Abstract
A collection of 4457 Saccharomyces cerevisiae mutants deleted for nonessential genes was screened for mutants with increased or decreased mobilization of the gypsylike retroelement Ty3. Of these, 64 exhibited increased and 66 decreased Ty3 transposition compared with the parental strain. Genes identified in this screen were grouped according to function by using GOnet software developed as part of this study. Gene clusters were related to chromatin and transcript elongation, translation and cytoplasmic RNA processing, vesicular trafficking, nuclear transport, and DNA maintenance. Sixty-six of the mutants were tested for Ty3 proteins and cDNA. Ty3 cDNA and transposition were increased in mutants affected in nuclear pore biogenesis and in a subset of mutants lacking proteins that interact physically or genetically with a replication clamp loader. Our results suggest that nuclear entry is linked mechanistically to Ty3 cDNA synthesis but that host replication factors antagonize Ty3 replication. Some of the factors we identified have been previously shown to affect Ty1 transposition and others to affect retroviral budding. Host factors, such as these, shared by distantly related Ty retroelements and retroviruses are novel candidates for antiviral targets.
The Ty retrovirus-like elements of Saccharomyces cerevisiae (Aye et al. 2001; Scholes et al. 2001; Garfinkel and Bailis 2002; Sandmeyer et al. 2002; Voytas and Boeke 2002; Aye and Sandmeyer 2003; Griffith et al. 2003) and Tf retrovirus-like elements of Schizosaccharomyces pombe (Levin 2002) have been studied in order to identify host functions that affect the retrotransposon lifecycle and, potentially, the retrovirus lifecycle. Host genes that activate or suppress viral replication represent a source of novel antiviral targets and therapeutics.
Ty elements are subject to host regulation at several levels. Ty1 and Ty3 share replication requirements for initiator tRNAMet, the primer for reverse transcription (Keeney et al. 1995) and the lariat debranching enzyme (Dbr1) (Chapman and Boeke 1991; Karst et al. 2000). Pmr1, a Mg++ transporter (Bolton et al. 2002), and Lhp1, an RNA chaperone (Aye and Sandmeyer 2003), have been implicated in Ty1 and Ty3 replication, respectively. Although the mechanism through which DNA maintenance genes affect Ty1 transposition is not clear, disruption of several of these genes is associated with increased amounts of Ty1 cDNA. These include genes encoding nucleotide excision repair components Ssl2 and Rad3 (Lee et al. 2000); RAD52 epistasis group proteins (including RAD50) (Rattray et al. 2000); and Rad27, the flap endonuclease (Sundararajan et al. 2003). The helicase Sgs1 acts to depress integration of Ty1 multimers (Bryk et al. 2001). Similar to nondividing mammalian cells, the fungal nuclear membrane does not break down during mitosis. Therefore the core particle is presumed to be remodeled to allow a preintegration complex to transit the nuclear pore. Tf1 requires Sin3, a deacetylase, and the nucleoporin Nup124 for nuclear entry (Levin 2002). Chromatin-related proteins have also been shown to affect the integration pattern of Ty elements; these include the histone E2 enzyme, Rad6, and nucleosome assembly factor, CAF-I (Voytas and Boeke 2002). In the case of the copialike element, Ty5, Sir4 interaction targets the preintegration complex to heterochromatic regions (Xie et al. 2001). Ty3 targeting to tRNA gene polymerase III initiation sites in vitro requires transcription factors TFIIIB and TFIIIC (Kirchner et al. 1995). Three high-throughput screens for Ty host factors have recently reported a large number of genes, including some of those mentioned above (Scholes et al. 2001; Griffith et al. 2003; Aye et al. 2004). In spite of the many host genes known to affect replication of Ty elements and Tf1, and the several dozen proteins identified in retrovirus virions or affecting retroviral replication (Goff 2001; Ott 2002; Pornillos et al. 2002; Dvorin and Malim 2003; Freed 2004), those studies have not yet shown overlap in the sets of host factors for anything other than very basic functions.
We report here on the results of a screen of 4457 S. cerevisiae knockout strains for mutants affected in transposition of Ty3. One hundred thirty mutants were identified that had consistent phenotypes in high- and low-copy Ty3-HIS3 transposition assays. The classes of genes that we identified suggest that trafficking functions are involved in retroelement activity as well as retroviral budding, that nuclear entry represents a control point for Ty3 cDNA synthesis, that Ty3 cDNA may be down-regulated by host replication functions, and that distantly related Ty elements and retroviruses overlap in host factor requirements.
Results and Discussion
Transposition of a high-copy, galactose-inducible Ty3-HIS3 (pNB2142) was assayed in 4457 of 4847 strains of the S. cerevisiae haploid knockout collection (Invitrogen) (Winzeler et al. 1999). The remaining strains (Supplemental Table 1) did not grow sufficiently well to be reliably tested. Each deletion strain was transformed with the high-copy Ty3-HIS3 plasmid, and duplicate independent transformant patches were assayed by two independent observers. In this assay, an active Ty3 element tagged downstream of POL3 with a copy of the HIS3 gene (Aye and Sandmeyer 2003) is induced by growth in galactose as a carbon source. Ty3-HIS3 RNA is reverse transcribed, and cDNA is transposed through integration or recombination into chromosomal DNA (Fig. 1A). Cells that have lost the plasmid but acquired a chromosomal copy of Ty3-HIS3 are identified by sequential replicaplating steps (Fig. 1B; Methods). Mutants identified in the first round of screening as having increased or decreased Ty3 transposition were rescreened in a less dense plating format. Based on mutants identified in these tests, a subset of 62 of the 4457 strains were retested for transposition phenotypes. Two hundred fifty-two mutants were identified as increased or decreased in high-copy transposition (Supplemental Table 2A). The assay is semiquantitative in this format; mutants with scores differing by one unit were estimated to be roughly twofold different from wild type in transposition frequency based on a spot dilution assay of representative mutants (data not shown). Of the 252 mutants, 24 represented 12 pairs of neighboring ORF deletion mutants, raising the possibility that deletion of neighboring coding sequence was responsible for a small subset of phenotypes (Supplemental Table 2B). In order to identify mutants in which the phenotype was independent of the high-copy Ty3-HIS3 vector, mutants identified in the original screen were transformed with low-copy Ty3-HIS3 plasmid pNB2155 and reassayed for transposition.
Screen strategy. (A) Ty3 replication. The S. cerevisiae knockout mutant collection was screened for strains that are increased or decreased for Ty3-HIS3 transposition as described in the Methods. Ty3-HIS3 was expressed by galactose induction from a high-copy, URA3-marked plasmid. Reverse transcription of the Ty3 RNA resulted in a cDNA that transposes by integration or recombination into chromosomal DNA. (B) Genetic assay of deletion mutant collection for transposition. Deletion mutants transformed with the high-copy Ty3-HIS3 plasmid were patched onto SD-Ura, -His medium (SD-UH), and replica plated as shown in the diagram. Papillations on the final selective medium are indicative of colonies derived from cells that have lost plasmids and undergone transposition. Two independent transformants were tested for each mutant. Mutants were rescreened with Ty3-HIS3 expressed from a low-copy plasmid. Up and down indicate patches of mutants with increased and decreased frequencies of transposition. (C) Distribution of Ty3 transposition phenotypes in knockout mutants using high- and low-copy Ty3-HIS3 assays. Mutants were originally screened by using a high-copy Ty3-HIS3 element (left). Mutants that displayed increased (up) and decreased (down) transposition relative to wild type were rescreened by using a low-copy plasmid (right). One hundred thirty mutants displayed consistent phenotypes in high- and low-copy Ty3-HIS3 assays. Up to down, up to wild type, down to up, and down to wild type refer to changes in transposition phenotype between the two vector systems, with the high-copy vector phenotype indicated first.
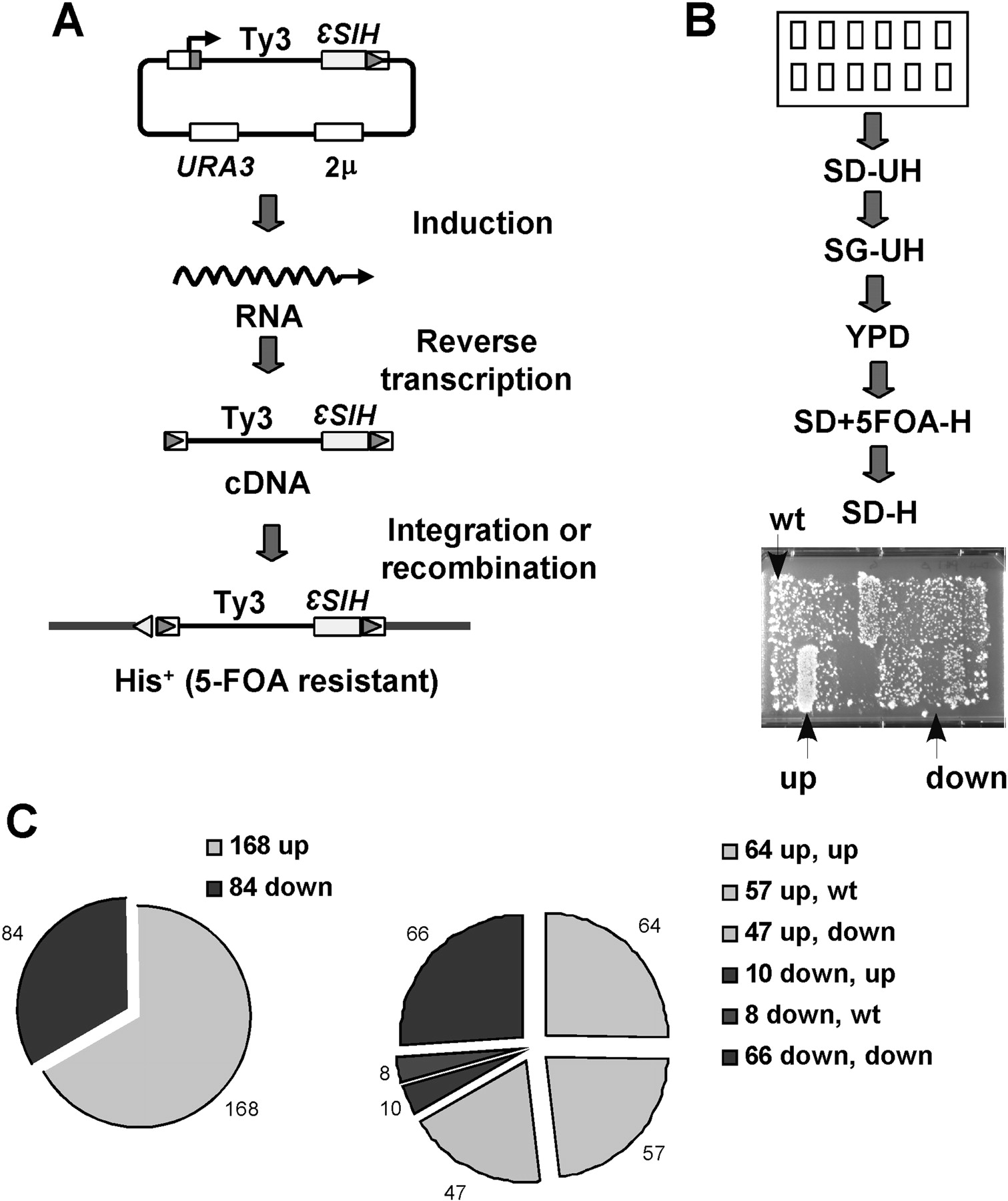
A total of 130 mutants that were affected consistently in both high- and low-copy Ty3-HIS3 transposition assays were identified (Table 1). Of the original 12 closely-spaced ORF pairs, six pairs remained after the low-copy screen. One of these pairs (MRC1) was merged by the time of the rescreen. Of the 130 mutants, 64 were increased in frequency and 66 were decreased in frequency compared with wild type (Fig. 1C). Mutants with decreased transposition in the high-copy screen retested more consistently in the low-copy screen than did mutants with increased transposition.
Distribution of functions of genes that affect Ty3 mobilization
Type | Deletion reduced transposition | Deletion enhanced transposition |
|---|---|---|
| Cell wall, membrane | FUR4, PKR1, FTR1,a FET3a | LDB7, ECM29, TIR3 |
| Chromatin modification and transcription | EAF7, IES6, CTK2, BAS1, SIN4,b,c SSN3 | HTL1, ADA2, SGF73, RSC1, BRE2, BDF1, MBP1, NGG1, GAL80,a CKB2, NCE4, SWD1b |
| DNA maintenance | RAD24 | ELG1,d MRC1, RAD6, HPR5, SGS1,d CSM3, EST3, CLN2, CLB2, BRE1, RRM3d |
| Nuclear transport | KAP120, ASM4/NUP59 | NPL3, SAC3, NUP133,c NUP84,c NUP120, NUP100 |
| RNA processing | DBP3,c DEG1, DHH1, KEM1, DBR1c | POP2c |
| Translation | ASC1, GCN20, RPL6Ac | SRO9, RPS24ba |
| Mitochondrial related | TCM62, SOD2, FLX1, ATP2, ATP18, | MRS1, FMP21 |
| Cytoskeletal | SLA1, ARP5, BUB1 | BEM2, BUD13, SRV2 |
| Secretion/vesicular trafficking/vacuolar | VAM7, ATG17,c VPH1,c VPS27, VPS28, VPS36, VPS25, SNF8/VPS22, VPS20, SNF7/VPS32, RIM20, BRO1,b VPS4, MNT4, FAB1, VPS9,c PEP7, CLC1, RIM13,b VPS51 | VAC8,d SEC22,c APL2, YEA4, YTA7, ATG2, VPS24 |
| Stress/chaperone/modification | CPR7, RUB1, BLM3 | RTT101,d TPS2,c HMF1 |
| Metabolic | DEP1,c SPO22, ARV1 | FCY22, LAG1, |
| Ty1 | RTT103,d RTT109d | |
| Other | BAR1 | |
| Unknown | YAL027W, YDL118W, YDR067C, YDR493W/FMP36, YFR012W, YGL057C, YJR120W, YNL224C, YOL083W, YPL225W, YMR158W-B | YIL090W, YLL044W,a YBL094C, YBR284W,b YHR177W, YLL030C, YLR235C, YBR277C, YDR290Wa,b,e |
a Change in transcript levels (Ideker et al. 2001) or low fitness (Giaever et al. 2002) in galactose carbon source.
b Aye et al. screen (Aye et al. 2004).
c Ty1: Griffith et al. screen (Griffith et al. 2003).
d Ty1: Scholes et al. screen (Scholes et al. 2001).
e ORF overlaps RTT103.
Sixty-six mutants were examined for Ty3 Gag3 proteins, IN and cDNA. The original screen was performed with a high-copy plasmid because Ty3 proteins expressed from a low-copy plasmid are at the limit of immunoblot detection in the knockout BY4741 strain background. Therefore, strains for Ty3 protein and cDNA analysis were transformed with pDLC201, a high-copy plasmid similar to that carrying the Ty3-HIS3 used for the original screen, but not interrupted with HIS3 (see Methods). Duplicate independent transformants were induced to express Ty3 for 24 h. At the end of that time, cells were harvested and extracts were examined by immunoblot analysis for Ty3 proteins, using anti-CA, and anti-IN. PGK was monitored as a protein loading control. Southern blot analysis was performed to quantify cDNA relative to the Ty3 plasmid. Results of these assays are summarized qualitatively in Figure 2. Data are available in Supplemental Figure 1, A through N, and Supplemental Table 3.
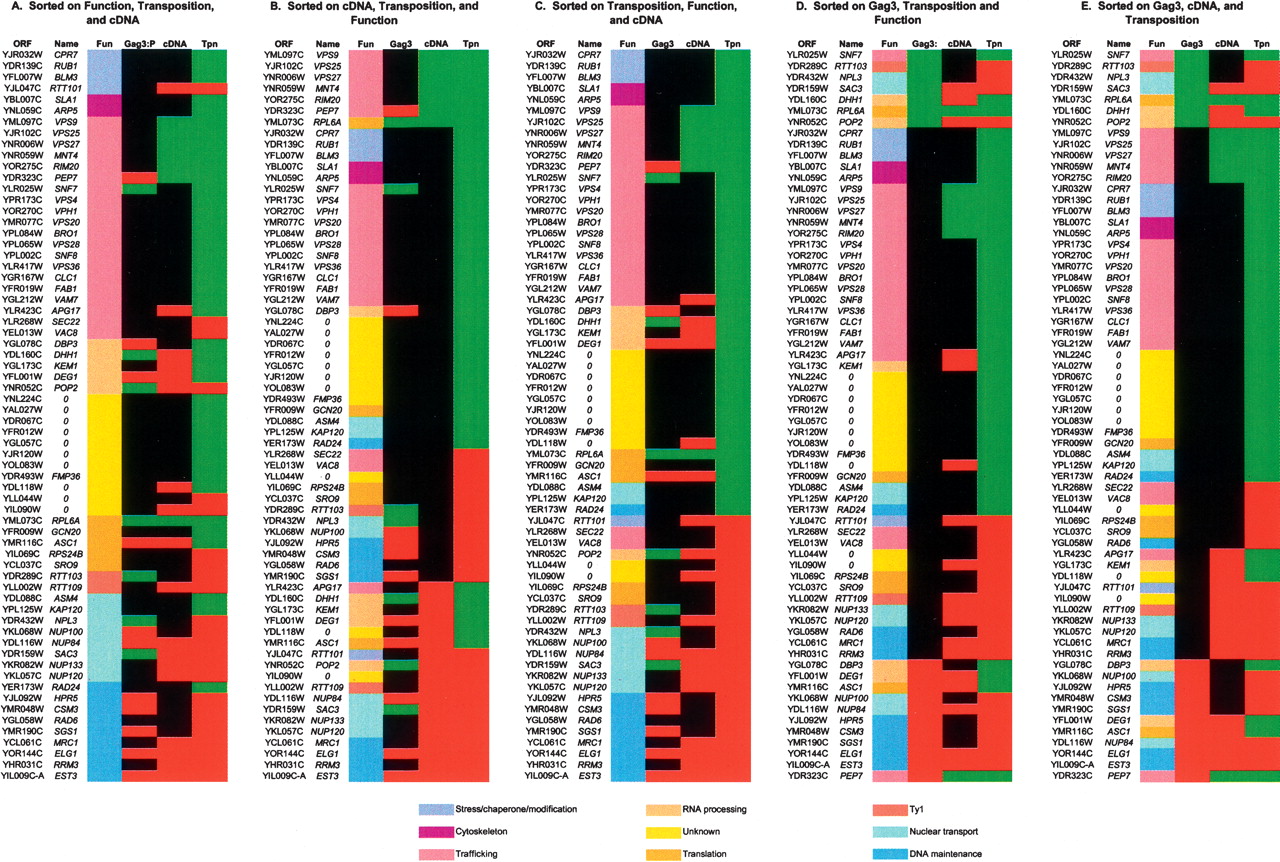
Summary of gene functions, Ty3 intermediate phenotypes, and transposition phenotypes for 66 mutants. Based on GO functions, a subset of 66 genes identified in the current Ty3 screen representing nine categories from Table 1: stress/chaperone, cytoskeletal, vesicular/vacuolar, RNA processing, unknown, translation, Ty1 transposition repressors, nuclear transport, and DNA maintenance were chosen for analysis of transposition intermediates (Fun). Transposition frequencies (Tpn) are indicated only as increased (red) or decreased (green). Gag3 was monitored by Western blot analysis, described in the Methods, of two independent transformants induced for expression of wild-type Ty3 for 24 h. Analysis used anti-CA and anti-IN (not shown in this figure). Gag3 species were considered increased (red, 4 or 5), decreased (green, 1 or 2) or the same as wild type (black, 3) on the same gel using PGK as a loading control (Supplemental Fig. 1). This analysis is qualitative rather than quantitative. Ty3 cDNA was monitored and quantified by Southern blot analysis, as described in the Methods, using a probe to the internal domain of Ty3 (Supplemental Fig. 1). The amount of Ty3 cDNA is expressed as the ratio of Ty3 cDNA to Ty3 plasmid in the same blot. The wild-type ratio was 0.57 ± 0.31. The cDNA/plasmid ratio (cDNA) was rated greater than wild type if the average ratio of two transformants was greater than wild type plus one standard deviation (red) and less than wild type if the average of two transformants was less than wild type minus one standard deviation (green). In three cases (VAM7, YOL083W, and SRO9) the measurement from one transformant was used because the second transformant did not show presence of the plasmid (Supplemental Fig. 1). Panels were sorted in Excel to show relationships of function, Gag3 protein, cDNA, and transposition frequency. Sorting priority is indicated in labels A–E. Genes are colored according to function as indicated in the legend in functional groups corresponding to those in Table 1.
GOnet analysis of strains affected for transposition
A computational approach was taken in order to identify patterns in the collection of mutants implicated in Ty3 transposition. A Web interface (GOnet) (see Methods) was developed to relate the 130 genes affecting high- and low-copy Ty3-HIS3 transposition to interacting genes identified in two-hybrid, affinity purification, and synthetic lethal assays by using publicly available information (e.g., GRID) (Allen et al. 2002; Damelin and Silver 2002; Breitkreutz et al. 2003; Bowers et al. 2004). GOnet was used to cluster genes identified in the screen and interacting genes according to GO terms (function, process, component), facilitating identification of genes representing components of the same complex or with common functions (Table 1; Supplemental Table 3). Interactions of genes were displayed by using Cytoscape (version 2.0) http://www.cytoscape.org/; Shannon et al. 2003). Groups of genes affecting chromatin modification and remodeling, mRNA export, translation and RNA processing, late endosome trafficking, and DNA maintenance emerged. Cellular complexes with involvement of multiple components included the ESCRT trafficking complexes (Fig. 3A), the nuclear pore (Fig. 3B), the P-body complex, DNA maintenance proteins (Fig. 3C), and DNA replication and repair clamp loading complexes (Fig. 3D). As indicated throughout Figure 3, there was overlap among the genes identified in the Ty3 screen and two high-throughput Ty1 screens (Scholes et al. 2001; Griffith et al. 2003).
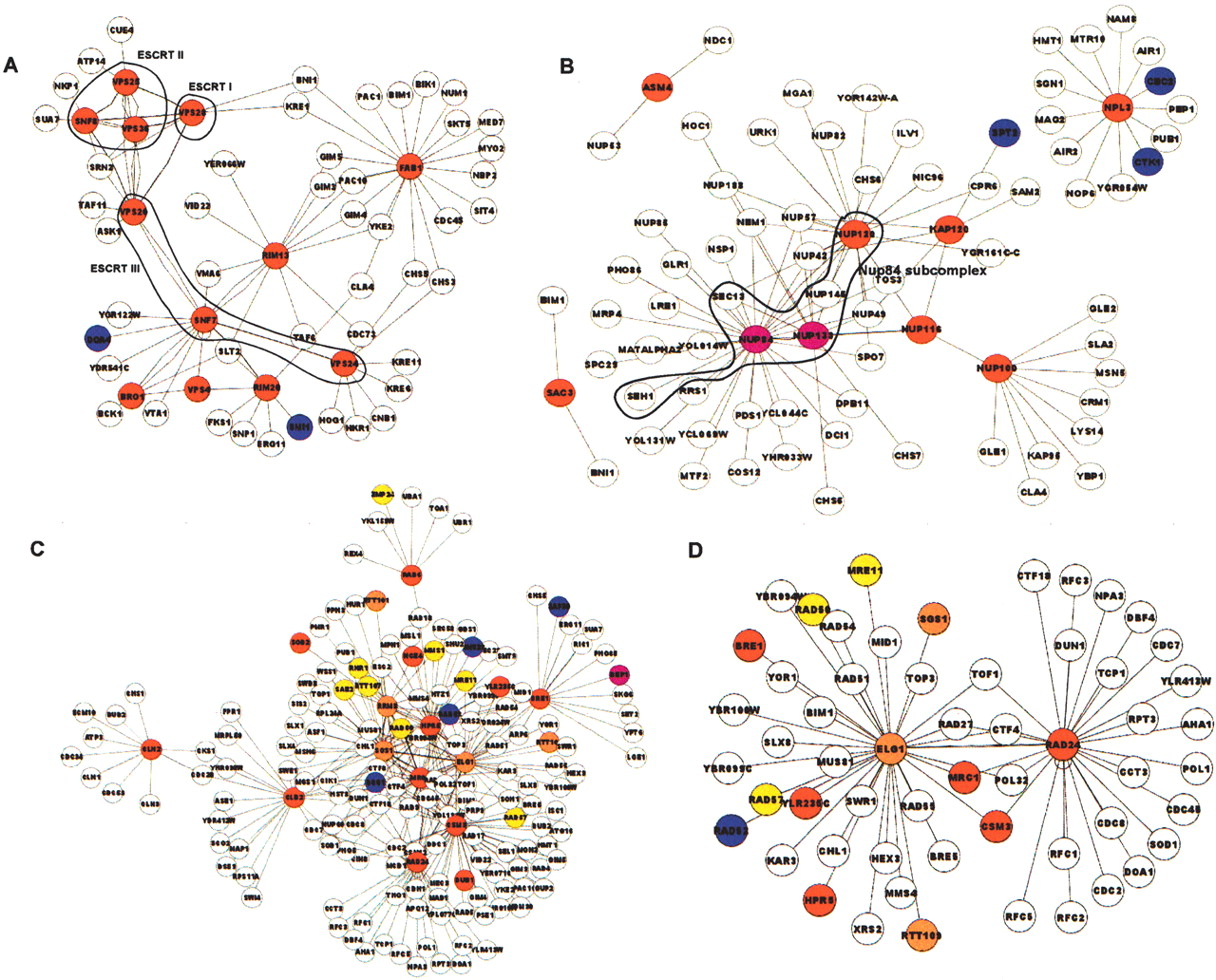
Clusters of interacting proteins implicated in Ty transposition. ORFs encoding proteins that affected transposition were entered into GOnet, and interacting protein pairs as determined by all methods (synthetic lethal, two hybrid, and affinity purification) were downloaded for display in Cytoscape. To illustrate clusters of interacting proteins in this figure, only relatively small subsets of genes were queried for interaction networks. (A) Eleven ESCRT-related protein genes (SNF8, VPS36, VPS28, VPS25, VPS20, SNF7, VPS24, VPS4, BRO1, RIM13 and RIM20); ESCRT complex genes are encircled. (B) Nuclear transport genes; Nup84 subcomplex component genes are encircled (Table 1). (C) DNA maintenance genes (Table 1). (D) ELG1 and RAD24, homologs of clamp loader subunit Rfc1. Red indicates those identified in the current Ty3 screen; blue, those identified in the Griffith et al. (2003) Ty1 screen; yellow, those identified in the Scholes et al. (2001) Ty1 screen; purple, those identified in the Griffith et al. screen and the Ty3 screen; and orange, those identified in the Scholes et al. screen and the Ty3 screen.
BLASTP batch (Altschul et al. 1997) (http://www.ncbi.nlm.nih.gov/Education/BLASTinfo/guide.html) searching of the 130 Ty3-related ORFs against human protein sequences in GenBank (automated in GOnet) showed that 87 have homologs scoring with P-values less than e–04 (Supplemental Table 4). Fifty-two of these have P-values less than e–20. Thus, a high proportion of the genes identified in the Ty3 transposition screen have meaningful homologs in the human genome.
Mutants potentially affected in expression of GAL1-10 UAS-regulated Ty3
The native Ty3 promoter is strongly induced by mating pheromones (Sandmeyer et al. 2002). In order to control Ty3 expression in growing strains, we used the GAL1-10 UAS. Because all strains were required to grow on galactose as a carbon source in order to be scored, it seemed likely that strains deficient for galactose induction would represent only a small fraction of the mutants identified. In order to test this notion, the 390 mutants that grew too poorly to be reliably tested (Supplemental Table 1) were inspected for representation of genes expected to have aberrant galactose-mediated expression of Ty3-HIS3. Ideker et al. (2001) identified 118 S. cerevisiae genes, transcripts of which significantly changed in amounts during growth on galactose as a carbon source. Of the genes identified as significantly different in expression based on a statistical maximum likelihood model and likelihood ratio test, ones deleted in mutants that were not tested in our study because of slow growth, included those for GAL7, GAL10, HIS4, HIS1, CEM1, KAP123, RPL1B, RPL11B, ADE3, HIS5, URA1, CAT5, and RPL21B. Comparison of the set of 252 deletion mutants identified in the high-copy Ty3-HIS3 screen to the 118 genes identified by Ideker et al. (2001) showed that only GAL2, RPL14A, YLL044W, RPS24B, and GAL80 genes were represented. Of these, only deletions of YLL044W, RPS24B, and GAL80, which caused increased transposition, tested consistently in high- and low-copy Ty3 assays (Table 1). GAL80 encodes a negative regulator of galactose metabolism.
Many nonessential genes that are important for growth on galactose as a carbon source are not induced (Giaever et al. 2002). Of the 390 strains that grew too slowly to be assayed in our screen, 14 were identified in a genomewide functional genomic assay to be attenuated for growth in galactose (deletion mutants representing GAL7, GAL10, GAL3, YLR202C, IRA2, YOR305W, GAL4, RIP1, CBP2, YHR116W, MRPS16, COX10, GAL4, and PET494) (Giaever et al. 2002). Comparison of the 252 genes identified in the high-copy Ty3-HIS3 assay showed that six (GAL2, YML090W, GEF1, YDR290W, FTR1, and FET3) had fitness scores >20, indicating significant deficiencies. Of these, only three (YDR290W, FTR1, and FET3) were retained in the final set of 130 Ty3 mutants that affect transposition in both high- and low-copy assays (Table 1). In summary, of the genes reported in Table 1, six (YLL044W, RPS24B, GAL80, YDR290W, FTR1, and FET3) have been demonstrated to either be regulated by growth on galactose or be required for competitive growth in galactose.
Chromatin- and transcription-related proteins
Deletions resulting in decreased transposition included those for genes that encode members of the Nu4A HAT complex (EAF7), INO80 remodeling complex (IES6 and ARP5), and mediator complex (SIN4). Deletions resulting in increased transposition included genes encoding members of the SAGA HAT complex (ADA2, SGF73, and NGG1) (Sterner and Berger 2000; Orphanides and Reinberg 2002), RSC remodeling complex (RSC1 and HTL1), COMPASS histone methyltransferase (BRE2), and a bromodomain protein (BDF1). Preliminary experiments suggested that several of these did not grossly affect Ty3 protein expression (data not shown). However, because Ty3 expression relied upon the GAL1 UAS, additional experiments will be required to determine if effects of these mutations are Ty3 specific. This class of proteins could affect initiation, or elongation or targeting of integration late in the lifecycle.
Transposition of Ty3 increased in three mutants with deletions of genes encoding mRNA elongation and transport factors (SAC3, NPL3, and NUP133) (Fig. 2; Table 1). SAC3 encodes a member of the TREX complex involved in coupling transcription elongation to mRNA export (Fischer et al. 2002; Jimeno et al. 2002; Strasser et al. 2002). Deletion of THP2, which encodes another member of the TREX complex, caused elevated transposition of Ty3-HIS3 expressed from the high-copy plasmid, but not the low-copy plasmid (Supplemental Table 2). TREX acts downstream of the promoter and does not affect initiation from the GAL1 promoter (Chavez et al. 2000). However, disruption of SAC3 and THP2 makes the transcribed region hyperrecombinogenic (Chavez et al. 2000; Gallardo et al. 2003). Elongation is linked to mRNA export through a network of proteins such that disruption of export can also interfere with elongation. NPL3 encodes another component of the mRNA export machinery that is associated with the transcription apparatus (Lei et al. 2001), and NUP133 encodes a nucleoporin required for wild-type mRNA export (Doye et al. 1994). We hypothesize that defects in those functions could activate recombination between the transcribed region of the Ty3-HIS3 plasmid and chromosomal Ty3 sequences, potentially resulting in elevated frequency of His+ cells in NPL3, SAC3, and NUP133 deletion mutants. Consistent with disruption of transcription elongation, cells deleted for NPL3 and SAC3 were decreased for Ty3 Gag3 and its derivatives as well as IN (Fig. 2; Supplemental Fig. 1A). However, surprisingly, the ratio of cDNA to plasmid, as well as the amount of cDNA, was increased in one of two NPL3 deletion mutant transformants, in both SAC3 deletion mutant transformants, and in both NUP133 deletion mutant transformants (Supplemental Fig. 1A,B). The phenotypes of these mutants and some other mutants suggested that Ty3 protein levels per se are not limiting and in some cases even vary inversely with cDNA levels or cDNA to plasmid ratios. Thus, either increased recombination of the plasmid-borne Ty3, stimulated by disruption of elongation and followed by chromosomal recombination, or increased recombination or integration of Ty3 cDNA due to increased amounts of cDNA could have caused the transposition phenotype. SAC3 and NUP133 encode components of the nuclear pore. Those deletion mutants exhibited greater elevation of cDNA. The phenotype of those mutants could also be related to disruption of the nuclear pore (see below).
Strains with deletions of THP2 and NUP133 were isolated in the Griffith et al. (2003) Ty1 screen, and NUP133 deletion increased Ty1 cDNA levels. However, both were decreased for Ty1 transposition. Ty1 transposition depends upon multiple elongator components (Griffith et al. 2003). The qualitative difference in effects of this deletion on Ty3 and Ty1 transposition could be partially reconciled if, in the case of Ty1, disruption of elongation did not activate recombination of the transcribed region.
Translation and mRNA turnover
Genes encoding translation initiation and cytoplasmic mRNA processing proteins were identified in the Ty3 screen. Deletion of genes for rRNA processing proteins (DEG1, DBP3), a translation initiation factor (ASC1), and a regulator of translation initiation (GCN20) decreased transposition without a similar effect on Ty3 protein or cDNA levels (Fig. 2; Supplemental Fig. 1C). Thus, it is not clear how these genes would affect Ty3 transposition if they act directly. Deletion of the gene encoding a ribosomal protein (RPL6A) decreased accumulation of Ty3 proteins and cDNA and decreased transposition. Double-stranded killer RNA of S. cerevisiae and positive-strand animal viruses are sensitive to deletions of redundant ribosomal protein genes (Ohtake and Wickner 1995; Kushner et al. 2003). Such deletions disrupt ribosome morphogenesis and can affect translation initiation efficiency. In addition, although tRNA gene–associated Ty3 LTRs do not appear to colocalize with 5S genes (Lin et al. 2001), at least some Ty3 and Ty1 targets (5S and tRNA genes) are likely to be located in the nucleolus (Bertrand et al. 1998; Thompson et al. 2003). If nucleolar structure were disrupted in a subset of these mutants, it could reduce transposition. Deletion mutants of several translation-related genes, including RPL6A and DBP3, were also identified as mutants with decreased frequency of Ty1 transposition (Griffith et al. 2003).
Translation factors and mRNA deadenylation and decapping factors and nucleases compete for access to cytoplasmic RNAs. Three strains with deletions of genes related to RNA deadenylation, decapping, and degradation (POP2, DHH1, KEM1) were affected in Ty3 transposition (Table 1). Strains deleted for POP2 and DHH1 had reduced levels of Ty3 protein and Ty3 plasmid (Fig. 2; Supplemental Fig. 1D). Disruption of the cytosolic Ccr4/NOT deadenylation 3′-5′ exonuclease complex (Decker and Parker 2002) by deletion of POP2 increased the ratio of cDNA to plasmid and transposition. Deletion of CCR4 also increased high-copy Ty3-HIS3 transposition (Supplemental Table 2). Deletion of DHH1 and KEM1, which encode a protein that couples deadenylation with decapping and the 5′ to 3′ cytoplasmic exonuclease, respectively, decreased transposition. Both of these factors localize to the P body decapping and degradation complex (Sheth and Parker 2003). Amounts of Ty3 Gag3, CA, and IN were slightly decreased in the DHH1 deletion mutant. However, in the case of the KEM1 deletion, levels of Gag3 proteins were not decreased (Fig. 2; Supplemental Fig. 1D). Although Gag3 processing in the case of one KEM1 deletion transformant was decreased dramatically, this result was not observed with three other transformants (data not shown). Amounts of IN were also decreased in the KEM1 mutant (Supplemental Fig. 1D), which would be consistent with disrupted protein processing. These experiments indicated that at least one RNA species involved in Ty3 transposition is likely to be sensitive to disruption of cytoplasmic processing as transposition decreased in both the DHH1 and KEM1 deletion mutants. However, since the ratio of cDNA to plasmid was not dramatically affected compared with wild type, it is not clear whether the effect of disruption of RNA processing on transposition is direct. Deletions of POP2 and two related genes, LSM1 and PAT1, which encode components of the decapping complex, decreased Ty1 transposition (Griffith et al. 2003). Genes related to RNA export and processing could act by affecting the equilibrium between messenger and genomic populations of RNAs.
Intracellular trafficking
One of the largest classes of mutants identified in the Ty3 screen were those defective in vesicular trafficking and vacuolar inheritance (Fig. 2; Table 1; Supplemental Tables 2, 3; Supplemental Fig. 1E–H). In the endosomal pathway, cargoes from the cell surface and from the Golgi are trafficked through vesicular budding and fusion (Katzmann et al. 2002; Horak 2003; Bonifacino and Glick 2004; Burri and Lithgow 2004). Ultimately, in a process that requires ESCRT complexes I, II, and III, these molecular cargoes bud from the limiting membrane into the lumen of the MVB. The MVB subsequently fuses with the vacuole. A number of trafficking genes, including genes encoding a clathrin coat component (CLC1); RAB5 GTPase homolog guanine nucleotide exchange factor (VPS9); homolog of the mammalian fusion facilitator EEA1 for Golgi to vacuole/lysosome trafficking (PEP7/VAC1); ESCRT cargo adapter (VPS27/HRS1); ESCRT-I subunit (VPS28); ESCRT-II subunits (VPS22, VPS25, VPS36); ESCRT-III subunits (VPS20, VPS24 and SNF7/VPS32); the AAA ATPase that recycles ESCRT complexes (VPS4); homologous factors that interact with Snf7 and Vps4 ESCRT components (BRO1/AIP1 and RIM20) (Gavin et al. 2002; Odorizzi et al. 2003; Bowers et al. 2004); the vacuolar Qc SNARE (VAM7); and the regulator of autophagy (ATG17) were all required for wild-type levels of transposition (Table 1). These genes have interrelated functions, and a number encode interacting proteins. Pep7/Vac1 is required for vesicular traffic to the vacuole, vacuolar inheritance (Weisman and Wickner 1992), and vacuolar acidic pH (Preston et al. 1992). It interacts with the RAB5 homolog Vps21, Sec1 homolog Vps45, and t-SNARE Pep12, and this complex is required for Golgi to endosome sorting (Burd et al. 1997; Webb et al. 1997; Tall et al. 1999). A mutant deleted for VPS9, which encodes a ubiquitylated guanine nucleotide exchange factor for one member of this complex, Vps21 (Hama et al. 1999), also showed decreased Ty3 transposition frequency (Table 1). Proteins are targeted to the endosomal pathway through interactions of a subset of proteins with specific phosphoinositides. Deletion of FAB1, which encodes the kinase that phosphorylates PtdIns(3)P to produce PtdIns(3,5)P2, also reduced Ty3 transposition. This activity is required for MVB formation (Odorizzi et al. 1998). Application of GOnet showed interactions among subsets of these genes. For example, genes encoding members of the three ESCRT complexes and associated proteins (Babst et al. 2002a,b; Katzmann et al. 2002; Odorizzi et al. 2003; von Schwedler et al. 2003; Bowers et al. 2004;) constituted a cluster of interacting genes in our screen (Fig. 3A).
Examination of the pattern of Gag3, p27, and CA and IN proteins produced in 20 trafficking mutants showed that most mutants were not affected in protein expression (Fig. 2; Supplemental Fig. 1F–H; Supplemental Table 3). Nonetheless, several mutants had distinctive phenotypes. The most striking phenotype was that of the mutant deleted for PEP7/VAC1. Deletion of PEP7/VAC1 decreased transposition and increased amounts of Ty3 Gag3 and IN. This increase was not associated with a comparable increase in cDNA. However, out of the 20 trafficking mutants decreased for transposition that were examined by Southern analysis, seven had ratios of cDNA to plasmid that were more than one standard deviation below wild type. These results suggest that the effect of trafficking disruption occurs relatively late in the Ty3 lifecycle.
Vesicular trafficking proteins and membranes are important for replication of many different types of viruses. Moloney murine leukemia virus genomic RNA, together with Gag and Gag-Pol, is trafficked through the endosomal system to the plasma membrane (Basyuk et al. 2003), and Type-D retroviruses, which assemble at perinuclear sites, use endosomal trafficking to direct intracellular core particles to the cell surface for release (Sfakianos and Hunter 2003). In addition, ESCRT functions identified in the Ty3 search are required for late stages of retroviral budding and for intracellular virion formation in some cell types (Kikonyogo et al. 2001; Gottwein et al. 2003; Strack et al. 2003; von Schwedler et al. 2003; Freed 2004). If trafficking proteins required for wild-type levels of Ty3 transposition act directly, we hypothesize that they could (1) promote delivery of excess Ty3 protein to the vacuole and prevent Ty3 expression from overwhelming normal cellular functions, (2) provide a cellular microenvironment that promotes VLP maturation—either assembly or uncoating, or (3) traffic Ty3 intermediates within the cell (Fig. 4A). The absence of a consistent phenotype throughout the set of trafficking mutants suggests that more than one process may be affected. Thus, the PEP7/VAC1 deletion mutant could be defective for degradation of Ty3 proteins, and the subset of mutants with modestly reduced cDNAs could be affected for some later stage of the lifecycle, such as nuclear delivery. Deletion of RIM20 produced the lowest cDNA/plasmid ratio of any mutant. Rim20 acts together with a subset of ESCRT proteins in activation of the Rim101 pH adaptation transcription factor, which is unrelated to MVB formation (Xu et al. 2004). However, Ty3 transposition was decreased by deletion of VPS27 and VPS4 which does not affect Rim101 activation. Therefore it seems unlikely that the effect of ESCRT complex disruption on Ty3 transposition is via disruption of the Rim101 pathway.
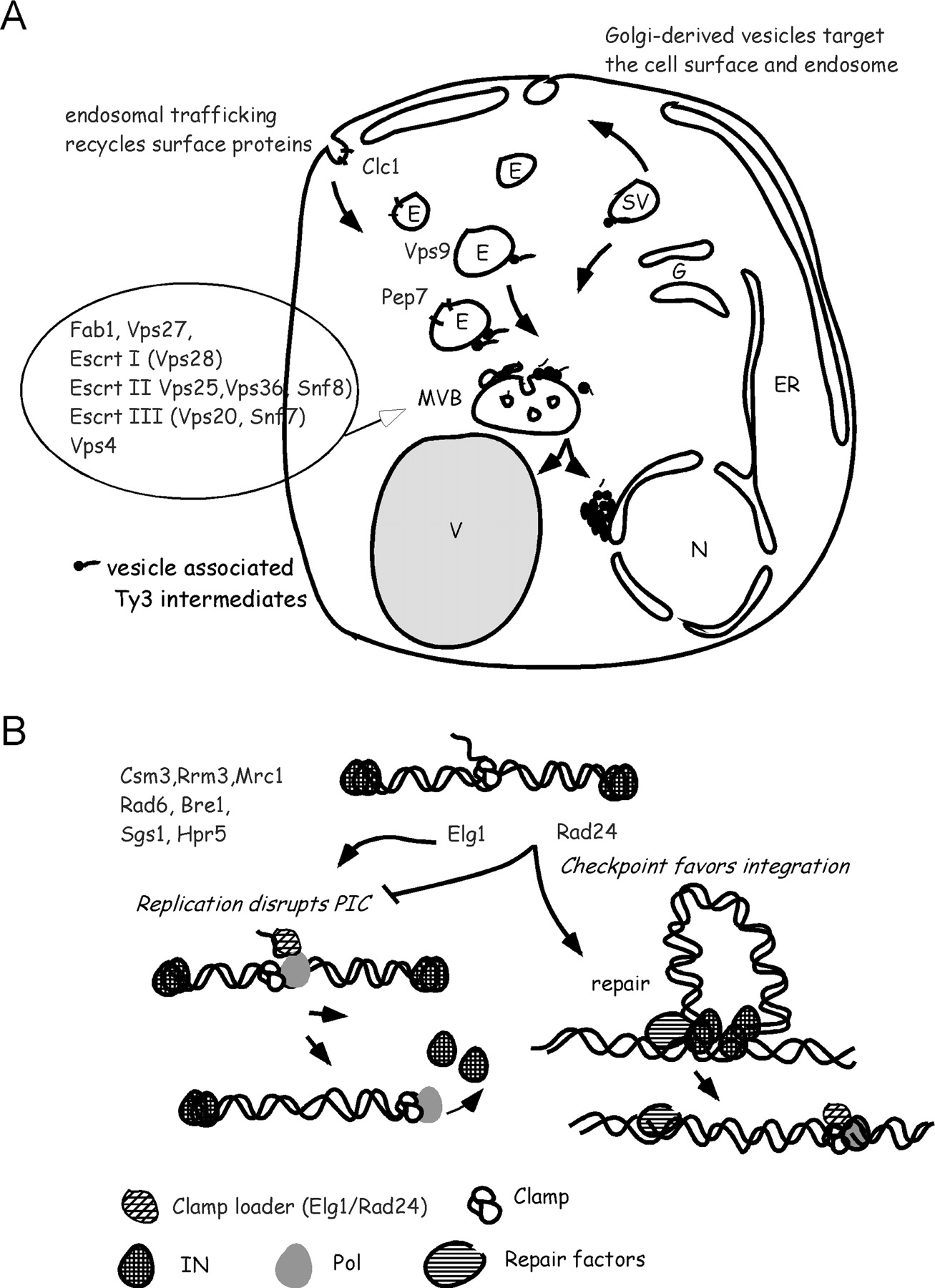
Hypothetical roles of gene function clusters in the Ty3 lifecycle. Genes identified in this screen for Ty3 mutants are shown. (A) Galactose-induced Ty3-HIS3 transposition depends upon trafficking functions. Ty3 VLPs form perinuclear clusters. Accumulation of VLPs in clusters is postulated to be directly or indirectly dependent upon an intact endosomal trafficking system. Disruption of this localization could interfere with nuclear delivery or uncoating. In this illustration MVBs are shown with associated ESCRT complexes, which mediate intralumenal vesicle budding, but ESCRT complexes could function separately from those specialized vesicles in facilitating Ty3 VLP formation or delivery. N indicates nucleus; ER, endoplasmic reticulum; G, Golgi; SV, secretory vesicle; E, endosome; MVB, multivesicular body; and V, vacuole. See text for details. (B) Ty3 transposition depends upon Rad24 function but is disrupted by proteins associated with the lagging strand clamp-loader, Elg1. Ty3 preintegration complexes entering the nucleus are postulated to distribute between disassembly by replication clamp-mediated activity and integration facilitated by repair. This competition is postulated to be influenced by the relative activity of Elg1 replication clamp-loading and Rad24 checkpoint and repair clamp loading activities.
The screen for disruption mutants affecting Ty3 also identified BRO1, RIM13, and VPS51/67 mutants as decreased in transposition (Aye et al. 2004). The majority of the vesicular trafficking genes identified in our study were not recovered in the large-scale Ty1 screens. However, Scholes et al. (2001) reported a modest increase in Ty1 transposition frequency in a VAC8 disruption mutant, and Griffith et al. (2003) described decreases in transposition in ATG17, SEC22, and VPS9 deletion strains. The relative absence of overlap in this category of genes could indicate that Ty1 and Ty3 are different with respect to the function of this pathway or that technical differences among the screens resulted in differential recovery of this class of mutants.
Nuclear pore functions
Because the fungal nuclear envelope does not break down during metaphase, as does mammalian nuclear envelope, the nuclear pore is likely to be intimately involved in Ty nuclear entry. Consistent with this, our analysis uncovered a number of genes related to nuclear transport and nucleoporins (Fig. 2; Table 1; Supplemental Tables 2, 3). The nuclear pore in yeast is comprised of ∼30 proteins, including FG and non-FG nucleoporins and membrane proteins (Macara 2001). Vertebrate nuclear pores have been shown to transport colloidal gold particles of 26 (Dworetzky et al. 1988) to 39 nm in diameter (Pante and Kann 2002) coated with karyophilic proteins. It is highly likely that the 40- to 50-nm size of the Ty3 VLP exceeds the transport capacity of yeast nuclear pore. Thus, Ty3 VLPs are probably reconfigured or uncoated prior to nuclear entry, as is also presumed for retroviral core particles in nondividing cells (Greene and Peterlin 2002; Dvorin and Malim 2003). Although a NLS required for Ty3 transposition was identified in the carboxyl-terminal domain of IN (Lin et al. 2001), Ty3 Gag3 expressed alone forms perinuclear clusters (M. Zhang, N. Beliakova-Bethell, and S.B. Sandmeyer, unpubl.), indicating that some direct interaction between Gag3 and the nuclear pore might also occur. One possibility is that Ty3 nuclear entry occurs in two stages, with contact mediated by Gag3 and entry by IN. In the case of Tf1, another gypsylike element, the Gag-like protein contains a nuclear localization signal and requires the FG nucleoporin Nup124 for nuclear entry (Dang and Levin 2000).
Genes for mRNA export proteins (SAC3 and NPL3), a nuclear karyopherin (KAP120), FG nucleoporins (NUP159, NUP59/ASM4, and NUP100), and non-FG Nup84 subcomplex members (NUP84, NUP133, and NUP120) (Siniossoglou et al. 2000; Allen et al. 2002) were identified in the Ty3 screen (Fig. 2; Table 1; Supplemental Tables 2, 3).
A subset of nucleoporins (Nup159, Nup42, and Nup82) are asymmetrically located on the cytoplasmic face of the nuclear pore, and two additional porins (Nup116 and Nup100) have a distribution biased toward the cytoplasmic side (Rout et al. 2000). In our disruption screen for Ty3 mutants, NUP159 and NUP116 were associated with increased and decreased Ty3 transposition, respectively (Aye et al. 2004), and in this screen, a mutant for NUP100 was identified with increased transposition. If they act directly, FG nucleoporins Nup159 and Nup100 might simply impede progress of the Ty3 preintegration complex through the pore channel. Deletion of KAP120, which encodes a karyopherin implicated in a late stage of 60S ribosomal subunit export (Stage-Zimmermann et al. 2000), and ASM4/NUP59, which encodes a nucleoporin symmetrically located on the nuclear and cytoplasmic faces of the pore (Rout et al. 2000; Allen et al. 2001), decreased transposition of Ty3 without dramatically decreasing cDNA/plasmid ratios, suggesting that the effects are late in the Ty3 lifecycle (Fig. 2; Supplemental Fig. 1).
The Nup84 complex (Nup84, Nup85, Nup120, Nup145, Nup133, Seh1, and Sec 13) comprises a non-FG substructure of the pore and plays an important role in nuclear biogenesis (Siniossoglou et al. 2000; Allen et al. 2002). Deletion of NUP133 or NUP120 causes clustering of nuclear pores (Doye et al. 1994; Aitchison et al. 1995; Siniossoglou et al. 2000). Deletion of each of three genes, NUP84, NUP133, and NUP120, resulted in elevated transposition frequencies and cDNA/plasmid ratios (Fig. 2; Fig. 3B; Supplemental Fig. 1A,B). Deletion of NUP84 caused the most dramatic increase in cDNA of any mutant studied. The simplest interpretation of these results is that Ty3 nuclear entry is enhanced either by disruption of the nuclear pore structure or by physical clustering of pores. Disruption of nuclear pore structure in these mutants might lead to relaxation of the pore structure and increased nuclear access for Ty3 intermediates. Ty3 VLPs accumulate in clusters (Hansen et al. 1992; data not shown). An alternative possibility is that association of Ty3 clusters with nuclear pore clusters has a synergistic effect on nuclear entry. The increase in Ty3 cDNA levels in these mutants is intriguing. If it is the case that these nuclear pore mutants have elevated Ty3 nuclear entry, it indicates that nuclear localization positively affects cDNA synthesis and may even constitute an important aspect of reverse transcription. It is possible that either the process of nuclear entry itself, for example, if it is linked to uncoating, or the nuclear environment rich in dNTPs promotes cDNA synthesis. In the case of Ty1, both NUP84 and NUP133 deletions decreased transposition; nonetheless, the NUP133 deletion mutant had elevated levels of cDNA (Griffith et al. 2003).
DNA maintenance functions
DNA maintenance proteins represented a major class of Ty3-related factors (Figs. 2, 3C; Table 1; Supplemental Tables 2, 3). Underscoring the significance of the DNA maintenance class of mutants is the prior implication of these genes in stability and integration of retrovirus and Ty1 cDNA. Processivity of replication and repair polymerases is enhanced by a sliding clamp mechanism (Maga and Hubscher 2003; Majka and Burgers 2004). The Rfc1-5 complex loads the replication processivity clamp Pol30/PCNA. This clamp also provides a platform for repair- and replication-related factors. GOnet analysis of the genes identified in our screen highlighted a cluster of clamp-related genes (Fig. 3D), including two Rfc1-related proteins, Rad24 (hRad17) and Elg1. Rad24, in addition to its checkpoint function, substitutes for Rfc1 in the repair clamp loading complex. Deletion of RAD24 decreased Ty3 transposition, indicating that Rad24 plays a positive role in transposition, either through the checkpoint (for example by extending a cell cycle period with high nucleotides and blocking cellular replication activity) or more directly through DNA repair processes, perhaps subsequent to integration.
Deletions of genes for nine DNA maintenance-related proteins, including (ELG1/RTT110) (Bellaoui et al. 2003; Ben-Aroya et al. 2003; Kanellis et al. 2003), MRC1 (Osborn and Elledge 2003), RAD6 (Hoege et al. 2002; Giannattasio et al. 2005), HPR5, SGS1 (Fabre et al. 2003), CSM3 (Mayer et al. 2004), EST3 (Hughes et al. 2000), BRE1 (Hwang et al. 2003; Giannattasio et al. 2005), and RRM3 (Schmidt et al. 2002) increased transposition. Deletions of ELG1/RTT110, HPR5, SGS1, CSM3, and EST3 increased Ty3 protein, and deletions of ELG1, MRC1, EST3, and RRM3 correlated with particularly high cDNA and elevated cDNA:plasmid ratios. Among these genes, ELG1/RTT110 is particularly interesting as it encodes the homolog of Rfc1 and Rad24 that is proposed to replace Rfc1 in the clamp loader for lagging strand replication (Bellaoui et al. 2003; Ben-Aroya et al. 2003; Kanellis et al. 2003). It was also identified in the Scholes et al. (2001) screen. Many of these gene products interact physically or genetically with Elg1 (Fig. 3D), and Hpr5, Sgs1, and Rrm3 have related replication helicase activities (Schmidt and Kolodner 2004).
Deletion of EST3, a component of telomerase, elevated Ty3 transposition. Disruption of functions that triggered the telomere checkpoint correlated strongly with enhanced Ty1 transposition (Scholes et al. 2003). Strains with deletions of RTT107, RTT103, and RTT109, which were identified in our high-copy screen as having increased Ty3 transposition frequency, were among the mutants increased in Ty1 transposition identified in the Scholes et al. (2001) screen, in which mutants in DNA maintenance factors were highly represented. The RTT107 deletion mutant phenotype was specific for high-copy transposition (Supplemental Table 2). Rtt103 is part of an exonuclease complex that enhances RNA polymerase II transcription termination downstream of poly(A) sites (Kim et al. 2004). The RTT103 deletion mutant had reduced levels of Ty3 protein and so might act similarly to NPL3 or SAC3 deletions. As reported for the effect of RTT109 disruption on Ty1, RTT109 deletion elevated Ty3 cDNA (Fig. 2; Supplemental Fig. 1I). Rtt109 is nuclear and has an RNA expression pattern similar to that of Cln2 (Sundin et al. 2004), but its function is not known. The mechanism through which DNA maintenance functions affect Ty1 cDNA is not completely understood but, given the common proteins and similar phenotypes, is likely to operate in the case of Ty3 as well. Increased cDNA production regulated by checkpoint activation (Scholes et al. 2003) and increased cDNA stability (Rattray et al. 2000) have been proposed as possible mechanisms in the case of rad50. In support of the former model, RAD24 deletion mutants are decreased for transposition, and RAD24 disruption attenuates the increased transposition of rad50 mutants (Scholes et al. 2003). However, other results suggest that direct interaction of cellular DNA maintenance proteins with incoming cDNAs do occur. For example, disruption of SGS1 increased multimerization of the Ty1 genome, indicating that cellular helicases might interact with Ty cDNA and antagonize its recombination. In retrovirus-infected cells, nonhomologous end joining factors circularize extrachromosomal cDNA (Li et al. 2001). Both checkpoint ATR and Ku repair proteins have been implicated in the retrovirus integration reaction (Daniel et al. 1999, 2003). Retroviral preintegration complex cDNA is not internally protected by IN (Miller et al. 1997) and contains gaps and displaced single-stranded regions (Yoder and Bushman 2000). Thus, nascent cDNA is a potential target for single-stranded binding proteins and associated replication and repair proteins. The fact that multiple genes for Pol30/PCNA-interacting proteins were identified as mutants with increased transposition in our screen and in a Ty1 screen (Scholes et al. 2001) suggests that replication-related activities could disrupt completion of reverse transcription and integration of the nascent cDNA, perhaps by active displacement of Ty proteins. This might destabilize the cDNA as proposed by Garfinkel and colleagues for Rad52 epistasis group and other repair-recombination factors (Rattray et al. 2000). Replication-based destabilization of the preintegration complex might be antagonized by the Rad24 replication checkpoint, consistent with the proposal of Scholes et al. (2003). In summary, we propose that completion of cDNA synthesis is a key control point, and that replication and integration-promoting repair proteins compete for access to the nascent cDNA (Fig. 4B).
Individual genes
Although no definitive clusters of cytoskeletal or chaperone genes emerged in our screen, there were individual mutants of potential interest (Supplemental Tables 2, 3). A strain deleted for SLA1, which encodes a component of the polarisome required for polarization of actin filaments along which vesicular, mRNA, and organelle trafficking occurs (Pruyne and Bretscher 2000), was among the mutants most severely decreased for transposition. Ty3 protein and cDNA levels were not grossly different from those in wild-type strains (Fig. 2; Supplemental Fig. 1K).
Deletion of genes for a prolyl isomerase (CPR7) (Duina et al. 1996), a ubiquitin-like protein (RUB1); a cullin found conjugated to Rub1 (RTT101) (Laplaza et al. 2004); and a subunit of the nuclear proteasome (BLM3) (Fehlker et al. 2003) decreased Ty3 transposition (Fig. 3; Table 1; Supplemental Fig. 1L; Supplemental Tables 2, 3). Similar to retroviruses, Ty3 contains a domain with a high percentage of proline residues. The potential role of Cpr7 in transposition is of interest because cyclophilin A, which is also a prolyl isomerase, enhances replication of HIV (Towers et al. 2003). Deletion of GIM3, which encodes one of the six prefoldin subunits (Cowan and Lewis 2002), decreased transposition in the case of the high-copy, but not the low-copy, Ty3 (Supplemental Table 2). Prefoldins deliver actin and tubulin to the TRiC complex for folding and assembly. The Hunter laboratory has implicated TRiC in perinuclear assembly of Type-D retroviruses (Hong et al. 2001). We speculate that prefoldins could act as chaperones for Ty3 assembly or uncoating. TRiC components are essential, and so TRiC involvement in Ty3 assemby was not addressed by our screen. Deletion of PFD1, which encodes another of the prefoldin subunits, decreased Ty1 transposition (Griffith et al. 2003), suggesting that these proteins have a general role in retrotransposition.
Genes of unknown function
Twenty genes of unknown function were identified in our Ty3 screen (Table 1; Supplemental Tables 2, 3). Eleven mutants representing genes that appeared to be required for transposition or that were strongly acting were chosen for Ty3 protein and cDNA analysis (Fig. 2; Supplemental Fig. 1M,N). Of these, the mutant deleted for YJR120W showed reduced Ty3 transposition and amounts of Ty3 Gag3 and IN, but wild-type ratios of cDNA to plasmid. The mutant deleted for YDL118W was also decreased for transposition, but displayed increased ratios of Ty3 cDNA to plasmid. The mutant deleted for YIL090W displayed both increased transposition and Ty3 cDNA.
Relationship of deletion mutations to Ty3 transposition
We observed differences in Ty3 intermediates between wild type and some, but not all, mutants. This was also the case in the other Ty3 screen and in the two Ty1 screens that monitored Ty1 cDNA. There are several potential explanations. First, there were differences in growth conditions used for sampling in genetic screens and biochemical assays (solid vs. liquid medium and colonies vs. early stationary phase liquid cultures). Second, Ty3 proteins were highly expressed. The current screen was initially performed with a high-copy plasmid because in the BY4741 background, it is difficult to detect Ty3 proteins and cDNA expressed from a low-copy plasmid, making it impossible to analyze mutants expressing the low-copy plasmid biochemically. Ty3 protein might not be limiting for transposition in the high-copy system. In contrast cDNA/plasmid ratios correlated with transposition in a number of mutants, particularly those with increased transposition, suggesting that cDNA might be the limiting intermediate under conditions of the high-copy screen.
What proportion of gene functions identified directly affect Ty3 transposition remains to be determined. This screen was performed in the haploid background because that is the context in which native Ty3 transposition is initiated during mating. Despite the power of knockout collection screens, a concern, particularly when using the haploid background, is the occurrence of suppressor mutations, which may attenuate or even cause specific phenotypes. Mutants of interest must be reconstructed or complemented with the gene of interest in order to rigorously demonstrate linkage to the Ty3 phenotype. We find it encouraging that GOnet analysis identified clusters of genes encoding proteins with related functions, as would be expected from bona fide effects of specific deletion mutations.
Comparison of Ty screens
Consideration of mutants recovered in the present screen and those in other screens for Ty3, Ty1, and retrovirus host factors suggests that the profile of genes identified is specific to the screening system, that many genes affect the retroelement lifecycle, and that we have not yet even identified all the genes that affect Ty transposition in S. cerevisiae. For example, a low-copy wild-type screen for Ty3 mutants using a random disruption mutant library based on a different S. cerevisiae strain and a position-specific target (Aye et al. 2004) recovered 50 mutants, almost all of which were decreased for transposition and showed only a few genes in common with the screen reported here. There was also no overlap between the two genomewide screens for Ty1 transposition mutants (Scholes et al. 2001; Griffith et al. 2003). However, each of these screens had unique technical features. The screen by Scholes et al. (2001) used a chromosomal Ty1 element under its native promoter, marked with his3AI to screen a haploid disruption library. The screen by Griffith et al. (2003) used a low-copy Ty1 under the GAL1 promoter and marked with HIS3 to screen the knockout collection in a different genetic background. In the former Ty1 screen, only mutants with increased transposition were described. In the latter screen, all but two of the 101 mutants exhibited decreased transposition. Overall, it seems likely that the search for genes affecting Ty3 transposition is not yet saturated.
In spite of the differences among the Ty screens, it is clear that there are a significant number of common host factors for Ty1 and Ty3. We observed effects of 2.9% of nonessential ORF deletions tested. These were evenly divided between mutants with increased and decreased transposition. That is likely to have played a role in our recovery of subsets of genes identified in each of the Ty1 screens. Out of the 29 disruption mutants recovered in the Scholes et al. (2001) screen, there were six genes in common with the current Ty3 screen (VAC8, RTT103, SGS1, RTT109, RRM3, and ELG1/RTT110). Of the 101 mutants recovered in the Ty1 screen by Griffith et al. (2003), 13 (DBP3, NUP84, DBR1, VPH1, NUP133, POP2, DEP1, SEC22, VPS9, ATG17, RPL6A, TPS2, and SIN4) were recovered in our screen. Overall of the 130 genes identified in the two Ty1 screens, we identified a total of 19 (15%) (Table 1). An additional seven genes were in common between our high-copy Ty3 screen and one of the two Ty1 screens, but did not affect transposition of the low-copy Ty3 (Supplemental Table 2).
Conclusions
Clusters of genes encoding RNA processing proteins, vesicular processing complexes, nuclear pore components, and DNA maintenance factors were identified in our search for mutations that affect Ty3 retrotransposition. Analysis of Ty3 Gag3, CA, and IN proteins suggested that under the conditions of our assay, Ty3 proteins were not limiting for transposition. Deletion of one gene encoding a protein required for Golgi to endosome trafficking increased Ty3 protein levels and decreased transposition. Although deletion of one nucleoporin gene appeared to act late and block transposition, disruption of the Nup84 subcomplex required for nuclear pore biogenesis increased Ty3 cDNA and transposition. These results suggest that nuclear entry acts to regulate Ty3 cDNA levels, either directly through facilitating the replication process or indirectly by providing access to the nuclear environment. Deletion of the gene encoding one homolog of the Rfc1 clamp loader involved in repair, Rad24, decreased transposition, but deletion of the gene for another, Elg1, and the genes for a number of interacting proteins increased transposition and cDNA. We hypothesize that cellular replication antagonizes completion of Ty3 reverse transcription. Many of the genes we identified have mammalian homologs, and a significant subset have been shown to affect transposition of the distantly related Ty1 retroelement or retroviruses. Such genes provide a novel source of antiviral target candidates.
Methods
Strains and culture conditions
Media and standard techniques for S. cerevisiae were as previously described (Burke et al. 2000). The collection of nonessential deletion mutants in the BY4741 background (MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0) was purchased from Research Genetics (Invitrogen Inc.). Colonies were suspended in liquid YPD and transformed by using a high-throughput modification of the Lazybones technique (Burke et al. 2000). Transformed cells were grown under noninducing conditions on SD -Ura -His to select for the plasmid and the tagged Ty3 element, respectively, or under inducing conditions on SG -Ura -His.
Plasmid constructions
Construction of a high-copy, URA3-marked plasmid, pNB2142, which carries a GAL1-10UAS–regulated, HIS3-marked Ty3 element (Ty3-HIS3) derived from a HIS3AI-marked element (Sadeghi et al. 2001) has been described (Aye et al. 2004). In order to make a low-copy, Ty3-HIS3 expression plasmid, the GAL1-10UAS and Ty3 in pNB2142 was amplified by PCR and cloned between the HindIII and Not1 sites of the low-copy, URA3 plasmid pRS316. This plasmid was designated as pNB2155.
Transposition assays
A total of 4847 knockout mutants were transformed with pNB2142, and two independent transformants for each strain were patched onto SD -Ura -His medium. Because of poor growth, 390 strains could not be assayed (Supplemental Table 1). After 1 d at room temperature, cells were replica plated to SG -Ura -His to induce transposition and, after 2 d, to YPD to allow plasmid loss. After 3 d on YPD, cells were replica plated to SD -His supplemented with 5FOA (2 mg/mL) in order to select for His+ cells that had acquired a genomic Ty3-HIS3 and lost the URA3-marked plasmid, and after 1 d, to SD -His. Ty3 transposition is proportional to the frequency of His+ papillations. Papillation density was evaluated by two independent observers. The independent transformants were repatched and retested. Mutants that displayed consistent changes from wild type in the high-copy Ty3-HIS3 assay (252) were transformed with the low-copy Ty3-HIS3 plasmid pNB2155, and two independent transformants of each deletion strain were retested as described above.
Immunoblot analysis
BY4741 cells and deletion mutants containing pDLC201 (pEGTy3-1) were grown under non-inducing (SR-ura) conditions to early log phase and then were grown under inducing conditions (SG-ura) for 24 h at 26°C. Whole-cell extracts (WCEs) were prepared from 10 mL cultures by vortexing with glass beads in denaturing buffer (9 M urea, 5 mM EDTA as previously described; Continental Labs). WCEs were fractionated by electrophoresis in a SDS–10% polyacrylamide gel. Proteins were transferred to Immobilon-P (Millipore) and were incubated with rabbit polyclonal anti-Ty3 CA or anti-Ty3 IN antibodies diluted 1:20,000 and 1: 2000, respectively (Menees and Sandmeyer 1994). Rabbit IgG was visualized by using ECL+Plus (Amersham) according to manufacturer's instructions. The blot used to probe for IN was also used to probe for phosphoglycerate kinase (PGK) using anti-PGK IgG (Molecular Probes).
Southern blot analysis
BY4741 cells were grown as described above for immunoblot analysis. Cells were suspended in extraction buffer (10 mM Tris-HCl at pH 8.0, 1 mM EDTA, 100 mM NaCl, 2% Triton X-100, 1% SDS) and vortexed vigorously with glass beads in the presence of phenol:chloroform. Nucleic acid was precipitated from the aqueous phase with ethanol, redissolved, and digested with RNase (Hoffman and Winston 1987). RNA-free DNA (0.5 μg) was digested with EcoRI, fractionated by electrophoresis in a 0.8% agarose gel, transferred to Duralon UV membrane (Stratagene), and immobilized by UV cross-linking in Stratalinker 1800 (Stratagene). Hybridization was performed with a 32P-labeled, internal BglII fragment of Ty3, which hybridizes with the full-length 5.4-kbp Ty3 cDNA, as well as Ty3 donor plasmid and chromosomal Ty3 elements. The filters were washed and exposed to a phosphorimager screen, and hybridization signals were quantitated by using Quantity One software (BioRad). The ratio of cDNA to plasmid signal was used to evaluate the level of cDNA.
GOnet
In order to expedite functional analysis of the genes identified in our screen, a database with a Web interface was developed that integrates genomic, proteomic, and interaction information across several existing sources, including the SGD (http://www.yeastgenome.org/; ftp://ftp.yeastgenome.org/yeast/, March 2004) (Issel-Tarver et al. 2002), the Gene Ontology Consortium (http://www.geneontology.org/), and the Yeast GRID (General Repository for Interaction Datasets) available through http://biodata.mshri.on.ca/yeast_grid (Breitkreutz et al. 2003). A suite of Perl scripts (http://www.perl.org/) was used to parse the ORF information files, extract useful information, and link each ORF with the corresponding GO information. Interaction information downloaded from GRID was symmetrized to mirror the Web version of GRID and to allow greater flexibility in identification of potential interactions. The Web interface to the database was developed by using JavaScript and PHP (http://www.php.net/), a scripting language suited for Web development with a built-in interface for MySQL. GOnet allows users to rapidly cluster genes according to interaction and GO hierarchy information and, in so doing, to identify functions, processes, or components of greatest interest and significance. Output genes can be batch searched by using BLASTP and the NCBI BLAST server (http://www.ncbi.nlm.nih.gov/BLAST/) to identify homologs in other species, or saved and formatted in a spreadsheet or as interaction files for visualization by using Cytoscape (http://www.cytoscape.org/) (Shannon et al. 2003). A detailed description of GOnet and instructions for use are available in the Supplemental data and at the UCI Institute for Genomics and Bioinformatics Web site (http://contact5.ics.uci.edu/gonet/sgdindex.html).
Notes
[6] [Supplemental material is available online at www.genome.org.]
[7] Article and publication are at http://www.genome.org/cgi/doi/10.1101/gr.3739005. Article published online ahead of print in April 2005. Freely available online through the Genome Research Immediate Open Access option.
[8] These authors contributed equally to this work.
[9] Corresponding author. E-mail [email protected]; fax (949) 824-2688.
We thank Roy Parker, Wes Sundquist, Scott Emr, Greg Odorizzi, Joan Curcio, Wes Hatfield, and David Goldfarb for helpful discussions. We thank Tom Menees for helpful discussions and for plasmid constructs. This work was supported by PHS grant GM33281 and UC AIDS Research Program Grant to S.B.S, by NSF MRI grant EIA-0321390 to P.B., and by NLM 5 T15 LM007443, Biomedical Informatics Training Program and UC Systemwide Biotechnology Research and Education Program Grants to P.B. and the Institute for Genomics and Bioinformatics.
References
- ↵Aitchison, J.D., Blobel, G., and Rout, M.P. 1995. Nup120p: A yeast nucleoporin required for NPC distribution and mRNA transport. J. Cell. Biol. 131: 1659–1675.
- ↵Allen, N.P.C., Huang, L., Burlingame, A., and Rexach, M. 2001. Proteomic analysis of nucleoporin interacting proteins. J. Biol. Chem. 276: 29268–29274.
- ↵Allen, N.P.C., Patel, S.S., Huang, L., Chalkley, R.F., Burlingame, A., Lutzmann, M., Hurt, E.C., and Rexach, M. 2002. Deciphering networks of protein interactions at the nuclear pore complex. Mol. Cell. Prot. 1: 930–946.
- ↵Altschul, S.F., Madden, T.L., Schaffer, A.A., Zhang, J., Zhang, Z., Miller, W., and Lipman, D.J. 1997. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 25: 3389–3402.
- ↵Aye, M. and Sandmeyer, S.B. 2003. Ty3 requires yeast La homologous protein for wild-type frequencies of transposition. Mol. Microbiol. 49: 501–515.
- ↵Aye, M., Dildine, S.L., Claypool, J.A., Jourdain, S., and Sandmeyer, S.B. 2001. A truncation mutant of the 95-kilodalton subunit of transcription factor IIIC reveals asymmetry in Ty3 integration. Mol. Cell Biol. 21: 7839–7851.
- ↵Aye, M., Irwin, B., Beliakova-Bethell, N., Chen, E., Garrus, J., and Sandmeyer, S.B. 2004. Host factors that affect Ty3 retrotransposition in Saccharomyces cerevisiae. Genetics 168: 1159–1176.
- ↵Babst, M., Katzmann, D.J., Estepa-Sabal, E.J., Meerloo, T., and Emr, S.D. 2002a. ESCRT-III: An endosome-associated heterooligomeric protein complex required for MVB sorting. Dev. Cell 3: 271–282.
- ↵Babst, M., Katzmann, D.J., Snyder, W.B., Wendland, B., and Emr, S.D. 2002b. Endosome-associated complex, ESCRT-II, recruits transport machinery for protein sorting at the multivesicular body. Dev. Cell 3: 283–289.
- ↵Basyuk, E., Galli, T., Mougel, M., Blanchard, J.M., Sitbon, M., and Bertrand, E. 2003. Retroviral genomic RNAs are transported to the plasma membrane by endosomal vesicles. Dev. Cell 5: 161–174.
- ↵Bellaoui, M., Chang, M., Ou, J.W., Xu, H., Boone, C., and Brown, G.W. 2003. Elg1 forms an alternative RFC complex important for DNA replication and genome integrity. EMBO J. 22: 4304–4313.
- ↵Ben-Aroya, S., Koren, A., Liefshitz, B., Steinlauf, R., and Kupiec, M. 2003. ELG1, a yeast gene required for genome stability, forms a complex related to replication factor C. Proc. Natl. Acad. Sci. 100: 9906–9911.
- ↵Bertrand, E., Houser-Scott, F., Kendall, A., Singer, R.H., and Engelke, D.R. 1998. Nucleolar localization of early tRNA processing. Genes & Dev. 12: 2463–2468.
- ↵Bolton, E.C., Mildvan, A.S., and Boeke, J.D. 2002. Inhibition of reverse transcription in vivo by elevated manganese ion concentration. Mol. Cell 9: 879–889.
- ↵Bonifacino, J.S. and Glick, B.S. 2004. The mechanisms of vesicle budding and fusion. Cell 116: 153–166.
- ↵Bowers, K., Lottridge, J., Helliwell, S.B., Goldthwaite, L.M., Luzio, J.P., and Stevens, T.H. 2004. Protein–protein interactions of ESCRT complexes in the yeast Saccharomyces cerevisiae. Traffic 5: 194–210.
- ↵Breitkreutz, B.J., Stark, C., and Tyers, M. 2003. The GRID: The General Repository for Interaction Datasets. Genome Biol. 4: R23.
- ↵Bryk, M., Banerjee, M., Conte, D., and Curcio, M.J. 2001. The Sgs1 helicase of Saccharomyces cerevisiae inhibits retrotransposition of Ty1 multimeric arrays. Mol. Cell. Biol. 21: 5374–5388.
- ↵Burd, G.C., Peterson, M., Cowles, C.R., and Emr, S.D. 1997. A novel Sec18p/NSF-dependent complex required for Golgi-to-endosome transport in yeast. Mol. Biol. Cell 8: 1089–1104.
- ↵Burke, D., Dawson, D., and Stearns, T. 2000. Methods in yeast genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
- ↵Burri, L. and Lithgow, T. 2004. A complete set of SNAREs in yeast. Traffic 5: 45–52.
- ↵Chapman, K.B. and Boeke, J.D. 1991. Isolation and characterization of the gene encoding yeast debranching enzyme. Cell 65: 483–492.
- ↵Chavez, S., Beilharz, T., Rondon, A.G., Erdjument-Bromage, H., Tempst, P., Svejstrup, J.Q., Lithgow, T., and Aguilera, A. 2000. A protein complex containing Tho2, Hpr1, Mft1 and a novel protein, Thp2, connects transcription elongation with mitotic recombination in Saccharomyces cerevisiae. EMBO J. 19: 5824–5834.
- ↵Cowan, N.J. and Lewis, S.A. 2002. Type II chaperonins, prefoldin, and the tubulin-specific chaperones. Protein Folding Cell 59: 73–104.
- ↵Damelin, M. and Silver, P.A. 2002. In situ analysis of spatial relationships between proteins of the nuclear pore complex. Biophys. J. 83: 3626–3636.
- ↵Dang, V.D. and Levin, H.L. 2000. Nuclear import of the retrotransposon Tf1 is governed by a nuclear localization signal that possesses a unique requirement for the FXFG nuclear pore factor nup124p. Mol. Cell. Biol. 20: 7798–7812.
- ↵Daniel, R., Katz, R.A., and Skalka, A.M. 1999. A role for DNA-PK in retroviral DNA integration. Science 284: 644–647.
- ↵Daniel, R., Kao, G., Taganov, K., Greger, J.G., Favorova, O., Merkel, G., Yen, T.J., Katz, R.A., and Skalka, A.M. 2003. Evidence that the retroviral DNA integration process triggers an ATR-dependent DNA damage response. Proc. Natl. Acad. Sci. 100: 4778–4783.
- ↵Decker, C.J. and Parker, R. 2002. mRNA decay enzymes: Decappers conserved between yeast and mammals. Proc. Natl. Acad. Sci. 99: 12512–12514.
- ↵Doye, V., Wepf, R., and Hurt, E.C. 1994. A novel nuclear-pore protein Nup133p with distinct roles in Poly(A)(+) RNA transport and nuclear-pore distribution. EMBO J. 13: 6062–6075.
- ↵Duina, A.A., Marsh, J.A., and Gaber, R.F. 1996. Identification of two CyP-40–like cyclophilins in Saccharomyces cerevisiae, one of which is required for normal growth. Yeast 12: 943–952.
- ↵Dvorin, J.D. and Malim, M.H. 2003. Intracellular trafficking of HIV-1 cores: Journey to the center of the cell. Curr. Top. Microbiol. Immunol. 281: 179–208.
- ↵Dworetzky, S.I., Lanford, R.E., and Feldherr, C.M. 1988. The effects of variations in the number and sequence of targeting signals on nuclear uptake. J. Cell Biol. 107: 1279–1287.
- ↵Fabre, F., Chan, A., Heyer, W.D., and Gangloff, S. 2003. Alternate pathways involving Sgs1/Top3, Mus81/Mms4, and Srs2 prevent formation of toxic recombination intermediates from single-stranded gaps created by DNA replication. Proc. Natl. Acad. Sci. 99: 11887–16892.
- ↵Fehlker, M., Wendler, P., Lehmann, A., and Enenkel, C. 2003. Blm3 is part of nascent proteasomes and is involved in a late stage of nuclear proteasome assembly. E. Reports 4: 959–963.
- ↵Fischer, T., Strasser, K., Racz, A., Rodriguez-Navarro, S., Oppizzi, M., Ihrig, P., Lechner, J., and Hurt, E. 2002. The mRNA export machinery requires the novel Sac3p-Thp1p complex to dock at the nucleoplasmic entrance of the nuclear pores. EMBO J. 21: 5843–5852.
- ↵Freed, E.O. 2004. HIV-1 and the host cell: An intimate association. Trends Microbiol. 12: 170–177.
- ↵Gallardo, M., Luna, R., Erdjument-Bromage, H., Tempst, P., and Aguilera, A. 2003. Nab2p and the Thp1p-Sac3p complex functionally interact at the interface between transcription and mRNA metabolism. J. Biol. Chem. 278: 24225–24232.
- ↵Garfinkel, D. and Bailis, A. 2002. Nucleotide excision repair, genome stability and human disease: New insight from model systems. J. Biomed. Biotechnol. 2: 55–60.
- ↵Gavin, A.C., Bosche, M., Krause, R., Grandi, P., Marzioch, M., Bauer, A., Schultz, J., Rick, J.M., Michon, A.M., Cruciat, C.M., et al. 2002. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415: 141–147.
- ↵Giaever, G., Chu, A.M., Ni, L., Connelly, C., Riles, L., Veronneau, S., Dow, S., Lucau-Danila, A., Anderson, K., Andre, B., et al. 2002. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418: 387–391.
- ↵Giannattasio, M., Lazzaro, F., Plevani, P., and Muzi-Falconi, M. 2005. The DNA damage checkpoint response requires histone H2B ubiquitination by Rad6-Bre1 and H3 methylation by Dot1. J. Biol. Chem. 280 9879–9886.
- ↵Goff, S.P. 2001. Intracellular trafficking of retroviral genomes during the early phase of infection: viral exploitation of cellular pathways. J. Gene Med. 3: 517–528.
- ↵Gottwein, E., Bodem, J., Muller, B., Schmechel, A., Zentgraf, H., and Krausslich, H.G. 2003. The Mason-Pfizer monkey virus PPPY and PSAP motifs both contribute to virus release. J. Virol. 77: 9474–9485.
- ↵Greene, W.C. and Peterlin, B.M. 2002. Charting HIV's remarkable voyage through the cell: Basic science as a passport to future therapy. Nat. Med. 8: 673–680.
- ↵Griffith, J.L., Coleman, L.E., Raymond, A.S., Goodson, S.G., Pittard, W.S., Tsui, C., and Devine, S.E. 2003. Functional genomics reveals relationships between the retrovirus-like Ty1 element and its host Saccharomyces cerevisiae. Genetics 164: 867–879.
- ↵Hama, H., Tall, G.G., and Horazdovsky, B.F. 1999. Vps9p is a guanine nucleotide exchange factor involved in vesicle-mediated vacuolar protein transport. J. Biol. Chem. 274: 15284–15291.
- ↵Hansen, L.J., Chalker, D.L., Orlinsky, K.J., and Sandmeyer, S.B. 1992. Ty3 GAG3 and POL3 genes encode the components of intracellular particles. J. Virol. 66: 1414–1424.
- ↵Hoege, C., Pfander, B., Moldovan, G.L., Pyrowolakis, G., and Jentsch, S. 2002. RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419: 135–141.
- ↵Hoffman, C.S. and Winston, F. 1987. A 10-minute DNA preparation from yeast efficiently releases autonomous plasmids for transformation of Escherichia coli. Gene 57: 267–272.
- ↵Hong, S., Choi, G., Park, S., Chung, A.S., Hunter, E., and Rhee, S.S. 2001. Type-D retrovirus Gag polyprotein interacts with the cytosolic chaperonin TRiC. J. Virol. 75: 2526–2534.
- ↵Horak, J. 2003. The role of ubiquitin in down-regulation and intracellular sorting of membrane proteins: insights from yeast. Biochim. Biophys. Acta.-Biomembranes 1614: 139–155.
- ↵Hughes, T.R., Evans, S.K., Weilbaecher, R.G., and Lundblad, V. 2000. The Est3 protein is a subunit of yeast telomerase. Curr. Biol. 10: 809–812.
- ↵Hwang, W.W., Venkatasubrahmanyam, S., Ianculescu, A.G., Tong, A., Boone, C., and Madhani, H.D. 2003. A conserved RING finger protein required for histone H2B monoubiquitination and cell size control. Mol. Cell 11: 261–266.
- ↵Ideker, T., Thorsson, V., Ranish, J.A., Christmas, R., Buhler, J., Eng, J.K., Bumgarner, R., Goodlett, D.R., Aebersold, R., and Hood, L. 2001. Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 292: 929–934.
- ↵Issel-Tarver, L., Christie, K.R., Dolinski, K., Andrada, R., Balakrishnan, R., Ball, C.A., Binkley, G., Dong, S., Dwight, S.S., Fisk, F.G., et al. 2002. Saccharomyces genome database. Methods Enzymol. 350: 329–346.
- ↵Jimeno, S., Rondon, A.G., Luna, R., and Aguilera, A. 2002. The yeast THO complex and mRNA export factors link RNA metabolism with transcription and genome instability. EMBO J. 21: 3526–3535.
- ↵Kanellis, P., Agyei, R., and Durocher, D. 2003. Elg1 forms an alternative PCNA-interacting RFC complex required to maintain genome stability. Curr. Biol. 13: 1583–1595.
- ↵Karst, S.M., Rutz, M., and Menees, T. 2000. The yeast retrotransposons Ty1 and Ty3 require the RNA lariat debranching enzyme, Dbr1p, for efficient accumulation of reverse transcripts. Biochem. Biophys. Res. Commun. 268: 112–117.
- ↵Katzmann, D.J., Odorizzi, G., and Emr, S.D. 2002. Receptor downregulation and multivesicular-body sorting. Nat. Rev. Mol. Cell Biol. 3: 893–905.
- ↵Keeney, J.B., Chapman, K.B., Lauermann, V., Voytas, D.F., Astrom, S.U., von Pawel-Rammingen, U., Bystrom, A., and Boeke, J.D. 1995. Multiple molecular determinants for retrotransposition in a primer tRNA. Mol. Cell. Biol. 15: 217–226.
- ↵Kikonyogo, A., Bouamr, F., Vana, M.L., Xiang, Y., Aiyar, A., Carter, C., and Leis, J. 2001. Proteins related to the Nedd4 family of ubiquitin protein ligases interact with the L domain of Rous sarcoma virus and are required for gag budding from cells. Proc. Natl. Acad. Sci. 98: 11199–11204.
- ↵Kim, M., Krogan, N.J., Vasiljeva, L., Rando, O.J., Nedea, E., Greenblatt, J.F., and Buratowski, S. 2004. The yeast Rat1 exonuclease promotes transcription termination by RNA polymerase II. Nature 432: 517–522.
- ↵Kirchner, J., Connolly, C.M., and Sandmeyer, S.B. 1995. Requirement of RNA polymerase III transcription factors for in vitro position-specific integration of a retroviruslike element. Science 267: 1488–1491.
- ↵Kushner, D.B., Lindenbach, B.D., Grdzelishvili, V.Z., Noueiry, A.O., Paul, S.M., and Ahlquist, P. 2003. Systematic, genome-wide identification of host genes affecting replication of a positive-strand RNA virus. Proc. Natl. Acad. Sci. 100: 15764–15769.
- ↵Laplaza, J.M., Bostick, M., Scholes, D.T., Curcio, M.J., and Callis, J. 2004. Saccharomyces cerevisiae ubiquitin-like protein Rub1 conjugates to cullin proteins Rtt101 and Cul3 in vivo. Biochem. J. 377: 459–467.
- ↵Lee, B.S., Bi, L., Garfinkel, D.J., and Bailis, A.M. 2000. Nucleotide excision repair/TFIIH helicases RAD3 and SSL2 inhibit short-sequence recombination and Ty1 retrotransposition by similar mechanisms. Mol. Cell Biol. 20: 2436–2445.
- ↵Lei, E.P., Krebber, H., and Silver, P.A. 2001. Messenger RNAs are recruited for nuclear export during transcription. Genes & Dev. 15: 1771–1782.
- ↵Levin, H.L. 2002. Newly identified retrotransposons of the Ty3/gypsy class in fungi, plants, and vertebrates. In Mobile DNA II (eds. N.L. Craig et al.), pp. 684–701. ASM, Washington, DC.
- ↵Li, L., Olvera, J.M., Yoder, K.E., Mitchell, R.S., Butler, S.L., Lieber, M., Martin, S.L., and Bushman, F.D. 2001. Role of the non-homologous DNA end joining pathway in the early steps of retroviral infection. EMBO J. 20: 3272–3281.
- ↵Lin, S.S., Nymark-McMahon, M.H., Yieh, L., and Sandmeyer, S.B. 2001. Integrase mediates nuclear localization of Ty3. Mol. Cell. Biol. 21: 7826–7838.
- ↵Macara, I.G. 2001. Transport into and out of the nucleus. Microbiol. Mol. Biol. Rev. 65: 570–594.
- ↵Maga, G. and Hubscher, U. 2003. Proliferating cell nuclear antigen (PCNA): A dancer with many partners. J. Cell Sci. 116: 3051–3060.
- ↵Majka, J. and Burgers, P.M.J. 2004. The PCNA-RFC families of DNA clamps and clamp loaders. Prog. Nucleic Acid Res. Mol. Biol. 78: 227–260.
- ↵Mayer, M.L., Pot, I., Chang, M., Xu, H., Aneliunas, V., Kwok, T., Newitt, R., Abersold, R., Boone, C., Brown, G.W., et al. 2004. Identification of protein complexes required for efficient sister chromatid cohesion. Mol. Biol. Cell 15: 1736–1745.
- ↵Menees, T.M. and Sandmeyer, S.B. 1994. Transposition of the yeast retroviruslike element Ty3 is dependent on the cell cycle. Mol. Cell. Biol. 14: 8229–8240.
- ↵Miller, M.D., Farnet, C.M., and Bushman, F.D. 1997. Human immunodeficiency virus type 1 preintegration complexes: Studies of organization and composition. J. Virol. 71: 5382–5390.
- ↵Odorizzi, G., Babst, M., and Emr, S.D. 1998. Fab1p PtdIns(3)P 5-kinase function essential for protein sorting in the multivesicular body. Cell 95: 847–858.
- ↵Odorizzi, G., Katzmann, D.J., Babst, M., Audhya, A., and Emr, S.D. 2003. Bro1 is an endosome-associated protein that functions in the MVB pathway in Saccharomyces cerevisiae. J. Cell Sci. 116: 1893–1903.
- ↵Ohtake, Y. and Wickner, R.B. 1995. Yeast virus propagation depends critically on free 60S ribosomal subunit concentration. Mol. Cell Biol. 15: 2772–2781.
- ↵Orphanides, G. and Reinberg, D. 2002. A unified theory of gene expression. Cell 108: 439–451.
- ↵Osborn, A.J. and Elledge, S.J. 2003. Mrc1 is a replication fork component whose phosphorylation in response to DNA replication stress activates Rad53. Genes & Dev. 17: 1755–1767.
- ↵Ott, D.E. 2002. Potential roles of cellular proteins in HIV-1. Rev. Med. Virol. 12: 359–374.
- ↵Pante, N. and Kann, M. 2002. Nuclear pore complex is able to transport macromolecules with diameters of about 39 nm. Mol. Biol. Cell 13: 425–434.
- ↵Pornillos, O., Garrus, J.E., and Sundquist, W.I. 2002. Mechanisms of enveloped RNA virus budding. Trends Cell Biol. 12: 569–579.
- ↵Preston, R.A., Reinagel, P.S., and Jones, E.W. 1992. Genes required for vacuolar acidity in Saccharomyces cerevisiae. Genetics 131: 551–558.
- ↵Pruyne, D. and Bretscher, A. 2000. Polarization of cell growth in yeast I. Establishment and maintenance of polarity states. J. Cell Sci. 113: 365–375.
- ↵Rattray, A.J., Shafer, B.K., and Garfinkel, D.J. 2000. The Saccharomyces cerevisiae DNA recombination and repair functions of the RAD52 epistasis group inhibit Ty1 transposition. Genetics 154: 543–556.
- ↵Rout, M.P., Aitchison, J.D., Suprapto, A., Hjertaas, K., Zhao, Y.M., and Chait, B.T. 2000. The yeast nuclear pore complex: Composition, architecture, and transport mechanism. J. Cell Biol. 148: 635–651.
- ↵Sadeghi, N., Rutz, M.L., and Menees, T.M. 2001. Thermal blockage of viruslike particle formation for the yeast retrotransposon Ty3 reveals differences in the cellular stress response. Arch. Virol. 146: 1919–1934.
- ↵Sandmeyer, S.B., Aye, M., and Menees, T.M. 2002. Ty3: A position-specific, gypsylike element in Saccharomyces cerevisiae. In Mobile DNA II (eds. N.L. Craig, et al.), pp. 663–682. ASM Press, Washington, DC.
- ↵Schmidt, K.H., Derry, K.L., and Kolodner, R.D. 2002. Saccharomyces cerevisiae Rrm3, a 5′ to 3′ DNA helicase, physically interacts with proliferating cell nuclear antigen. J. Biol. Chem. 277: 45331–45337.
- ↵Schmidt, K.H. and Kolodner, R.D. 2004. Requirement of Rrm3 helicase for repair of spontaneous DNA lesions in cells lacking Srs2 or Sgs1 helicase. Mol. Cell. Biol. 24: 3213–3226.
- ↵Scholes, D.T., Banerjee, M., Bowen, B., and Curcio, M.J. 2001. Multiple regulators of Ty1 transposition in Saccharomyces cerevisiae have conserved roles in genome maintenance. Genetics 159: 1449–1465.
- ↵Scholes, D.T., Kenny, A.E., Gamache, E.R., Mou, Z., and Curcio, M.J. 2003. Activation of a LTR-retrotransposon by telomere erosion. Proc. Natl. Acad. Sci. 100: 15736–15741.
- ↵Sfakianos, J.N. and Hunter, E. 2003. M-PMV capsid transport is mediated by Env/Gag interactions at the pericentriolar recycling endosome. Traffic 4: 671–680.
- ↵Shannon, P., Markiel, A., Ozier, O., Baliga, N.S., Wang, J.T., Ramage, D., Amin, N., Schwikowski, B., and Ideker, T. 2003. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13: 2498–2504.
- ↵Sheth, U. and Parker, R. 2003. Decapping and decay of messenger RNA occur in cytoplasmic processing bodies. Science 300: 805–808.
- ↵Siniossoglou, S., Lutzmann, M., Santos-Rosa, H., Leonard, L., Mueller, S., Aebi, U., and Hurt, E. 2000. Structure and assembly of the Nup84p complex. J. Cell Biol. 149: 41–53.
- ↵Stage-Zimmermann, T., Schmidt, U., and Silver, P.A. 2000. Factors affecting nuclear export of the 60S ribosomal subunit in vivo. Mol. Biol. Cell 11: 3777–3789.
- ↵Sterner, D.E. and Berger, S.L. 2000. Acetylation of histones and transcription-related factors. Microbiol. Mol. Biol. Rev. 64: 435–459.
- ↵Strack, B., Calistri, A., Craig, S., Popova, E., and Gottlinger, H.G. 2003. AIP1/ALIX is a binding partner for HIV-1 p6 and EIAV p9 functioning in virus budding. Cell 114: 689–699.
- ↵Strasser, K., Masuda, S., Mason, P., Pfannstiel, J., Oppizzi, M., Rodriguez-Navarro, S., Rondon, A.G., Andres, A.K., Struhl, K., Reed, R., et al. 2002. TREX is a conserved complex coupling transcription with messenger RNA export. Nature 417: 304–308.
- ↵Sundararajan, A., Lee, B.S., and Garfinkel, D.J. 2003. The Rad27 (Fen-1) nuclease inhibits Ty1 mobility in Saccharomyces cerevisiae. Genetics 163: 55–67.
- ↵Sundin, B.A., Chiu, C.H., Riffle, M., Davis, T.N., and Muller, E.G.D. 2004. Localization of proteins that are coordinately expressed with Cln2 during the cell cycle. Yeast 21: 793–800.
- ↵Tall, G.G., Hama, H., Dewald, D.B., and Horazdovsky, B.F. 1999. The phosphatidylinositol 3-phosphate binding protein Vac1p interacts with a Rab GTPase and a Sec1p homologue to facilitate vesicle-mediated vacuolar protein sorting. Mol. Biol. Cell 10: 1873–1889.
- ↵Thompson, M., Haeusler, R.A., Good, P.D., and Engelke, D.R. 2003. Nucleolar clustering of dispersed tRNA genes. Science 302: 1399–1401.
- ↵Towers, G.J., Hatziioannou, T., Cowan, S., Goff, S.P., Luban, J., and Bieniasz, P.D. 2003. Cyclophilin A modulates the sensitivity of HIV-1 to host restriction factors. Nat. Med. 9: 1138–1143.
- ↵von Schwedler, U.K., Stuchell, M., Muller, B., Ward, D.M., Chung, H.Y., Morita, E., Wang, H.E., Davis, T., He, G.P., Cimbora, D.M., et al. 2003. The protein network of HIV budding. Cell 114: 701–713.
- ↵Voytas, D.F. and Boeke, J.D. 2002. Ty1 and Ty5 of Saccharomyces cerevisiae. In Mobile DNA II (eds. N.L. Craig et al.), pp. 631–683. ASM, Washington, DC.
- ↵Webb, G.C., Zhang, J.Q., Garlow, S.J., Wesp, A., Riezman, H., and Jones, E.W. 1997. Pep7p provides a novel protein that functions in vesicle-mediated transport between the yeast Golgi and endosome. Mol. Biol. Cell 8: 871–895.
- ↵Weisman, L.S. and Wickner, W. 1992. Molecular characterization of Vac1, a gene required for vacuole inheritance and vacuole protein sorting. J. Biol. Chem. 267: 618–623.
- ↵Winzeler, E.A., Shoemaker, D.D., Astromoff, A., Liang, H., Anderson, K., Andre, B., Bangham, R., Benito, R., Boeke, J.D., Bussey, H., et al. 1999. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285: 901–906.
- ↵Xie, W., Gai, X., Zhu, Y., Zappulla, D.C., Sternglanz, R., and Voytas, D.F. 2001. Targeting of the yeast Ty5 retrotransposon to silent chromatin is mediated by interactions between integrase and Sir4p. Mol. Cell Biol. 21: 6606–6614.
- ↵Xu, W.J., Smith, F.J., Subaran, R., and Mitchell, A.P. 2004. Multivesicular body-ESCRT components function in pH response regulation in Saccharomyces cerevisiae and Candida albicans. Mol. Biol. Cell 15: 5528–5537.
- ↵Yoder, K.E. and Bushman, F.D. 2000. Repair of gaps in retroviral DNA integration intermediates. J. Virol. 74: 11191–11200.
Web site references
- http://www.cytoscape.org/; Cytoscape.
- http://www.geneontology.org; Gene Ontology Consortium.
- http://biodata.mshri.on.ca/yeast_grid/; Yeast GRID.
- http://www.ncbi.nlm.nih.gov/BLAST/; BLAST server.
- http://contact5.ics.uci.edu/gonet/sgdindex.html; UCI Institute for Genomics and Bioinformatics Web site.

