The Evolution of Trichromatic Color Vision by Opsin Gene Duplication in New World and Old World Primates
Abstract
Trichromacy in all Old World primates is dependent on separate X-linked MW and LW opsin genes that are organized into a head-to-tail tandem array flanked on the upstream side by a locus control region (LCR). The 5′ regions of these two genes show homology for only the first 236 bp, although within this region, the differences are conserved in humans, chimpanzees, and two species of cercopithecoid monkeys. In contrast, most New World primates have only a single polymorphic X-linked opsin gene; all males are dichromats and trichromacy is achieved only in those females that possess a different form of this gene on each X chromosome. By sequencing the upstream region of this gene in a New World monkey, the marmoset, we have been able to demonstrate the presence of an LCR in an equivalent position to that in Old World primates. Moreover, the marmoset sequence shows extensive homology from the coding region to the LCR with the upstream sequence of the human LW gene, a distance of >3 kb, whereas homology with the human MW gene is again limited to the first 236 bp, indicating that the divergent MW sequence identifies the site of insertion of the duplicated gene. This is further supported by the presence of an incomplete Alu element on the upstream side of this insertion point in the MW gene of both humans and a cercopithecoid monkey, with additional Alu elements present further upstream. Therefore, these Aluelements may have been involved in the initial gene duplication and may also be responsible for the high frequency of gene loss and gene duplication within the opsin gene array. Full trichromacy is present in one species of New World monkey, the howler monkey, in which separate MW and LW genes are again present. In contrast to the separate genes in humans, however, the upstream sequences of the two howler genes show homology with the marmoset for at least 600 bp, which is well beyond the point of divergence of the human MW and LW genes, and each sequence is associated with a different LCR, indicating that the duplication in the howler monkey involved the entire upstream region.
[The sequence data described in this paper have been submitted to GenBank under accession nos. AF155218, AF156715, and AF156716.]
Among the mammals, primates are unique in possessing trichromatic color vision. In humans and Old World primates, this is based on three classes of cone photoreceptor within the retina, each containing a different visual pigment. Visual pigments are composed of a chromophore, retinal, covalently bound by a protonated Schiff base to a protein moiety, opsin. Therefore, spectral differences between primate pigments are entirely attributable to differences in the amino acid sequence of the opsin proteins.
The spectral peaks (λmax) of the three visual pigments in primates are in the violet [short wave (SW)], green [middle wave (MW)], and yellow-green [long wave (LW)] regions of the spectrum. In Old World primates, the SW opsin is encoded by an autosomal gene and the MW and LW opsins by separate genes on the X chromosome (Nathans et al. 1986; Ibbotson et al. 1992). Sequence comparison between these two genes in different Old World species (Ibbotson et al. 1992) indicates that they arose by gene duplication at the base of the Old World lineage ∼40 mya (Goodman et al. 1998; Hunt et al. 1998). The two genes show substantial sequence identity throughout their coding (Nathans et al. 1986) and intronic (Shyue et al. 1995; Zhao et al. 1998) regions. They are organized into a head-to-tail tandem array (Nathans et al. 1986; Vollrath et al. 1988; Feil et al. 1990), which is bound on the upstream side by a so-called locus control region (LCR) (Wang et al. 1992), the presence of which is critical for the expression of either gene (Nathans et al. 1989) and may be involved in the selective expression of one or other gene in particular cone photoreceptors (Shaaban and Deeb 1998).
The situation in New World monkeys is much more variable. The nocturnal monkey Aotus trivirgatus lacks a functional SW gene and is monochromatic (Jacobs et al. 1996a). Most New World species exhibit a trichromacy that is based on only two opsin genes, an autosomal SW gene as in Old World primates, and a polymorphic X-linked MW/LW gene with up to three allelic forms that encode pigments with differing λmax values. Male monkeys combine the SW gene with just one of the different allelic forms of the X-linked gene and can generate only two cone pigments. Therefore, they are dichromats. In contrast, those females that inherit a different form of the MW/LW gene from each parent possess trichromatic vision, as X-inactivation will ensure that only one allele is expressed per cell (Mollon et al. 1984). Therefore, there is no need for an additional mechanism for the selective expression of the different allelic forms of the polymorphic X-linked gene in different cone photoreceptors.
An exception to this polymorphism-based trichromacy of New World primates has been identified by Jacobs et al. (1996b) in the howler monkey Alouatta seniculus and Alouatta caraya, where separate MW and LW genes are present, thus conferring full trichromacy in both males and females. Sequence analysis of the two genes indicates that this is a more recent duplication than in Old World primates (Hunt et al. 1998), which occurred after the separation of the Old World and New World lineages. For a functional trichromacy to be present in this genus, a system for the selective expression of either the MW or the LW gene in particular photoreceptors, is now required.
In this paper we have addressed the question of the origin of the duplication in Old World primates and in the howler monkey. The original report of the sequence of the human MW and LW genes by Nathans et al. (1986) included ∼450 bp of the upstream sequence of both genes. This region shows only 14 single nucleotide substitutions in the first 236 bp, but then diverges completely, indicating that this is the point of insertion of the duplicated opsin gene. To confirm this, we have sequenced the upstream region of the single X-linked gene in the marmoset Callithrix jacchus, a New World monkey with a polymorphism-based trichromacy. The absence of any opsin gene duplication in this species means that the upstream sequence will reflect the ancestral arrangement. We have also examined the conservation of substitution within the region of homology of the MW and LW genes in humans, chimpanzee, and in two species of Old World cercopithecoid monkeys.
The presence of two distinct upstream regions to the X-linked opsin genes in the howler monkey has already been established (Kainz et al. 1998). We have extended this sequence across the region of divergence in the Old World primate genes and also into the LCR. From these sequences, we have been able to establish that the opsin gene duplication in this genus is very different from that in Old World primates.
RESULTS
The 5′-Flanking Regions of the Separate MW and LW Opsin Genes in Old World Primates
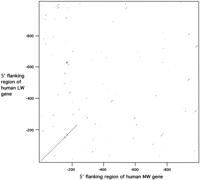
The first gene in the opsin gene array on the X chromosome is the LW gene and there is a distance of ∼3.5 kb between the upstream LCR and the transcription start site for this gene [accession nos. S44029(Wang et al. 1992) and Z68193 (Sanger Centre data bank 1995)]. Within this 5′-flanking sequence, a number of repeat elements are present including an Alu at −1991 to −2232, three L1 elements at −1360 to −1622, −1781 to −1970, and −2261 to −2341, and a MER12 element at −2438 to −2617. In the case of the MW gene, Nathans et al. (1986) originally published 450 bp of the human 5′-flanking region; this was extended by ∼700 bp to −1185 byDulai (1996) and further extended to −2513 by Hanna et al. (1997; accession no. U93721). A dot matrix plot of the first 1000 bp of the 5′-flanking regions of the MW and LW genes (Fig. 1) confirms that homology is restricted to the first 236 bp. Further analysis of the 5′-flanking region of the human MW gene shows that it contains four Alu repeat elements (Table 1), a partial repeat of 120 bp at position −234 immediately adjacent to the point of sequence divergence between the MW and LW 5′ flanking regions, and three more complete elements further upstream. To establish whether these Alu elements are also present in the upstream region of the MW gene of another Old World primate, a fragment of ∼750 bp of the upstream region of the MW gene in the diana monkey was amplified and cloned. Alignment with the human sequence shows that this fragment contains Alu elements at identical positions to the elements present in the human MW gene at positions −234 to −358, and −369 to −652 (Fig. 2).
Dot matrix plot of human MW and LW upstream region. The parameters for comparison were set at 60% identity for a sliding window of 10 nucleotides. The presence of a diagonal line that passes through the origin indicates homology at the same position in both sequences. Note that no continuous lines are present beyond position −236.
Alu Repeat Elements in the 5′-Flanking Region of the Human MW Opsin Gene
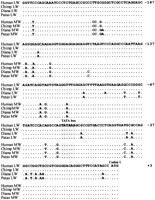
Alignment of the upstream region of the MW gene in humans and diana monkeys. The Alu repeat elements present in both sequences are shaded. Identity between the human and diana sequences is indicated by a dot.
The homologous region of the human MW and LW upstream sequences show a total of 14 substitutions between the start of the coding sequence and the point of sequence divergence at −237. To determine whether these differences are conserved, we have sequenced the region in the chimpanzee Pan troglodytes and in two species of cercopithecoid monkeys, the diana monkey Cercopithecus dianaand the Patas monkey Erythrocebus patas. As shown in Figure3, with the exception of the substitution at the transcription start, all of the differences between the human MW and LW upstream sequences in this region of homology are conserved, although the cercopithecoid MW and LW sequences differ at a few other positions.
Nucleotide sequences of the homologous upstream regions of the MW and LW genes of Old World primates. Identity to the human LW sequence is indicated by a dot.
The 5′-Flanking Region of the Single MW/LW Opsin Gene in a New World Primate
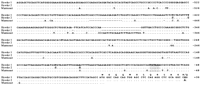
Using a combination of gene walking and conventional PCR, we have sequenced the entire 5′-flanking region of the marmoset MW/LW opsin gene from the translation start site to the conserved core sequence of the LCR. Figure 4 shows the alignment of the marmoset sequence with the upstream region of the human LW gene. Our amplification strategy involved using a primer designed to the 5′ half of the conserved core sequence of the LCR; therefore, the sequence extends only as far as the 3′ half of this region. The available sequence from the marmoset is sufficient to conclude, however, that an LCR is present upstream of the single MW/LW gene in a New World monkey.
Nucleotide sequence of the 5′-flanking regions of the human LW gene and the marmoset MW/LW gene, plus the homologous region of the human MW gene. Identity to the human LW sequence is indicated by a dot. Indels are indicated by dashes in the deleted sequence. The shaded regions identify the LCR core, the deleted region of the L1 repeat element in the marmoset, and the TATA box. Marmoset sequence, GenBank accession no. AF155218.
Sequence identity between the marmoset and human LW sequences is substantial throughout the entire length; the overall identity is 77%, and there is no evidence of duplicated regions within either sequence. Although the marmoset sequence is 291 bp shorter than the human LW sequence, alignment requires a number of insertions/deletions (indels) in both sequences; only one of these indels, a deletion of 42 bp at −1581 in the marmoset sequence, exactly coincides with a repeat element in these sequences, in this case with the 5′ end of a L1 repeat element at −1622 in the human sequence. In contrast, the human MW sequence shows essentially no homology with the marmoset sequence beyond the first 236 bp of upstream sequence. This region of the human MW sequence has also been included in Figure 4. Within this region, there are 14 differences between the human MW and LW genes, with the marmoset sequence showing identity to the human MW sequence at nine of these sites and to the human LW sequence at the other five sites. Within this region, therefore, the marmoset promoter shows a closer homology to the human MW than to the human LW gene.
The absence of any homology between the marmoset and human MW upstream sequences beyond position −236 effectively rules out the possibility that divergence beyond this region between the human MW and LW genes is the result of selective changes within the promoter regions of these two genes. Rather, the divergence must be the consequence of gene duplication and therefore identifies the breakpoint for the gene insertion that gave rise to the separate MW and LW genes in Old World primates.
Duplication of the X-Linked Opsin Genes in the Howler Monkey
A fragment of ∼700 bp was amplified that included 620 bp of the 5′-flanking region of the X-linked opsin genes in the howler monkey. The sequencing of nine clones of this amplified fragment showed that they fell into two classes distinguished by four nucleotide differences in the 5′-flanking region at −111, −125, −422, and −431, and a single difference in the coding region at +38 (Fig.5). This latter substitution will result in an arginine/glutamine amino acid difference between the two howler opsins. The substitution at −111 has been reported previously by Kainz et al. (1998) in a shorter direct-sequenced PCR-amplified fragment from the howler upstream region, but the second substitution at −90 reported by these investigators (these sites are incorrectly numbered in Kainz et al. 1998) was not present in our sequence. Therefore, the presence of two distinct upstream sequences is consistent with the presence of two X-linked opsin genes in this species of New World monkey and we conclude that one is derived from the upstream region of the MW gene and the other from the upstream region of the LW gene. This fragment extends well beyond the 236 bp of homology of the 5′ flanking regions of the human MW and LW genes, therefore, indicating that the duplication in the howler monkey differs from that in Old World primates. A 350-bp fragment of the region immediately downstream of the LCR was amplified successfully from howler monkey DNA by using a primer to a conserved region of the marmoset and human LW upstream sequences at position −2896. Cloning and sequencing yielded two distinct sequences that are identical in the LCR core sequence but differ by two nucleotide substitutions in the less conserved 3′ flanking region (Fig. 6). We conclude from this that there are two X-linked LCRs in the howler monkey. Therefore, the duplication in this species must include the entire 5′ flanking region of the gene and the LCR. Finally, by using the howler monkey sequence data downstream of the LCR and upstream of the transcription start site to design PCR primers AL− and AL+ (Table2), a fragment of ∼2.8 kb that overlapped with both sequences was amplified and cloned. Sequencing of the ends of these clones enabled us to associate, respectively, the upstream sequences 1 and 2 shown in Figure 5 with LCR sequences 1 and 2 shown in Figure 6. The additional sequence that this generated is included in Figure 6.
Nucloetide sequence of the upstream region of the X-linked opsin genes in the howler monkey, aligned with the equivalent region of the marmoset gene. Identity of howler sequence 2 and marmoset to howler sequence 1 is indicated by a dot. Sequence deletions are indicated by dashes. Howler monkey sequence, GenBank accession nos. AF156715 andAF156716.
Nucleotide sequence of the two locus control regions identified in the howler monkey, aligned with the corresponding region in the marmoset. The numbering of the marmoset sequence corresponds to the full sequence shown in Fig. 3. Identity of the other two sequences to howler sequence 1 is indicated by a dot.
Sequence and Position of PCR Primers
DISCUSSION
Trichromacy in all Old World primates depends on the presence of two separate X-linked opsin genes that encode a MW and LW opsin, with λmax values of ∼530 and 560 nm, respectively (Bowmaker et al. 1991). The close homology between these two genes indicates that they represent a relatively recent duplication from a single ancestral gene (Hunt et al. 1998) and the presence of the duplication in all Old World primates examined to date (Ibbotson et al. 1992; Deeb et al. 1994; Dulai et al. 1994) places the duplication event at the base of the catarrhine lineage ∼40 mya (Goodman et al. 1998). A recent analysis of the coding sequence of these two genes would suggest that the evolution of the spectral shift between the visual pigments encoded by these two genes occurred subsequent to duplication and did not draw on different allelic forms of the polymorphic gene now present in New World primates (Hunt et al. 1998). The single X-linked opsin gene that is present in most New World monkeys is clearly orthologous to both genes in Old World species (Ibbotson et al. 1992; Williams et al. 1992;Dulai et al. 1994; Hunt et al. 1998) and therefore, most likely represents the ancestral organization before gene duplication.
Previous sequence analysis of the 5′ flanking regions of the human MW and LW genes has demonstrated that the two regions are homologous for only 236 bp immediately upstream of the translation start site (Nathans et al. 1986). This raises the question of the origin of the different sequences beyond this region. By sequencing the 5′ flanking region of the marmoset, a New World monkey with a single X-linked opsin gene and a polymorphism-based trichromacy, we have been able to demonstrate that this region shows extensive homology to the human LW gene but not to the human MW gene. In fact, the homology between the marmoset and the human MW sequences is limited to the same 236 bp immediately upstream of the translation start as found for the two human sequences. From this, we can deduce that the duplicated opsin gene in Old World primates includes only this short upstream region, and that the 5′ flanking region of the human LW gene represents the ancestral sequence.
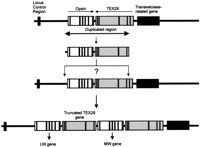
Recent work by Hanna et al. (1997) has demonstrated that, in humans, a novel gene called TEX28, which is expressed only in the testes, lies at the 3′ end of the opsin gene array on the X chromosome. A truncated copy of this gene is also present immediately downstream of the opsin gene closest to the LCR, and any additional copies of MW genes are also flanked by truncated copies ofTEX28. It is only the final opsin gene at the 3′ end of the array that is flanked by a complete copy of TEX28. Therefore, the duplication in Old World primates encompasses a 35-kb region that includes a complete copy of the opsin-coding region, 236 bp of the upstream flanking sequence, and a truncated copy of theTEX28 gene. With the exception of the opsin gene adjacent to the LCR, all other genes within the array now possess a minimal promoter region of ∼190 bp of the original flanking region upstream of the transcription start site. This duplicated region was inserted either just upstream of the original opsin gene or into the first intron of the TEX28 gene, as depicted in Figure7. If the former alternative is correct, then the duplication must have occurred by an unequal crossover mechanism that avoided any sequence duplication or deletion at the point of insertion, as there is no evidence for any indels in the marmoset and human LW sequences in the immediate vicinity of the insertion. The preservation of the exact sequence might have been less important if the insertion was into the intron of the TEX28 gene. It has been suggested that the LW opsin was most probably ancestral in primates (Nei et al. 1997). This does not, however, distinguish between the above two alternatives as, subsequent to duplication, evolution of a MW sequence could have occurred in either gene and it does not follow that the present LW gene was the ancestral sequence.
Origin of the opsin gene duplication on the X chromosome of Old World primates. The opsin and TEX28 genes are shown as open boxes showing the approximate positions of exons and introns. The LCR and transketolase-related gene are shown as filled boxes. Details of theTEX28 and transketolase-related genes are from Hanna et al. (1997).
A striking feature of the upstream region of the human MW gene beyond the region of homology is the presence of a number of Alurepeat elements, and at least the first two of these are also present in identical positions in the upstream region of the MW gene of the diana monkey. This would indicate that the juxtapositioning of these elements with the MW opsin gene occurred at the same time as the gene duplication rather than as a result of a more recent insertion. The first of these elements, a half Alu, is found at the point of insertion of the original duplication. Because Alu repeat elements have been implicated in unequal crossing over within other genes (e.g., see Campbell et al. 1995; Rudiger et al. 1995; Harteveld et al. 1997; Levran et al. 1998) and in the insertion event that resulted in the “long” intron 1 of the human LW opsin gene (Meagher et al. 1996), it is possible that the presence of Alu sequences was a contributory factor in the generation of the original opsin gene duplication, and in the generation of additional copies of the MW gene (Drummond-Borg et al. 1989; Ibbotson et al. 1992). By the same mechanism, these sequences may also be important in promoting unequal crossing over within the opsin gene array that leads to either the complete loss of opsin genes or the generation of hybrids. Such changes are considered to be responsible for anomalous trichromacy and dichromacy in humans (Nathans et al. 1986; Deeb et al. 1992).
The close proximity of the duplication insertion breakpoint to the transcription start site of the MW gene would imply that the promoter for this gene is contained within the remaining 199 bp of upstream sequence. This has been confirmed by an in vitro study of the activity of this region in driving the expression of the human MW and LW genes in transfected WERI Rb1 cells, a retinoblastoma cell line that is known to express cone opsins (Shaaban and Deeb 1998). The minimal promoter, numbered from immediately upstream of codon one, was defined in this study as the region from −64 to −1, with positive elements within the region from −83 to −171, and negative elements from −171 to −240. This study was unable to provide, however, any information on the mechanism of selective gene expression that ensures that only one visual pigment is produced in a single cone photoreceptor, although differences in the activity of the two promoters were clearly apparent. Given the divergence of both promoters from that of the marmoset, it is tempting to speculate that the control of differential gene expression in Old World primates may also reside in this region, and our finding that all the upstream differences, except that at position +1 within this minimal promoter region in the human and chimpanzee MW and LW genes, are conserved in two species of cercopithecoid monkeys is supportive of this interpretation.
The presence of an LCR upstream of the single X-linked opsin gene in the marmoset is consistent with the idea that this highly conserved region is required for the activation of gene transcription in this chromosomal region. It has been suggested that the LCR interacts by a looping mechanism with the adjacent gene promoter to form a transcriptionally active chromatin domain (Festenstein et al. 1996;Milot et al.1996) through protein–protein interactions of transcription factors that bind to both the promoter and LCR (Shaaban and Deeb 1998). In this case, sequence differences in the MW and LW opsin promoters may determine specificity of gene expression through the differential binding of such transcription factors. However, this would require a further level of gene regulation and the alternative possibility that the interaction of the LCR with one or other promoter region is mutually exclusive, provides a simpler explanation. The presence of an LCR in New World monkeys argues, however, that the primary role of this region is not the selection of either a MW or LW opsin gene for expression as only a single gene is present in most New World primates; the presence of the LCR must therefore predate the Old World opsin gene duplication.
The howler monkey is unique among New World primates in possessing separate MW and LW genes. Therefore, in this respect, it resembles the Old World primates. The presence of flanking regions to the two X-linked opsin genes in this species that differ at a small number of sites was first reported by Kainz et al. (1998) and the more extensive sequence analysis of this region presented here confirms their observation. The extensive identity within the 620-bp region immediately upstream of the howler genes contrasts with the divergence of the upstream region within 237 bp of the translation start site for the human MW and LW genes. The howler monkey and Old World primate duplications are, therefore, quite distinct and the homology in this region between the two howler genes, with only four single nucleotide differences in >600 bp of DNA, indicates that the howler monkey duplication is much more recent than that in Old World primates. In addition, we have been able to clone and sequence two distinct LCRs from the DNA of a single male howler monkey that differ by two nucleotide substitutions in the 3′ flanking region of the conserved core sequence of the LCR, and to link these two LCR sequences to the two sequences of the immediate upstream region, thus defining two complete 5′ flanking regions of the howler opsin genes. Therefore, it would appear that the duplication in the howler monkey involved not only the complete coding region of the opsin gene but also the entire upstream region, including the LCR.
In typical New World monkeys with a single polymorphic X-linked opsin gene, trichromacy is achieved in heterozygous females by random X inactivation. In contrast, separate MW and LW opsin genes will generate trichromacy only if a mechanism exists to limit expression to only one gene per cone photoreceptor. The presence of two LCRs in the howler monkey would indicate that the mechanism of selective opsin gene expression may differ from that in Old World primates. Whether the limited sequence differences within the promoter regions and LCRs of these two genes in howler monkeys are sufficient for this process remains to be determined.
METHODS
DNA Samples
DNA was isolated from a blood sample taken from a male black howler monkey Alouatta seniculus. DNA was isolated from liver tissue of a male marmoset Callithrix jacchus, with the P563 allele of the X-linked opsin gene, using a standard phenol–chloroform method as described previously (Williams et al. 1992). DNA from the two cercopithecoid monkeys, the diana monkey Cercopithecus dianaand the patas monkey Erythrocebus patas, was isolated as described by Ibbotson et al. (1992), and from the chimpanzee as described by Dulai et al. (1994).
Amplification of the 5′-Flanking Regions of Opsin Genes
The sequences of all primers used in PCR amplifications are listed in Table 2.
Human and Diana Monkey MW Gene
Three clones containing the upstream region of the human MW gene were isolated from a human cosmid library (Cachon-Gonzales 1991), using exon 1 of the human MW gene as a probe. The presence of the MW coding region was confirmed by PCR amplification of exon 2. One of these clones was prepared for sequencing by digestion with EcoRI, followed by the ligation of EcoRI Vectorette 1 linkers. The Vectorette 1 primer was then used in combination with a gene-specific primer (Op1−) to generate a fragment of ∼2.3 kb. This was direct sequenced using Op1− as a sequencing primer. The sequenced region was extended in two more steps to −1185 through the use of two additional sequencing primers designed from the sequenced region. Approximately 750 bp of the upstream region of the diana monkey MW gene was amplified using primer pair MW− and MW+. The resulting fragment was cloned and sequenced.
Marmoset MW/LW Gene
A combination of conventional PCR and gene walking was used to amplify the upstream region of the marmoset gene. Gene walking allows unknown flanking regions to be amplified from a known sequence. Initially, a 245-bp fragment that included the first exon and extended upstream to −151 bp was amplified with primers Op2+ and Op1−, designed from the X-linked human opsin genes (Nathans et al. 1986). This sequence was then extended upstream to −205 bp, using an unpredictably primed PCR developed by Domínguez and López-Larrea (1994) and modified by Bellingham et al. (1998). Gene-specific nested primers used in the gene walk were Op1−, Op2−, and the nested “universal” walking primers were UNI-33 and UNI-17. The first round UP–PCR-1 mix contained 10 ng of DNA, 12.5 pmoles of primer UNI-33, 0.5 nmole each of dATP, dCTP, dGTP, and dTTP, 3 mm MgCl2, and 5 μl of 10× NH4buffer in a total volume of 23 μl. The mix was heated at 94°C for 60 sec to denature the DNA, during which 0.25 units of Taqpolymerase was added in a volume of 1 μl. The incubation temperature was then lowered to 80°C for 30 sec, then rapidly to 15°C for 120 sec, and finally held at 25°C for 10 min. The sample tubes were then placed in a thermal cycler, initially at 72°C for 60 sec before DNA denaturation at 94°C for 60 sec during which 12 pmoles of the outer Op1− primer was added in a volume of 1 μl. The sample was then heated at 94°C for 1 sec, 62°C for 30 sec, and 72°C for 30 sec for 35 cycles, followed by a final extension at 72°C for 5 min. The UP–PCR-2 contained 1 μl of 1:1000 (vol/vol) dilution of the UP–PCR-1, 12.5 pmoles each of primers Op2− and UNI-17, 0.5 nmole each of dATP, dCTP, dGTP, and dTTP, 3 mm MgCl2, and 5 μl of 10× NH4buffer in a total volume of 24 μl. Taq polymerase (0.25 units) in 1 μl was added during the DNA denaturation step, as done previously, and the sample cycled at 94°C for 1 sec, 56°C for 1 sec, and 72°C for 30 sec for 35 cycles, followed by a final extension at 72°C for 5 min.
A further round of UP–PCR with a new set of primers [Up(A−) and Up(B−)] together with UNI-33 and UNI-17 generated a fragment of ∼550 bp, which extended upstream to base −659. On the basis of the sequence obtained from this fragment, further primers [NWM LCR−and MarUp(A+)] were designed to allow amplification both upstream and downstream back into the gene. Primer NWM LCR− was combined with primer LCR(C+) to amplify a fragment of ∼3000 bp, which extended from the LCR core to a region in the proximal promoter. This fragment was fully sequenced using a set of nested primers [MarLCR(C−), MarLCR(D+), MarLCR(E+), MarLCR(D−), and MarLCR(E−)].
The Conserved Region of the Chimpanzee, Diana Monkey, and Patas Monkey MW and LW Genes
The sequence of the LW gene of the chimpanzee, diana and patas monkeys was obtained by gene walking using Op1− and Op2− as gene-specific primers. This generated fragments of ∼400 bp that were both direct sequenced using Op2− and UNI-17 as sequencing primers, and sequenced after cloning. The MW sequences were obtained by conventional PCR using reverse primer Op2− and forward primer Op1+ that sits at the boundary of the region of homology between the MW and LW genes. The fragments were both directly sequenced and sequenced after cloning.
Howler MW and LW Genes
The upstream regions were initially amplified as two discrete blocks by conventional PCR using primer pairs MarUp(A+) with Op2−, and LCR(C+) with Up(C−), using standard PCR parameters with annealing temperatures of 61°C and 57°C, respectively. Fragments of ∼700 bp and 900 bp, respectively, were generated by these amplifications. Each fragment was both directly sequenced and sequenced after cloning. A primer pair (AL− and AL+) was designed from these two sequences that would amplify the intervening region of ∼2.8 kb. The resulting fragment was cloned and a ∼350-bp region at each end was sequenced.
DNA Cloning and Sequencing
All amplified fragments were sequenced directly on an Applied Biosystems model 373a automated sequencer. Before cycle sequencing, the PCR mix was cleaned with a Centricon (Princeton, Inc.) column. Approximately 100 ng of DNA was combined with 4 μl of ABI PRISM Dye Terminator and 3.5 pmoles of an appropriate sequencing primer, and cycled on a Perkin Elmer 9600 PCR thermal cycler. Selected PCR fragments were cloned into pTAg or pGEM-T-Easy, before cycle sequencing. Alignments were performed using GeneWorks.
Acknowledgments
This work was supported by a grant from the Medical Research Council. We are grateful to Dr. C. Julliot for the sample of howler monkey DNA.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
Footnotes
-
↵3 Corresponding author.
-
E-MAIL d.hunt{at}ucl.ac.uk; FAX 0171 608 6863.
-
- Received November 10, 1998.
- Accepted May 25, 1999.
- Cold Spring Harbor Laboratory Press