The Mouse Aire Gene: Comparative Genomic Sequencing, Gene Organization, and Expression
- Karin Blechschmidt1,
- Michal Schweiger2,
- Karin Wertz3,
- Richard Poulson4,
- Hoang-My Christensen2,
- Andre Rosenthal1,
- Hans Lehrach2, and
- Marie-Laure Yaspo2,5
- 1Institute of Molecular Biotechnology, Department of Genome Analysis, D-07745 Jena, Germany; 2Max-Planck Institute for Molecular Genetics, D-14195 Berlin-Dahlem, Germany; 3Max-Planck Institute for Immunobiology, D-79108 Freiburg, Germany; 4Imperial Cancer Research Fund; Histopathology Unit, London, UK
Abstract
Mutations in the human AIRE gene (hAIRE) result in the development of an autoimmune disease named APECED (autoimmunepolyendocrinopathycandidiasis ectodermaldystrophy; OMIM 240300). Previously, we have cloned hAIRE and shown that it codes for a putative transcription-associated factor. Here we report the cloning and characterization of Aire, the murine ortholog of hAIRE. Comparative genomic sequencing revealed that the structure of the AIRE gene is highly conserved between human and mouse. The conceptual proteins share 73% homology and feature the same typical functional domains in both species. RT–PCR analysis detected three splice variant isoforms in various mouse tissues, and interestingly one isoform was conserved in human, suggesting potential biological relevance of this product. In situ hybridization on mouse and human histological sections showed that AIRE expression pattern was mainly restricted to a few cells in the thymus, calling for a tissue-specific function of the gene product.
There is a wide range of human autoimmune diseases, but the molecular background of autoimmunity remains poorly understood (Ollier 1992). Despite the identification of a number of genetic susceptibility factors, the etiology of most autoimmune diseases remains elusive. In this context, the study of autoimmune conditions with Mendelian inheritance could provide a boost for unraveling pathogenic pathways involved in human autoimmunity.
Autoimmune polyendocrinopathy candidiasis ectodermal dystrophy [APECED, Online Mendelian Inheritance in Man (OMIM) 240300] is an autosomal recessive disease resulting in a variable combination of failure of the parathyroid glands, adrenal cortex, gonads, and pancreatic β cells (Ahonen 1985). Ectodermal dystrophies, vitiligo, and chronic mucocutaneous candidiasis are also frequently observed among APECED patients (Ahonen et al. 1990; Perheentupa 1996). APECED is a rare disease particularly enriched in genetic isolates, such as the Finnish population, Iranian Jews, and Sardinians (Ahonen 1985;Zlotogora and Shapiro 1992; Clemente et al. 1997). After demonstration of genetic linkage and locus homogeneity on chromosome 21q22.3 (Aaltonen et al. 1994; Bjorses et al. 1996), the APECED gene was cloned and called AIRE for autoimmuneregulator (The Finnish–German APECED Consortium 1997;Nagamine et al. 1997).
The AIRE gene encodes a nuclear factor of unknown function that harbors two PHD zinc fingers—a modular domain found in many proteins involved in chromatin-mediated regulation of transcription (Schindler et al. 1993; Aasland et al. 1995). Moreover, striking structural similarities were observed between AIRE and human Sp100/Sp140 proteins, suggesting that they derive from a common ancestor. In addition to the zinc fingers, AIRE and Sp140 also share a putative DNA-binding domain called SAND domain and a stretch of 90 amino acids in their amino-terminal region (Gibson 1998). Sp100 proteins localize to specific nuclear structures called nuclear bodies and represent a target of autoantibodies in patients with primary biliary cirrhosis (Szostecki et al. 1990).
As a first step toward investigating AIRE biochemical properties and for engineering a mouse model for APECED, we have cloned and characterized the murine AIRE homolog. Here we report comparative genomic sequence analysis of the AIRE loci andAIRE expression pattern on mouse and human histological sections.
RESULTS
Identification of the Mouse Aire Gene
We have isolated the mouse homolog of the human AIRE gene by cross-species screening of mouse genomic libraries with a human cDNA containing the complete AIRE coding sequence [B1-1pA (The Finnish–German APECED Consortium 1997), referred to here as hAIRE]. Six positive clones were isolated: one PAC (RPCIP711H2150), four P1s (ICRFP703A23152, A10129, G23152, and J2183), and one cosmid (MPMGc121L12287). After restriction digest withEcoRI and hybridization with hAIRE, all clones showed a similar pattern of four EcoRI fragments totaling a size of 20.6 kb, except for P1 A10129, which showed an AIRE pattern of only 13.54 kb (data not shown). Hybridization with the most 5′ end and 3′ ends of the human cDNA indicated that A10129 was missing at least the first exon, whereas the five other genomic clones contained the whole Aire gene (data not shown).
Comparative Genomic Sequencing and AIRE Gene Organization
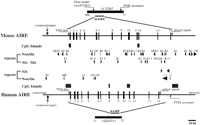
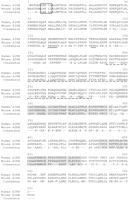
We have sequenced cosmid MPMGc121L12287 (GenBank accession no.AF073797) and deduced the mouse Aire gene structure by comparative analysis with the previously published hAIRE locus (cosmid LLNCO22G11; EMBL accession no. HSAJ9610). L12287 contained the 14 Aire exons spanning 13,276 bp from the proposed initiation codon to the termination codon. This compares with a length of 11,714 bp for the human gene (Fig. 1). The mouse Aire intron/exon boundaries were confirmed experimentally after alignment of L12287genomic sequence and mouse cDNA sequence (see below) using the EST:GENOME program (Mott 1997). In both species, splice acceptor and donor sequences were found to conform to the GT-AG rule, and the intron phase is completely conserved (Table 1). The GC content of AIRE coding sequence is 61% in mouse versus 68% in human. Genomic information was analyzed by first-pass automatic annotation using the Rummage package (http://genome.imb-jena.de/rummage.html). Features conserved between the two loci include a CpG island overlapping with AIRE first exon and a potential promoter associated with a TATA box located 200 bp upstream of the proposed translation initiation site (Fig. 1). Two other genes were identified in the mouse cosmid, the PFKLpromoter and a novel C2H2 zinc finger gene predicted in silico 6 kb proximal to AIRE on the opposite DNA strand (Fig. 1). This gene model is incomplete but shows significant EST matches (GenBank accession no. AA413561) and strong homology (78%) with a human trapped exon previously located 60 kb proximal from thePFKL promoter on 21q22.3 (HC21EXc32; D86111) (Kudoh et al. 1997). PFKL is 3 kb telomeric to AIRE in human (Fig.1), and data indicate that the linkage group HC21EXc32, AIRE,PFKL is conserved in human and mouse. To detect potentially conserved elements, the murine and human sequences were plotted on a dot matrix using the DOTTER program (Fig. 2a; Sonnhammer and Durbin 1995). Most of the exons were identified, albeit exons 4, 7, and 10 are barely distinguishable because of to their shorter size (exons 4 and 7) or sequence divergence. Interestingly, a highly conserved region of 90 nucleotides was identified 3 kb upstream of the AIRE first exon, displaying a stretch of 40 nucleotides with 80% identity (Fig. 2A,B). This region sharing no homology with other known regulatory elements may have a role in modulating AIRE expression, but this hyppothesis has to be formally demonstrated.
Comparative genomic organization of the AIRE locus. Exons are represented by solid boxes numbered from 1 to 14. Repetitive elements are depicted by arrowheads. CpG islands are represented by solid boxes. Putative TATA box promoter and conserved region are indicated by arrows.
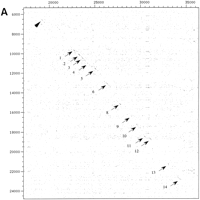
Human and Mouse Gene Structure Information
(A) Dot-matrix comparative analysis of hAIRE and mouseAire genomic sequences: HSAJ9610 is represented on thex-axis and AF073797 on the y-axis. Arrows denote exons; the arrowhead points to a conserved region located ∼3 kb upstream of the gene. (B) Alignment of the conserved nucleotide sequence identified in A. Numbers at the end of lines indicate nucleotide positions. Consensus sequence is drawn below the alignment. Box corresponds to the core conserved sequence.
Localization of Aire to Chromosome 10
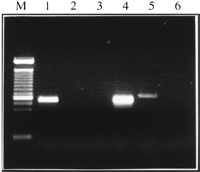
Comparative mapping between mouse and human has shown that human chromosome 21q22.3 shares conserved synteny with mouse Chromosomes 10 and 17 (Irving et al. 1994). The chromosomal localization ofAire was determined by PCR analysis of monochromosomal hybrids containing mouse Chromosome 10 or 17. The primer set Mforw2/Mrev32 amplified a specific product of the expected size in total mouse genome and Chromosome 10 DNAs (Fig. 3), in agreement with the expected conserved synteny in this region around the Pfkllocus (Irving et al. 1994).
Mapping of Aire to mouse Chromosome 10. Electrophoresis of PCR amplification of mouse DNAs with primers Mforw2 and Mrev32.(Lane1) Hybrid SN17C3 (Chr. 10), (Lane 2) SN11CS3 (chr. 3); (Lane 3) EJ167 (chr. 3+17); (Lane 4) mouse genomic DNA; (Lane 5) human genomic DNA; (Lane 6) no DNA. (M) 100-bp ladder (Life Technologies).
The Predicted Mouse Aire Protein
Genomic information allowed in silico characterization of the murine cDNA sequence and corresponding conceptual protein. Nucleotide sequence identity between mouse and human AIRE coding sequences is 77%. hAIRE encodes a 545-amino-acid protein. The predicted mouse Aire protein is 552 residues with a calculated pI of 8.43 and a theoretical molecular mass of 59 kD. The overall identity between the mouse and human AIRE proteins is 73% and similarity is 76% (Fig.4). The two proteins appear remarkably conserved and harbor the same modular domains: a SAND domain, two PHD zinc fingers, a LXXLL motif, which is a signature for nuclear receptor binding site (Heery et al. 1997), and a nuclear targeting signal (Fig. 4).
Amino acid aligment of hAIRE and mouse AIRE proteins. The LXXLL motif is shown by an open box; the nuclear localization signal is underlined; the SAND domain is shown by a broken line. Shaded boxes indicate the PHD zinc fingers.
AIRE Gene Expression
Using primers designed from genomic sequence information, mouseAire cDNA fragments were isolated by PCR amplification of a cDNA source prepared from ES cells. A cDNA sequence of 2015 bp deduced from overlapping PCR products contained an open reading frame (ORF) of 1656 bp (GenBank accession no. AJ132243). Northern blot analysis using a PCR product spanning exons 1–7 failed to detect any transcripts in the panel of mouse tissues analyzed containing heart, brain, spleen, lung, liver, skeletal muscle, kidney, and testis, indicating thatAire is seldom expressed in these tissues (data not shown). The screening of EST databases using BLAST (Altschul et al. 1990) identified only one partially processed cDNA from a 4-week mouse thymus (GenBank accession no. AA866822).
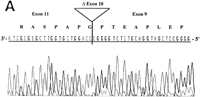
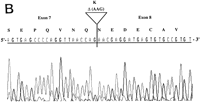
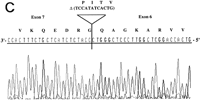
RT–PCR amplification was performed on a panel of mouse normalized first-strand cDNAs. Sequencing of cloned PCR products indicated the presence of Aire transcripts at 11 dpc and in adult heart, spleen, lung, skeletal muscle, and testis. Three potentially functional alternatively spliced transcripts (type I, II, and III) were seen in some tissues (Table 2). Type I isoform corresponds to the skipping of exon 10 (Fig. 5A). Type II splice variant shows a 3-bp deletion at the splice acceptor site in exon 8 (Fig. 5B), leading to a predicted protein lacking Lys-296. Type III isoform has an in-frame 12-bp deletion at exon 6 splice donor site, and the putative peptide is lacking Val-265, Thr-266, Ile-267, and Pro-268 (Fig. 5C). No Aire transcripts could be obtained from 7 dpc, 17 dpc, or from adult brain or kidney. Control Hprt gene PCR amplification led to a single product of comparable intensity in all tissues from the panel (not shown). In human, direct sequencing of uncloned RT–PCR products generated from a panel of tissues (see Methods) identified the type II transcript in spleen and bone marrow (data not shown). However, our data did not address whether type I and type III isoforms were conserved.
Summary of the Sequenced RT–PCR Products
Differential splicing of Aire transcripts. (A) Deletion of exon 10. Sequence is reversed as indicated 3′ → 5′; (B) deletion of 3 nucleotides at the start of exon 8; (C) deletion of 12 nucleotides at the end of exon 6. Sequence is reversed as indicated 3′ → 5′.
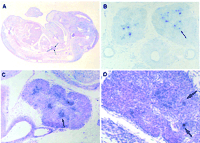
Spatial AIRE distribution was investigated by in situ hybridization on histological sections. In mouse embryo, Airecould be detected from 14.5 dpc, in which a peculiar pattern of expression was confined to a few cells in the developing thymus (Fig.6). The cells expressing Aire are located in the medulla of the organ anlage but cannot be correlated with a particular cell type. This restricted staining pattern could either reflect Aire expression in only a very limited subset of cells or for a very short period of time in a larger cell population. In human, the spatial expression profile was found comparable with a signal restricted to foci of cells in the lobule of juvenile thymus medulla (Fig. 7).
Expression of Aire at 14.5 dpc is restricted to few cells in the thymus. RNA in situ hybridization with Aire antisense riboprobe recognizing exons 1–7. No signal was detected upon hybridization with a sense probe.
(A) Sagittal section through 14.5 dpc. mouse embryo, counterstained with eosin. (B) Transverse section of thymic lobes of a 14.5 dpc. embryo. (C) Sagittal section of 14.5 dpc. thymus, counterstained with eosin. (D) Sagittal section of 14.5 dpc. thymus, counterstained with hematoxylin and eosin, at higher magnification.
Arrows point to single cells or cell groups expressing Aire.
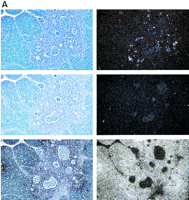
Expression of hAIRE in human juvenile thymus sections counterstained with Giemsa. Expression is restricted to a few cells in the medulla of the thymic lobule. (Top) The antisense probe; (middle) the sense probe; (bottom) the control β-actin probe. (Left) Bright field; (right) dark field. (A) Magnification, 100×; (B) another section with magnification, 500×.
DISCUSSION
We present here the cloning and characterization of the mouse ortholog of human AIRE, the gene causative for APECED disease. Comparative genomic sequencing indicated that the gene organization was highly conserved in human and mouse featuring 14 exons spanning 13 kb of genomic DNA, a TATA box promoter associated with a CpG island, and a potential controller element located 3 kb upstream of the first exon. The mouse and human AIRE genes are highly homologous at both the nucleotide and amino acid levels, and the two proteins contain similar structural hallmarks. By virtue of two PHD zinc fingers shared by a number of chromatin-associated transcriptional regulators, it was postulated that AIRE may have a role in gene regulation. PHD fingers are often found together with other functional domains, such as a RING zinc finger in KRIP-1 (Kim et al. 1996) or a helicase domain in the Mi2 autoantigen identified in some dermatomyositis patients (Ge et al. 1995; Seelig et al. 1995). AIRE’s closest structural homolog is Sp100, which localizes to discrete nuclear dots (Grotzinger et al. 1996;Zuchner et al. 1997). hAIRE protein localizes to speckled domains in the cell nucleus (Rinderle et al. 1999), and its murine counterpart probably exhibits a similar subcellular localization. However, AIRE function is as yet elusive even if it provides the third example of a PHD finger protein involved in autoimmunity.
It is of paramount importance to determine the temporal and spatial distribution of AIRE for gaining insights into its primary function. Controversy revolved around the expression of humanAIRE assessed previously by Northern blot. We reportedAIRE expression as a 2-kb cDNA in a range of tissues with most prevalent expression in thymus, pancreas, and adrenal cortex, using a probe spanning exons 2–5 or with the whole cDNA (The Finnish–German APECED Consortium 1997). However, we identified a strong 2.4-kb signal in fetal liver but not in other tissues, using a probe spanning exons 11–13 (M.L. Yaspo, unpubl.). Nagamine et al. (1997) reported a cDNA of 2.4 kb in fetal liver, and several transcripts of 2, 3, and 4 kb in lymph node and thymus, using a probe spanning exons 12–14. The difference between these results can be explained in part by GC-rich regions found in the 5′ end of the probe, which may hybridize with nonlegitimate transcripts. In mouse, Northern blot analysis failed to detect Aire in the tissues analyzed, indicating a rare or restricted expression profile. In human, AIRE protein expression investigated by Western blot analysis failed to detect the gene product in a range of fetal and adult tissues that included organs affected in APECED, such as thyroid and parathyroid (M.L. Yaspo et al. unpubl.) Although we (M.L. Yaspo et al. unpubl.) and others (Nagamine et al. 1997) detected AIRE mRNA expression in human fetal liver, this could not be confirmed by Western blot. Correlation may be difficult to draw from different samples if the gene product is expressed during a short period of time and/or at a particular developmental stage. Interestingly, in situ hybridization performed at 14.5 dpc in the mouse indicated that Aire is expressed in only a few cells of the thymus, which are probably located in the medulla. Analysis of histological sections originating from human juvenile thymus corroborate this observation. Taken together, data confirm that AIRE is seldom expressed in most tissues. In situ hybridization data would explain the RT–PCR negative results at 17 dpc on whole mouse embryo, considering the very low proportion of cells expressing the gene. RT–PCR analysis detected three potentially functional isoforms. These variants occur with a relatively high frequency in independent PCR reactions and are unlikely to represent artifacts. Isoform type I would lead to an in-frame deletion of 59 residues between the two PHD fingers. Examples of such splice variants in zinc finger proteins have been reported previously. For instance, alternate splicing isoforms of WT1 occurring in the hinge region spacing two Krüppel zinc fingers are associated with differential subnuclear localization (Larsson et al. 1995). However, the significance of Aireisoforms remains to be addressed formally, sensitivity of RT–PCR may reflect residual activity of a “leaky” promoter rather than true physiological expression. In situ analysis of the expression pattern on histological sections appears to be the most informative approach for tackling the temporal and spatial AIRE expression pattern. Identification of those cells expressing AIRE in the thymus will be of fundamental relevance for shedding light onto some of the pathological mechanisms leading to autoimmunity.
Mutations in AIRE represent the primary genetic defect leading to APECED, presumably because of a defective AIRE protein. AIREexpression profile in embryo and adult tissues suggest that if proven to act as a mediator of transcription, AIRE is not a global transcription factor but rather is involved in modulating the expression of tissue-specific genes, for example, in the thymus. Highly conserved protein structure and similar spatial expression profile in the thymus argue for a comparable function of AIRE in human and mouse. Characterization of the Aire murine ortholog will thus provide a tool for exploring AIRE function and for engineering a murine model of APECED
METHODS
Isolation of Mouse Aire Genomic Clones
Mouse clones were screened by hybridization of mouse genomic libraries with a human cDNA probe containing the complete AIREcoding sequence. Six positive mouse clones were isolated: PAC RPCIP711H2150 (129/SvevTACfBr); P1 clones ICRFP703A23152, A10129, G23152, and J2183 (C57/Black6); and cosmid MPMGc121L12287 (129/Ola).
Genomic Sequencing
Cosmid DNAs were isolated using a standard lysis method and purified on a CsCl gradient. DNA was sonicated, size fractionated, and ligated into M13 vector for shotgun sequencing using Thermo Sequenase (Amersham) and dye-terminator chemistry (Perkin Elmer). Data were collected using ABI 377 automated sequencers and assembled with Gap4 (Staden 1996). Gaps were closed by resequencing the M13 templates with ET (Energy Transfer) dye primers (Amersham).
Computer Analysis
Repeats were identified with the Repeat masker programhttp://ftp.genome.washington.edu/RM/RepeatMasker.html); (A.F.A Smit and P. Green). Homology searches were performed using BLAST version 1.4 (Altschul et al. 1990) and FASTA version 2.0 (Pearson and Lipman 1988). Programs GRAIL2 (Uberbacher and Mural 1991), XPOUND (Thomas and Skolnick 1994), MZEF (Zhang 1997), and GENSCAN (Burge and Karlin 1997) were used for exon prediction. Promoter predictions were done with Promoter Scan II (Prestridge 1995) and Transcription Start Site using both Ghosh/Prestridge (TSSG) and Wigender (TSSW) motif databases (http://dot.imgen.bcm.tmc.edu:9331/gene-finder/gf.html); (V.V. Solovyev, A.A. Salamov, and C.B. Lawrence). Dot matrix comparison was performed on a DEC-α station using the DOTTER program (Sonnhammer and Durbin 1995), and analysis was done using set default parameters.
RNA and RT–PCR analysis
Analysis of Mouse Aire
A Northern blot containing 2 mg of poly(A)+ RNA was purchased from Clontech. RT–PCR analysis was performed on ES cells cDNA and on a normalized first-strand cDNA panel from mouse multiple tissues (Clontech). Primers Mforw4 (5′-TGGCAGGTGGGGATGGAA-3′) and Mrev15 (5′-GGAGGGATGGAAGGGGAGGA-3′) amplified a product spanning exons 1–7. PCR reactions were performed in a Biometra UNO II thermocycler. An initial denaturation at 94°C for 2 min was followed by 35 cycles at 94°C for 45 sec, 56°C for 40 sec, 72°C for 1 min, and a final extension at 72°C for 5 min. Primers Mforw6 (5′-AAAGCCAGTGGTCCGAGCCAA-3′) and Mrev34. (5′-GGAAGTGGCAGCGCCAGT-3′) amplified exons 6–11. Primers Mforw7 (5′-TGGTCCGAGCCAAGGGAG-3′) and MR4 (5′-GCCACCTGTCATCAGGAAGAG-3′) were used to amplify a cDNA fragment spanning exon 7–14 and extending in the 3′ direction outside of the translated region. Conditions for all PCRs were basically identical with the exception of the annealing temperature specific for each primer pair.
Analysis of hAIRE
RT–PCR analysis was performed on a normalized first-strand cDNA panel from human immune system tissue (Clontech). Primers B127FR4-21 (5′-GGCTTCTGAGGCTGCACC-3′) and B127FR4-29 (5′-GCTCTGGATGGCCTACTG C-3′) were used to amplify a 1.6-kb fragment. Nested PCRs were performed using primers B127FR4-17 (5′-AGAAGTGCATCCAGGTTGGC-3′) and B127FR4-33 (5′-GTGTGCTCGCTCAGAAGGG-3′) and products were sequenced directly.
Sequencing of PCR Products
Products from PCR amplifications were purified using the QIAquick PCR Purification Kit (Qiagen). Purified products were sequenced using the dye-terminator chemistry on an ABI 377 automated sequencer (Perkin Elmer).
Chromosomal Localization of Aire
PCR amplifications were performed using mouse-specific primers Mforw2 (5′-TCCCACCTGAAGACTAAGC-3′) and Mrev32 (5′-TCACAGCTC TCTGGACAGAA-3′) on hybrids SN11CS3 and SN17C3 containing mouse chromosome 3 and 10, respectively (Sabile et al. 1997) and hybrid EJ167 containing mouse chromosomes 17 and 3 (Cox et al. 1991). PCR reactions were performed in a Biometra UNO II thermocycler. Initial denaturation at 94°C for 2 min was followed by 35 cycles at 94°C for 45 sec, 51°C for 40 sec, 72°C for 2 min, and a final extension at 72°C for 5 min.
In Situ Hybridization
In situ hybridization on mouse sections was performed essentially according to Henry et al. (1996). The cloned RT–PCR product used for riboprobe synthesis recognizes all splice variants and spans exons 1–7. Hybridization stringency was 5× SSC, 50% formamide, at 65°C. Final washing stringency was 1× SSPE, 50% formamide, at 50°C. The sections were stained for 2–3 days with BM-Purple (Boehringer). Counterstaining with eosin or with hematoxylin and eosin was performed according to standard procedures. In situ hybridization on human sections was performed with 35S radiolabeled riboprobe spanning human exons 7–14 (sense and antisense), using a standard protocol described previously (Poulsom et al. 1988; Senior et al. 1988;). Sections were 4 μm thick from formaline, paraffin-embedded samples. Final washing stringency was 0.5× SSC at 65°C. Counterstaining with Giemsa was performed according to standard procedures. Exposure was for 31 days for AIRE and 10 days for β-actin control.
Acknowledgments
We thank the Resource Center Team (RZPD) for providing mouse library filters and genomic clones. We thank Dr. Tilman Vogel (UniKlinik, Duesseldorf) for providing tissue samples. We thank Dr. Heinz Himmelbauer for the mouse monochromosomal hybrid DNAs and Dr. Michael Wiles for providing ES cell first-strand cDNA. Margit Teuchtler and Bärbel Ukena are appreciated for excellent technical assistance. This work was supported by the Deutsche Human Genome Projekt (grants BMBF 01KW9608 and 01KW9617).
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
Footnotes
-
↵5 Corresponding author.
-
E-MAIL yaspo{at}impimg-berlin-dahlem.mpg.de; FAX 49-30-8413-1380.
-
- Received October 26, 1998.
- Accepted January 11, 1999.
- Cold Spring Harbor Laboratory Press