRNAs between X and autosomes. This led to the long-term X chromosome demasculinization, probably through sexual antagonism and MSCI. Finally, we identified distinctive chromosomal preferences between young and old male-biased incRNAs. This pattern of male-biased incRNAs further generalizes the uneven pattern of male-biased gene content on Drosophila autosomes and X chromosomes and suggests that the pattern is shaped by natural selection acting on male functions. Our results contribute to a global picture of sex chromosome evolution in the Drosophila genome.
Methods
Data sources
Genome sequence and annotations were obtained from the UCSC Genome Browser database (http://genome.ucsc.edu; dm2). Gene model annotations were obtained from FlyBase (http://www.flybase.org; v5.46, downloaded in March, 2013). The Affymetrix array BPMAP annotation was downloaded from the Affymetrix website (http://www.affymetrix.com/products_services/arrays/specific/drosophila_tiling1_0r.affx). The 12 sequenced Drosophila genome sequences (CAF1) were downloaded from http://rana.lbl.gov/drosophila/.
Sample preparation and microarray hybridization
We extracted total RNA from whole bodies, thoraxes, digestive tracts (guts), testes, accessory glands, and ovaries from virgin Oregon R adults using the Qiagen RNeasy Mini Kit with on-column DNase digestion. All flies were virgin and aged for 1–6 d post-eclosion before being used in extractions or dissections. Whole body and thorax RNA was immediately extracted from 50 males or females. Gut tissue was completely removed from thoraxes. Testes, accessory glands, ovaries, and guts were dissected, placed in RNAlater (Qiagen), and stored at −20°C until RNA extraction. Roughly 200 testes, accessory glands or ovaries, or 100 guts were required for each replicate. Testes were separated from all other reproductive tract tissues (seminal vesicles, accessory glands, and ducts). For statistical independence, all tissues were harvested from independent sets of flies. Each of the biological replicates was labeled with GeneChip WT Sense Target Labeling and Control Reagents (Part#: 900652) and hybridized to an Affymetrix D. melanogaster genome tiling array as previously reported by Manak et al. (2006). Labeling and hybridizations were performed at the University of Chicago Functional Genomics Facility. Three replicates of hybridization were performed for each tissue.
Tiling array data processing and analysis
Affymetrix Tiling Analysis Software (TAS v1.1.02) was used to process raw tiling array data (Cheng et al. 2005; Manak et al. 2006). Raw data were normalized by quantile normalization and the median of target intensities was scaled to 100. As suggested in the Affymetrix TAS user manual (http://www.affymetrix.com/support/developer/downloads/TilingArrayTools/index.affx), each probe position was analyzed in a local smoothing window with bandwidth (BW) equal to 50 bp (resulting in a window width of 101 bp) for better statistical power. To assess the performance of replicates, standard Pearson correlation coefficients between replicates were calculated pairwise. The significant correlation (Spearman correlation rho > 0.98, P-value = 1 × 10−5) indicated reasonable consistency between replicated samples.
A one-tail Wilcoxon signed-rank test on all the probes in the window was performed with the alternative hypothesis that the true intensity difference between the perfect match (PM) probe and mismatched probe is significantly greater than zero. Only probes with a P-value < 0.1 were called “positive.” Neighboring positive probes with max-gap 50 bp and a minimum run of 90 bp were grouped as transfrags (transcribed fragments), then Unified Transfrags, or UTS, were further derived by assembling overlapped transfrags (“supporting transfrags”) in different samples as suggested by previous literature (Cheng et al. 2005; Manak et al. 2006). “Present” call was produced for each unified transfrag of UTS. A unified transfrag will be called “Present” in a given sample if and only if at least one of its supporting transfrags is identified in this sample. Moreover, the median intensity value of all comprising probe sets within each unified transfrag was calculated for each sample. Among UTS, ∼48% unified transfrags are tissue-specifically transcribed, and 78% consist of multiple supporting transfrags with ≥50% overlap. Further inspection indicated that 73.52% (68,310 of 92,916) of coding exons on autosomes (chr2L, 2R, 3L, and 3R) and X chromosome annotated in FlyBase were detected as expressed in at least one of the five D. melanogaster samples (adult male/female whole body and three reproductive organ tissues [testis, ovary and accessory gland]) with ≥70% coverage.
An integrated procedure was applied to assess sample-biased expression for each unified transfrag (Suppplemental Fig. S5). To be conservative, a global Kruskal-Wallis test (“nonparametric one-way ANOVA”) among the five samples was applied before the pairwise Mann–Whitney U-test. To assess the detection power of our procedure, we compared the identified sex-biased protein-coding transfrags to the Sebida database that integrates data derived from multiple previous high-throughput studies comparing male versus female protein-coding genes expression in D. melanogaster (Gnad and Parsch 2006). Up to 88% (613 of 695) male-biased and 84% (663 of 789) female-biased genes in the Sebida twofold high quality data set were also identified by our procedure with ≥70% length coverage, suggesting a high consistency of our procedure with previous studies.
Annotation of detected transfrags
Transfrags were classified as “coding” or “intergenic” based on their genomic coordinates. Because our work focused on noncoding genes, as a conservative estimation, we only consider a transfrag as “intergenic” if it does not overlap with any annotated FlyBase protein-coding gene models on both the sense and antisense strands. We further assessed the coding potentials of these intergenic transfrags by a SVM-based classifier (Kong et al. 2007); the results indicated that >98% intergenic transfrags are truly noncoding. After excluding 945 transfrags showing putative coding potential at either sense or antisense strand, we classified the remaining transfrags as “intergenic noncoding transfrags.” The intronic transfrags were excluded from follow-up analyses as current annotations might not distinguish true intronic noncoding RNAs from novel exons of protein-coding genes or nondegraded intronic transcripts. Moreover, in case of potential bias resulting from the relatively larger exon number in protein-coding genes (when comparing to noncoding RNAs), we re-ran analysis in gene level by assigning transfrags to annotated FlyBase genes according to the coordination.
Assessing the relationship between expression breadth and gene age for protein-coding genes
According to Meisel et al. (2012), microarray signal intensities from 14 adult D. melanogaster tissues were obtained from FlyAtlas (Chintapalli et al. 2007). Expression breadth was calculated according to the tissue specificity index, τ (Yanai et al. 2005). Genes were considered as narrowly (tissue-specific) and broadly expressed depending on their τ value (τ > 0.7 and τ ≤ 0.4, respectively). Testis-specific genes were considered to be encoded proteins in the sperm proteome if they were found in at least one of the two sperm proteomes (Dorus et al. 2006; Wasbrough et al. 2010). Gene age was obtained by crosslinking CG identifiers with information available from Zhang et al. (2010a).
Comparative genomics analysis to infer evolutionary ages along the Drosophila phylogeny
In silico comparative sequence analysis was performed with all 12 sequenced Drosophila genomes similar to the procedure reported by Sturgill et al. (2007). We ran NCBI BLAST against genomic DNA of each species. To handle the relatively low sequence conservation of noncoding genes, optimized BLAST parameters were employed as suggested in previous literatures (Korf et al. 2003; Freyhult et al. 2007). A D. melanogaster incRNA is called “absent” in another species if there are no hits with E-value < 1 × 10−4 and coverage >80% found in that species. After making the “present”/“absent” call for each incRNA, we dated their origination along the Drosophila genus phylogenetic tree (Supplemental Fig. S7) following the parsimony principle described below.
IncRNA x is assigned to branch X if and only if it is called “present” in all in-group species of branch X and “absent” in all out-group species of X. For example, branch 0 includes incRNAs that are “present” in all 12 sequenced Drosophila genomes; branch 4 includes incRNAs that are “present” in D.mel, D.sim, D.sec, D.yac, and D.ere, but “absent” in D.ana, D.pse, D.per, D.wil, D.moj, D.vir, and D.gri.
Assessment of the uneven distribution of the transfrags among X chromosome and autosomes
We applied the Fisher's exact test to assess the uneven chromosomal distribution using the estimated odds ratio (OR) as an intuitive measurement (see Table 1).
Test for chromosomal distribution of transfrags
Independent RT-PCR assay
RNA extraction
Adults were collected within 10 d after eclosion. Tissues like heads, ovaries, and testes were separately dissected into tubes with TRIzol (Invitrogen). RNAs were extracted following TRIzol reagent instructions. About 10 μg sample RNA was mixed with 20 μL RQ1 RNase-free DNase (1unit/μL; Promega), 10 μL 10× DNase buffer, 2 μL RNase inhibitor HPR1 (Takara) and DEPC water (up to 100 μL), and incubated for 3 h at 37°C. Contamination of RNAs with DNA was ruled out by PCR amplification of two pairs of primers for Gapdh2 and II171a, taking the extracted RNAs as template.
Reverse transcription
Five tenths micrograms 6-mer random primer (Takara Company) and 5 μL dNTPs (2.5 mM, Takara Company) per microgram of RNA sample were mixed in a total volume of ≤15 μL in a tube; the tube was heated for 5 min to 70°C to melt secondary structure; the tube was cooled immediately on ice for at least 3 min to prevent a secondary structure from reforming; 5 μL M-MLV 5×Reaction Buffer (Promega Company), 5 μL dNTPs (2.5 mM, Takara Company), 0.5 μL RNase Inhibitor (HPR I, Takara Company), and 1 μL M-MLV RTase (Promega Company) were added to the annealed primer/template. Then DEPC water was added up to the volume of 25 μL. PCR of primer II171a was conducted to guarantee no genomic DNA contamination, and PCR of Gapdh2 was conducted to guarantee the quality of cDNA.
Data access
The data discussed in this publication have been submitted to the NCBI Gene Expression Omnibus (GEO; http://www.ncbi.nlm.nih.gov/geo/) (Edgar et al. 2002) under accession number GSE53421.
Acknowledgments
L.W. was supported by the National Outstanding Young Investigator award from the Natural Science Foundation of China (31025014), 973 grant from the China Ministry of Science and Technology (2012CB837600), and the China Ministry of Education 111 Project (B06001). The research of G.G. was supported in part by the National Outstanding Youth Talent Initiative Program. M.L. is currently supported by NIH grants 1R01GM100768-01A1 and NSF1051826 for investigating genetics and evolution in Drosophila. M.D.V. was supported by the Pew Latin America Fellowship and the Brazilian Council of Technological and Scientific Development (CNPq). N.W.V. was partially supported by National Institutes of Health Grant T32 GM007197.
Footnotes
-
↵10 Corresponding authors
E-mail weilp{at}mail.cbi.pku.edu.cn
E-mail mlong{at}uchicago.edu
-
[Supplemental material is available for this article.]
-
Article published online before print. Article, supplemental material, and publication date are at http://www.genome.org/cgi/doi/10.1101/gr.165837.113.
Freely available online through the Genome Research Open Access option.
- Received September 1, 2013.
- Accepted December 28, 2013.
This article, published in Genome Research, is available under a Creative Commons License (Attribution-NonCommercial 3.0 Unported), as described at http://creativecommons.org/licenses/by-nc/3.0/.