Abstract
We determined the genome-wide distribution of the nucleoid-associated protein Fis in Escherichia coli using chromatin immunoprecipitation coupled with high-resolution whole genome-tiling microarrays. We identified 894 Fis-associated regions across the E. coli genome. A significant number of these binding sites were found within open reading frames (33%) and between divergently transcribed transcripts (5%). Analysis indicates that A-tracts and AT-tracts are an important signal for preferred Fis-binding sites, and that A6-tracts in particular constitute a high-affinity signal that dictates Fis phasing in stretches of DNA containing multiple and variably spaced A-tracts and AT-tracts. Furthermore, we find evidence for an average of two Fis-binding regions per supercoiling domain in the chromosome of exponentially growing cells. Transcriptome analysis shows that ∼21% of genes are affected by the deletion of fis; however, the changes in magnitude are small. To address the differential Fis bindings under growth environment perturbation, ChIP-chip analysis was performed using cells grown under aerobic and anaerobic growth conditions. Interestingly, the Fis-binding regions are almost identical in aerobic and anaerobic growth conditions—indicating that the E. coli genome topology mediated by Fis is superficially identical in the two conditions. These novel results provide new insight into how Fis modulates DNA topology at a genome scale and thus advance our understanding of the architectural bases of the E. coli nucleoid.
The Escherichia coli genome forms a highly condensed structure called a “nucleoid body” (Robinow and Kellenberger 1994), whereas the genomic DNA in eukaryotic cells is packed in a nucleus as a chromatin structure (Kornberg 1974). The compact nucleoid body in a bacterial cell is extensively bound by several nucleoid-associated proteins, which include H-NS, HU, IHF, Fis, and the stationary-phase-specific DNA-binding protein Dps (Murphy and Zimmerman 1997; Azam et al. 2000; Schneider et al. 2001; Dame 2005). The involvement of the nucleoid-associated proteins in organizing the genetic material within the bacterial nucleoid has been widely accepted, as well as their involvement in regulating transcription (Ussery et al. 2001; Dorman and Deighan 2003; Blot et al. 2006).
The Fis protein is a general host nucleoid-associated DNA bending factor comprising 98 amino acids that was first identified because of its critical role in promoting site-specific DNA recombination (Johnson et al. 1986). The Fis protein contains a helix–turn–helix motif, which binds in the major groove and bends DNA by between 50° and 90° (Kostrewa et al. 1991; Pan et al. 1996). Its bending activity stabilizes DNA looping, either directly or through protein–protein interactions, to enhance transcription as well as to promote DNA compaction (Travers and Muskhelishvili 1998; Skoko et al. 2006). The intracellular level of Fis protein is growth-dependent and changes from less than 100 copies in stationary phase to more than 60,000 copies per cell in log phase (Ball et al. 1992; Azam et al. 1999). A variety of evidence suggests that the Fis protein plays a variety of roles in regulating DNA transactions and modulating DNA topology (Ussery et al. 2001). Recently, Fis has been implicated in the control of the gene expression involved in metabolism, transport, flagellar biosynthesis, and virulence in E. coli and Salmonella typhimurium (Kelly et al. 2004; Blot et al. 2006; Croinin et al. 2006). The regulation mechanism widely accepted is that Fis influences transcription by directly or indirectly affecting the activity of RNA polymerase and by modulating the level of DNA supercoiling in the cell. For example, at the promoters rrnB P1 and proP P2, Fis directly stimulates transcription by contacting the C-terminal domain of the RNA polymerase α subunit (RNAP α, also known as RpoA) (Bokal et al. 1997; McLeod et al. 2002). On the other hand, Fis negatively autoregulates its own operon by hindering RNA polymerase binding (Ninnemann et al. 1992). In the case of bacteriophage λ DNA excision, Fis appears to play an architectural role by contributing to a higher-order nucleoprotein complex that facilitates DNA cleavage and excision (Landy 1989).
There are 53 Fis-binding sites (Keseler et al. 2005) that have been directly experimentally determined. Robison and coworkers applied the recognition matrices developed from the experimentally derived Fis-binding sequences to search for Fis-binding sites across the E. coli genome sequence and reported more than 10,000 binding sites (Robison et al. 1998). Using hidden Markov models (HMMs), Ussery and coworkers reported 6000 strong Fis-binding sites in the E. coli genome (Ussery et al. 2001). Information analysis used by Hengen and coworkers estimated 68,000 Fis-binding sites, or one site per 230 bases (Hengen et al. 1997). The huge variance of Fis-binding sites predicted by three different computational methods reflects the fact that only weak binding site profiles are obtained when Fis-binding site sequences are aligned.
The relationship between the global effect of Fis on DNA topology and its local effects exerted on particular promoter regions is not well understood. The global interactions between the E. coli genome and Fis can be addressed by the direct measurement of Fis–DNA complexes by chromatin immunoprecipitation coupled with microarrays (ChIP-chip). The ChIP-chip approach is particularly well suited since unambiguous identification of the location of the proteins is possible by in vivo measurement of the protein–DNA complex (Ren et al. 2000). A recent genome-wide analysis of Fis association in E. coli cells identified 224 binding regions (Grainger et al. 2006) but was limited in the ability to define binding motif because of the resolution limitation of the low-density microarrays used. Here we improve on the resolution of this approach and use a ChIP-chip approach with fully tiled high-density microarrays to determine the distribution of the Fis-binding sites on a genome-scale. Our data enable the refinement of the Fis-binding motif and new insight into the functional behavior of the Fis protein. We also determined the effects of fis deletion on the transcription state of the cell.
Results
Immunoprecipitation of the DNA fragments associated with Fis, σ70, and RNAP
Prior to microarray hybridization, we used qPCR to determine the quality of immunoprecipitated DNA from the strain harboring myc-tagged Fis protein (BOP608), which has been shown to be highly resistant to stringent washing conditions and to retain its regulatory function in vivo (Cho et al. 2006a). The cross-linked DNA–protein complexes were immunoprecipitated by using antibodies against myc-tag, σ70 (also known as RpoD), or core RNAP (β subunit, also known as RpoB) from the cultured cells in the minimal media. Following reversal of DNA–protein cross-links, the immunoprecipitated DNA (IP DNA) was randomly amplified using PCR (Herring et al. 2005). In order to determine the enrichment of the IP DNA, qPCR was used to measure the relative levels of promoter and gene regions of known Fis-binding sites using nrfA, nirB, rrsA, sdhC, and dmsA as controls.
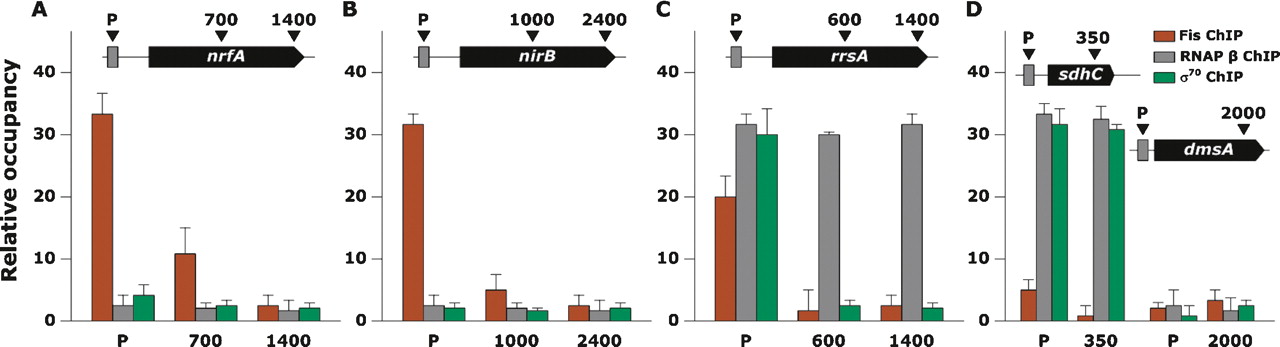
The relative occupancy of Fis at the promoter regions of nrfA, nirB, and rrsA was 34, 32, and 20, respectively (Fig. 1), which is consistent with previous studies (Wu et al. 1998; Browning et al. 2002; Paul et al. 2004). We also determined the association of σ70 and the core RNAP at the promoter and gene regions of nrfA, nirB, and rrsA under the same conditions. The association of σ70 and core RNAP was found only at the promoter of rrsA. Interestingly, the association of σ70 was only shown at the promoter, whereas the core RNAP was not only shown at promoter but at gene regions as well. These observations are in strong agreement with previous studies, such that nrf and nir operons are repressed by Fis (Wu et al. 1998; Browning et al. 2002), and Fis acts as a classical activator at the rrsA promoter (Paul et al. 2004). As control experiments, we determined the relative occupancy of Fis, σ70, and core RNAP at the promoters and gene regions of sdhC and dmsA (Fig. 1D). The Fis levels at promoters and gene regions of sdhC and dmsA remained at background levels. As expected, there was a large increase in σ70 and core RNAP association with the promoter and gene regions of sdhC due to its biological role in central metabolism under our growth conditions (Park et al. 1997). On the other hand, very low levels of σ70 and core RNAP were measured at the promoter and gene regions of the dmsA gene. This agrees with the known strong repression of the dmsA gene under the aerobic condition (Bearson et al. 2002). Altogether, these results demonstrate that Fis-bound DNA fragments were selectively immunoprecipitated from the exponentially growing E. coli cells.
Association of Fis with promoter regions of nrfA, sdhC, dmsA, nirB, and rrsA in mid-log growth phase under aerobic growth conditions. Relative occupancy on Y-axis represents the ratio of the immunoprecipitated DNA with and without antibodies using quantitative PCR. (A) In mid-log growth phase, Fis is present at the promoter region of nrfA, while the RNAP β levels remain at background levels. (B,C) Fis ChIP shows high occupancies at the promoter regions of nirB and rrsA. Owing to the transcriptional repression of the nirB gene under the conditions, RNAP β levels remain at background levels across the gene. However, rrsA is highly expressed in rapidly growing cells, so the RNAP β occupancy plateaus at the high level across the gene. (D) Fis is not associated with the promoter regions of sdhC and dmsA. RNAP β levels are high at the promoter and ORF regions of sdhC but are not present at the promoter regions of dmsA.
Genome-wide mapping of Fis-binding regions
To identify Fis-binding regions on a genome scale, we next performed a ChIP-chip analysis using custom-designed whole-genome tiling microarrays (NimbleGen) that contained a total of 371,034 oligonucleotides to represent the E. coli genome with 50-bp probes in overlapping by 25 bp on both forward and reverse strands (Herring et al. 2005). Our results identify regions of the genome enriched in the IP DNA samples, allowing us to construct a genome-wide map of in vivo interactions between Fis and the E. coli genome (Fig. 2A). Using a peak detection algorithm based on the double-regression model (Kim et al. 2005) together with manual curation, 894 unique peaks of Fis association were identified. The complete list of 894 Fis-binding regions is summarized in Supplemental Table S1.
Genome-wide mapping of Fis-binding regions in E. coli. (A) An overview of Fis-binding profiles across the E. coli chromosome at exponential state under aerobic growth conditions. The log2 enrichment ratio on the Y-axis was calculated from Cy5 (IP DNA) and Cy3 (mock IP DNA) signal intensity of each probe and plotted against each location on the 4.64-Mb E. coli chromosome on the X-axis. (B) Determination of genuine Fis-binding peaks on the selected regions. Promoter region of (i) nrfA, (ii) nuoA, (iii) aldB, and (iv) nrdA are occupied by Fis at exponential state under aerobic growth conditions. The peak height of the identified Fis-binding peak is the log2 enrichment ratio calculated from Cy5 (IP DNA) and Cy3 (mock IP DNA) signal intensity of the probe corresponding to the identified peak.
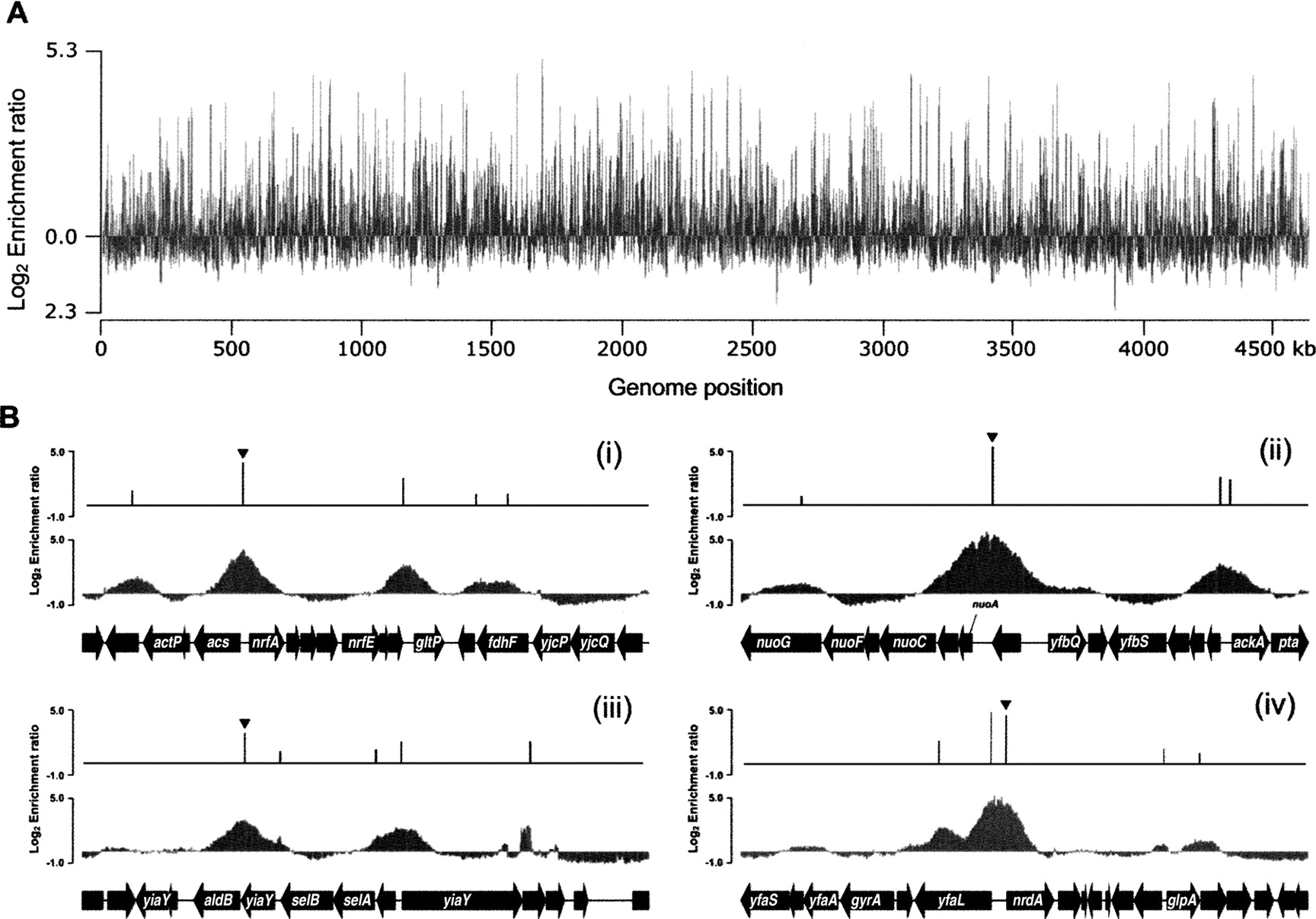
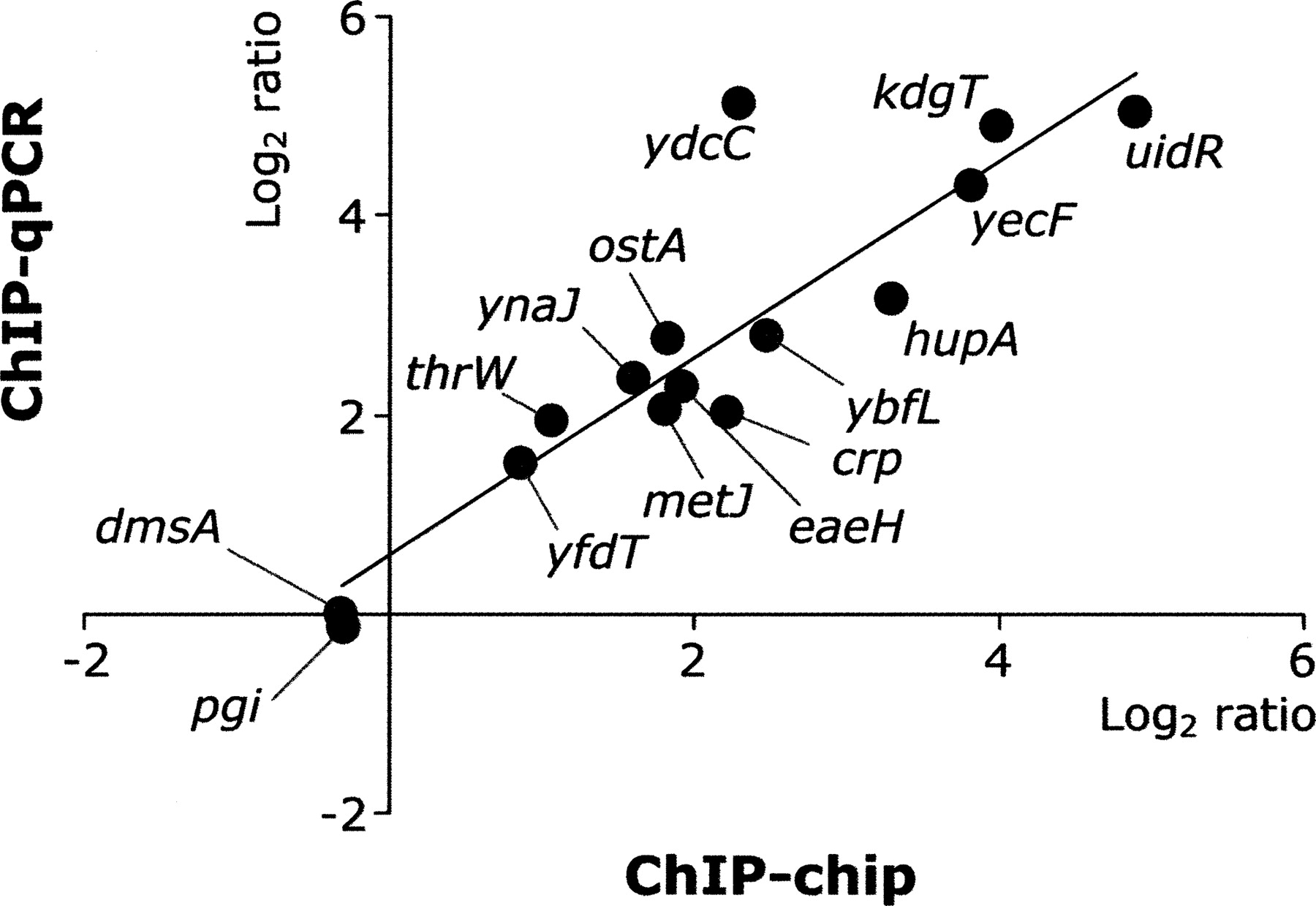
The ChIP-chip analysis of Fis was also in agreement with the literature, showing binding at the promoters of acs, nrfA, nuoA, aldB, and nrd (Fig. 2B) (Augustin et al. 1994; Xu and Johnson 1995; Browning et al. 2004, 2005; Zhang et al. 2004). Prior to this study, only 53 Fis-binding sites had previously been reported, 43 (81%) of which were identified in this study (Supplemental Table S3) (Keseler et al. 2005). The exceptions were lpdA, hupB, lysT-valT-lysW, adhE, osmE, gyrA, rnpB, gyrB, bglGFB, and glnALG. In order to determine whether the failure to detect Fis binding at these 10 sites was due to the sensitivity of the microarrays, we performed conventional ChIP assays followed by qPCR analysis and detected binding of the Fis protein to the promoter region of only bglG. Since these known Fis-binding sites would be considered false negatives of our ChIP-chip analysis (Heintzman et al. 2007), we estimate the sensitivity of our approach to be ∼98% (43 out of 44). Validation of the ChIP-chip results was then done using qPCR on 13 randomly selected sites of the 894 Fis-binding regions (uidR, kdgT, hupA, yecF, eaeH, ybfL, ydcC, crp, thrW, ynaJ, otsA, metJ, and yfdT) and two control regions (pgi and dmsA). All of the selected Fis-binding regions exhibited enrichment as a log2 ratio range of 1.5∼5.1, while the two control regions showed no significant enrichment (Supplemental Table S4). Assuringly, there was a strong correlation between the signal intensities obtained from ChIP-chip analysis and the real-time qPCR (Fig. 3). On the basis of this analysis, we concluded that the majority of Fis-binding peaks identified here are bona fide binding sites.
Verification of ChIP-chip results by real-time quantitative PCR. Thirteen Fis-binding regions were randomly selected from the list of the identified Fis-binding regions. The promoter regions of dmsA and pgi were selected as control regions.
Properties of Fis-binding regions
To assess the properties of Fis-binding regions, we analyzed the position of Fis-binding regions against the current annotated genome information (NC_000913). Fis-binding regions were not only observed within intergenic (IG) regions, but were just as likely to be found within open reading frames (ORFs). From the Fis-binding pattern, we classified three binding categories: IG1, IG2, and ORF. The IG1 category consists of Fis-binding peaks found within promoter regions, while the IG2 consists of Fis-binding peaks found within the intergenic region between convergently transcribed genes (Fig. 4A). All of the remaining sites found within ORFs are thus members of the ORF category. Among a total of 894 unique Fis-binding sites, 547 peaks (∼61%) were within IG1 regions. A significant portion of the Fis-binding sites was also present in the IG2 (48 peaks) and ORF (299 peaks). Thus, although many sites (67%) are present in the intergenic regions (IG1 and IG2), 33% of Fis-binding sites are also located at other regions within a gene (Fig. 4B). To validate these sites shown in IG2 and ORF regions, we performed ChIP analysis followed by qPCR to measure the association of Fis protein with four targets within ORF regions (uidR, ydcC, crp, and otsA) and two targets in the IG2 region (metJ-metB and yfdT-dsdC). The ChIP-qPCR results indicated that each of those regions is a genuine Fis-binding target.
Properties of Fis-binding regions. (A) Classification of Fis-binding regions based on the binding patterns. (i) Fis bound near the promoter of brnQ is a member of the IG1 class, which encompasses Fis bound near promoter regions of the currently annotated genes. (ii) Fis bound between hemN and glnG is a member of the IG2 class, which encompasses Fis bound within the region of two divergently transcribed genes. (iii) Fis bound within the nusA gene is a member of the ORF class, which binds within the open reading frame of annotated genes. (B) Distribution of Fis binding between the three classes. (C) Many of the Fis-binding regions (IG1, IG2, and ORF) are also occupied by RNAP and σ70.
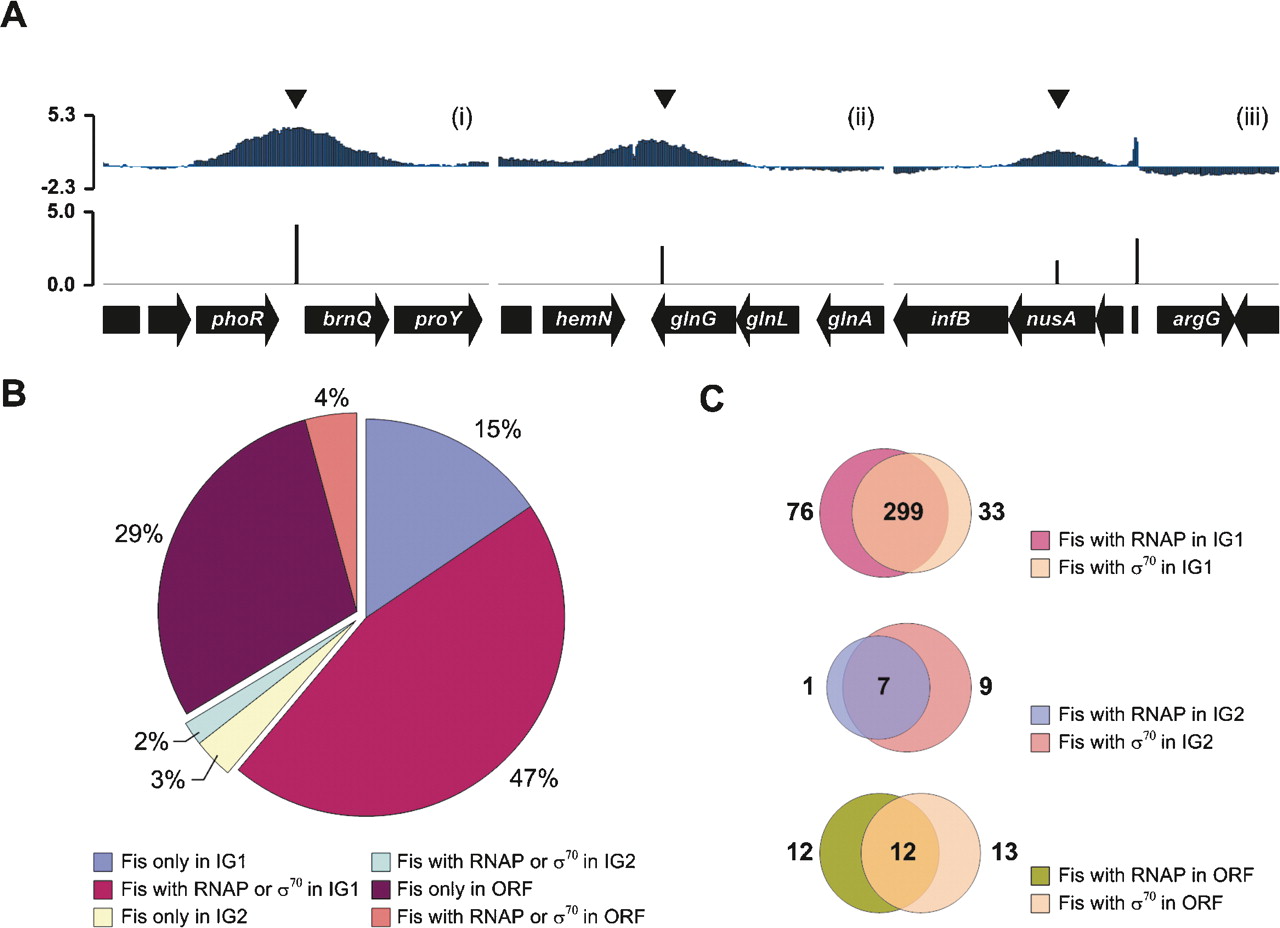
We now compare Fis-binding regions to core RNAP and σ70 binding sites discovered in previous experiments. In a previous study (Herring et al. 2005), we measured the genome-wide association of core RNAP (β′-subunit [also known as RpoC]) using the same microarray under aerobic growth conditions. Since the core RNAP ChIP-chip analysis was performed with rifampicin treatment to trap RNAP at promoter sites, the core RNAP-binding peaks detected represent most of the promoters (both active and inactive). Recently, the genome-wide association of σ70 with the E. coli genome was also revealed by using the similar whole-genome tiling microarray (Reppas et al. 2006). Using all of these data, we found core RNAP or σ70-binding peaks in 462 Fis-binding regions. Most of the core RNAP or σ70-binding peaks were located in the IG1 region (408 peaks). Interestingly, 37 and 17 Fis-binding peaks in the ORF and IG2 regions also have the RNAP or σ70 bindings, respectively (Fig. 4B). Of the 161 σ70 sites that were determined to be within the coding sequences of genes (ORF region) or between convergently transcribed genes (IG2 region) (Reppas et al. 2006), 41 also contained a Fis-binding peak within the same region (Fig. 4C). This result suggests that many of the σ70-dependent transcripts in E. coli are regulated by Fis protein in vivo.
The effect of a fis deletion on changes in the E. coli transcriptome
Given that Fis binds 894 regions of the E. coli genome, we expected the deletion of fis to result in a substantial effect on the global gene expression patterns during exponential growth phase. To address this issue, we isolated total RNA from the parental strain (MG1655) and its isogenic fis deletion mutant during exponential growth phase and hybridized the cDNA obtained from the total RNA onto Affymetrix microarrays. A comparison of the gene expression levels between cells grown in the presence and absence of the Fis protein revealed that 923 genes (21% of current annotated E. coli genes) exhibit differential expression using a 1% FDR (Benjamini and Hochberg 1995) (Supplemental Table S1). In order to determine whether Fis was directly responsible for the differential expression, the correlation between Fis-binding sites and differential gene expression levels was examined (Table 1; Supplemental Table S1). Of the 923 genes that were differentially expressed between the parental strain and fis deletion mutant, only 281 (∼30%) exhibited Fis binding to the region. Of these, 234 were members of the IG1 class, and 47 were members of the ORF class. In regard to the mode of regulation, 84 ORFs (∼9%) were repressed by Fis, and 150 (∼16%) were activated.
Direct and indirect regulation mediated by Fis
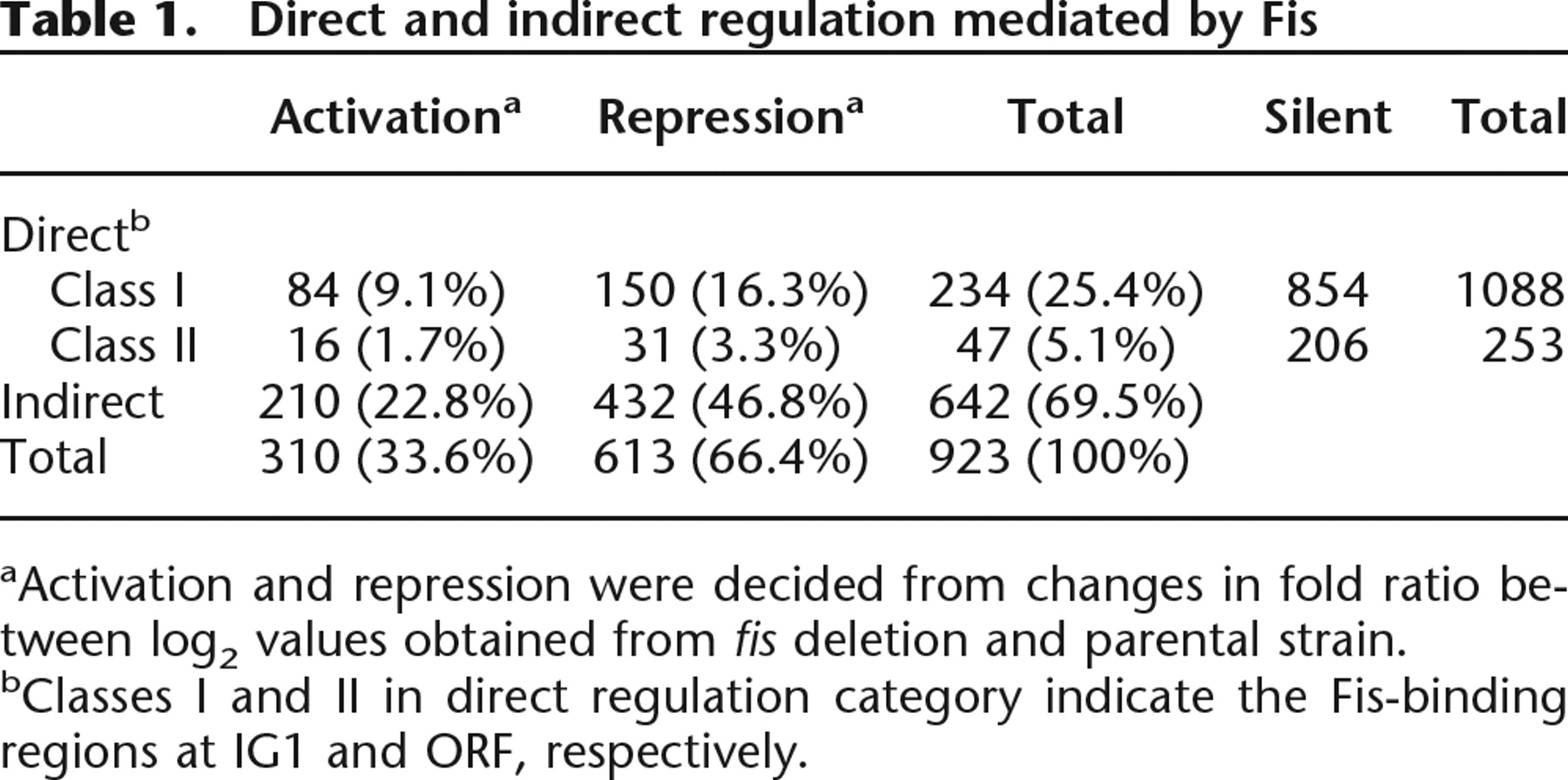
aActivation and repression were decided from changes in fold ratio between log2 values obtained from fis deletion and parental strain.
bClasses I and II in direct regulation category indicate the Fis-binding regions at IG1 and ORF, respectively.
One would expect that the Fis-binding sites within the IG1 class are regulating gene expression through close interaction with the promoter and RNA polymerase, while the Fis within the ORF class are likely to be regulating expression indirectly through local genome architecture. Surprisingly, of the 1341 genes bound by Fis, the expression of only 281 genes was significantly affected when the fis gene was deleted. In addition, 642 (∼70%) genes showing differential expression had no Fis binding and are presumably regulated through an indirect method. This work thus lends support to previous suggestions that the primary role of Fis is in organizing and maintaining nucleoid structure (Schneider et al. 2001), with its direct regulatory role as a secondary function.
Genome-wide mapping of growth-condition-dependent Fis-binding regions
The amount of Fis protein in a cell is known to be growth-phase-dependent (Azam et al. 1999). The dramatic increase in levels of Fis during exponential growth phase is controlled at the transcriptional level, which responds directly to an increase in growth rate. The fact that Fis concentration varies tremendously under different growth phases clearly points to an important regulatory implication of the Fis protein for cell physiology. However, under different growth conditions (e.g., aerobic to anaerobic growth condition shift), its regulatory role or binding regions have not been investigated. To address this issue, genome-wide Fis-binding regions were mapped under aerobic and anaerobic growth conditions in exponential growth phase. Interestingly, the Fis-binding regions identified from the ChIP-chip analysis of anaerobically grown cells were almost identical with those from aerobically grown cells. Complete Fis-binding sites of anaerobic growth conditions are also summarized in Supplemental Table S1. Next, to investigate the effect that Fis has on gene expression, we measured the expression profiles of a fis deletion strain and its parental strain under aerobic and anaerobic conditions. A two-way ANOVA analysis with a 1% FDR (P-value = 0.0001) revealed 48 genes to be regulated by Fis across the aerobic/anaerobic shift. The ChIP-chip data further suggested that 21 of these genes were directly regulated by Fis, while the remaining 27 genes appeared to be regulated indirectly. Of the 21 genes, 19 were members of class I (IG1), and the other two were members of class II (ORF). Interestingly, when comparing the Fis binding of these 21 sites under anaerobic and aerobic conditions, there was no evidence of differential Fis binding between the two conditions. It thus remains unclear whether Fis does, indeed, directly regulate these genes.
Analysis of the length distribution of Fis peak intervals
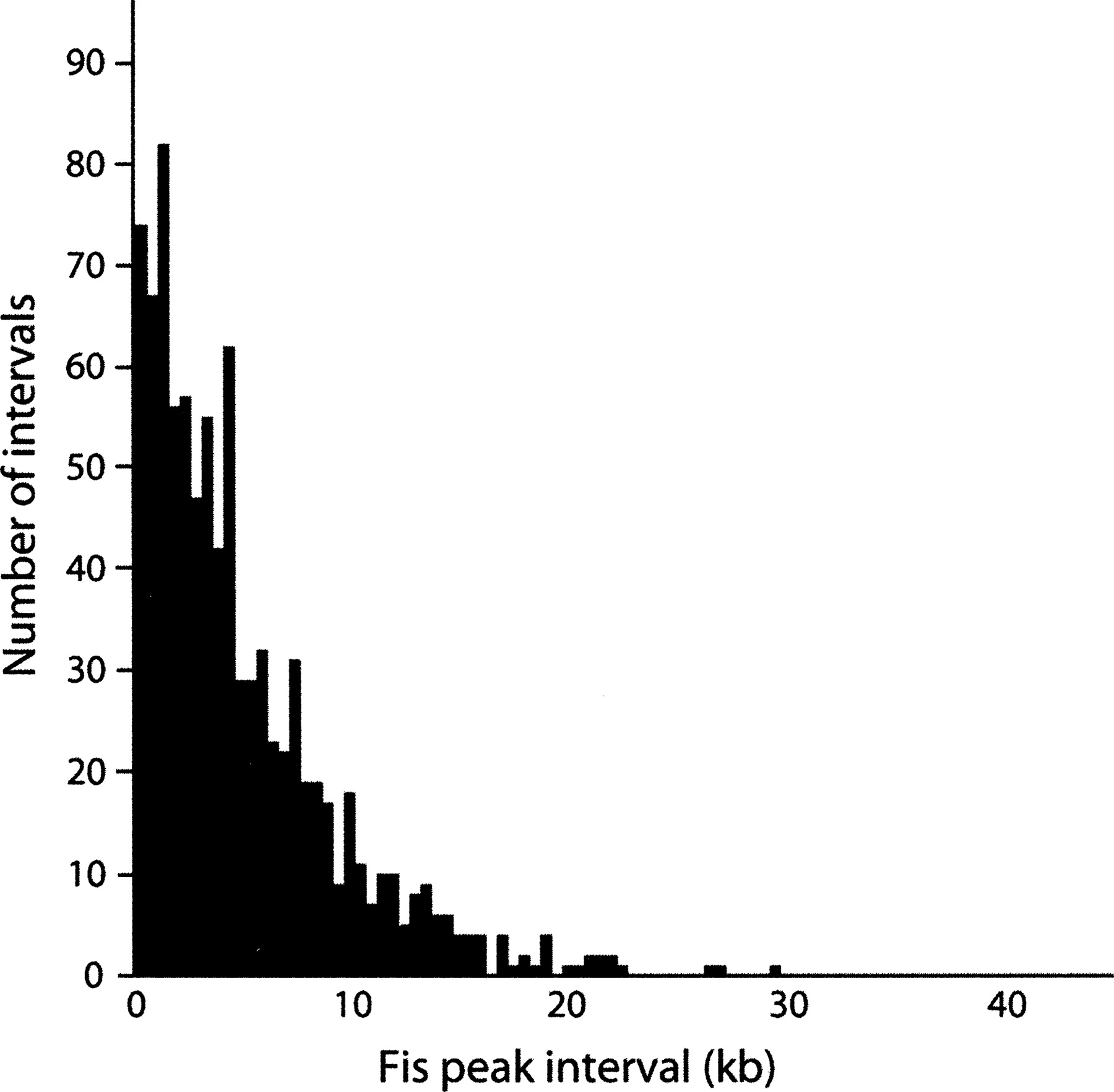
The length distribution and number of supercoiled loop domains in E. coli have been determined via manual measurements of loops seen in electron micrographs, spread-of-supercoiling relaxation experiments (Postow et al. 2004), and resolvase half-lives (Stein et al. 2005). The consensus of these studies is that the average size of ∼400–450 dynamically distributed domains is 10 kb. Since Fis is presumed to be instrumental in defining these domains, we created a histogram of the measured interval sizes between neighboring ChIP-chip Fis peaks. As can be seen in Figure 5, the distribution is similarly exponential in nature. Importantly, the average interval size is 5.15 kb, almost exactly half of the directly measured average domain size (Postow et al. 2004).
A histogram of the lengths of the intervals between Fis-binding sites identified by ChIP-chip experiments.
Determination of the Fis-binding-site position weight matrix (PWM)
We used the large number of Fis-binding regions discovered in this study to reappraise previously estimated Fis-binding site preferences (Finkel and Johnson 1992; Hengen et al. 1997). As a first step in doing this, we manually identified individual binding peaks and then computationally determined the minimal contiguous chromosomal regions corresponding to 70% of the (log ratio) area under each peak. We performed this refinement to minimize the effect of non-bound DNA duplex that is the result of the sonication step in the ChIP-chip protocol. Each such refined chromosomal region was then classified according to the log ratio of its corresponding peak. We then performed motif searches in these chromosomal regions for different log ratio cutoffs. These different log ratio cutoffs corresponded to different levels of conservative searching, with the assumption that chromosomal regions corresponding to Fis peaks with larger log ratio values were more likely to contain more or stronger motif signals. Since Fis binds as a homodimer, we performed two rounds of searches wherein the palindromic motif was and was not mandated.
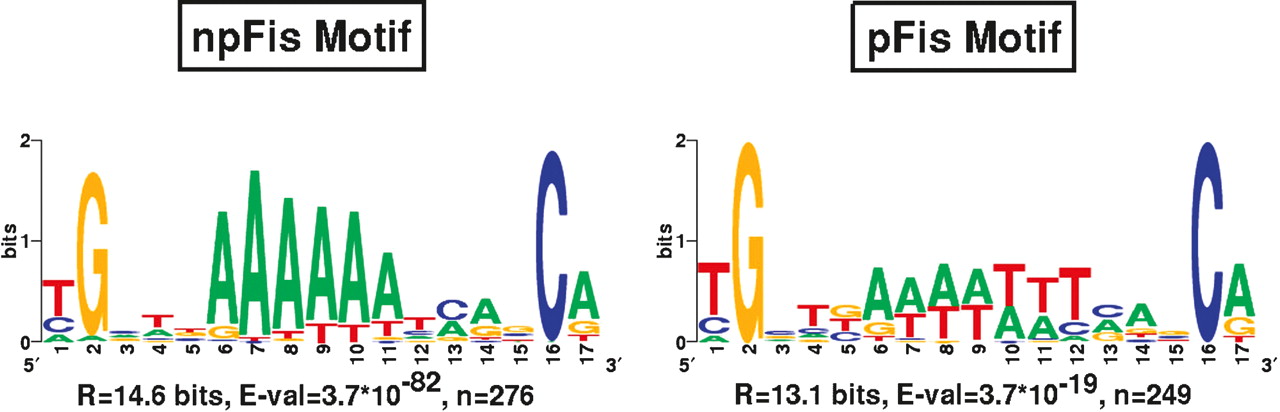
Figure 6 shows the logo representation (Schneider and Stephens 1990) of the sequence found in both the non-palindromic (npFis) and palindromic (pFis) motif searches for the log ratio ≥2 set of sequence. (Supplemental Fig. 3 shows the results for all sequence sets.) Three important results are contained in Figure 6 and Supplemental Figure 3. First, while the npFis motifs found in each of the sequence sets are very significant, the pFis motifs all have much less significant E-values. Second, the information content values of the npFis motifs are larger than the values for the corresponding pFis motifs. Third, both the npFis and pFis motifs contain at their core a strong A-tract and AT-tract, respectively. These results are discussed below in the context of Fis binding and patterning along the chromosome.
The most significant non-palindromic (npFis) and palindromic (pFis) motifs found in the chromosomal sequence regions under the Fis ChIp-chip peaks with log ratios ≥ 2. (Motifs estimated using different conservativeness levels are very similar; see Supplemental Figure 3.) The information content, significance value, and number of sites used to estimate the motif are displayed underneath each motif.
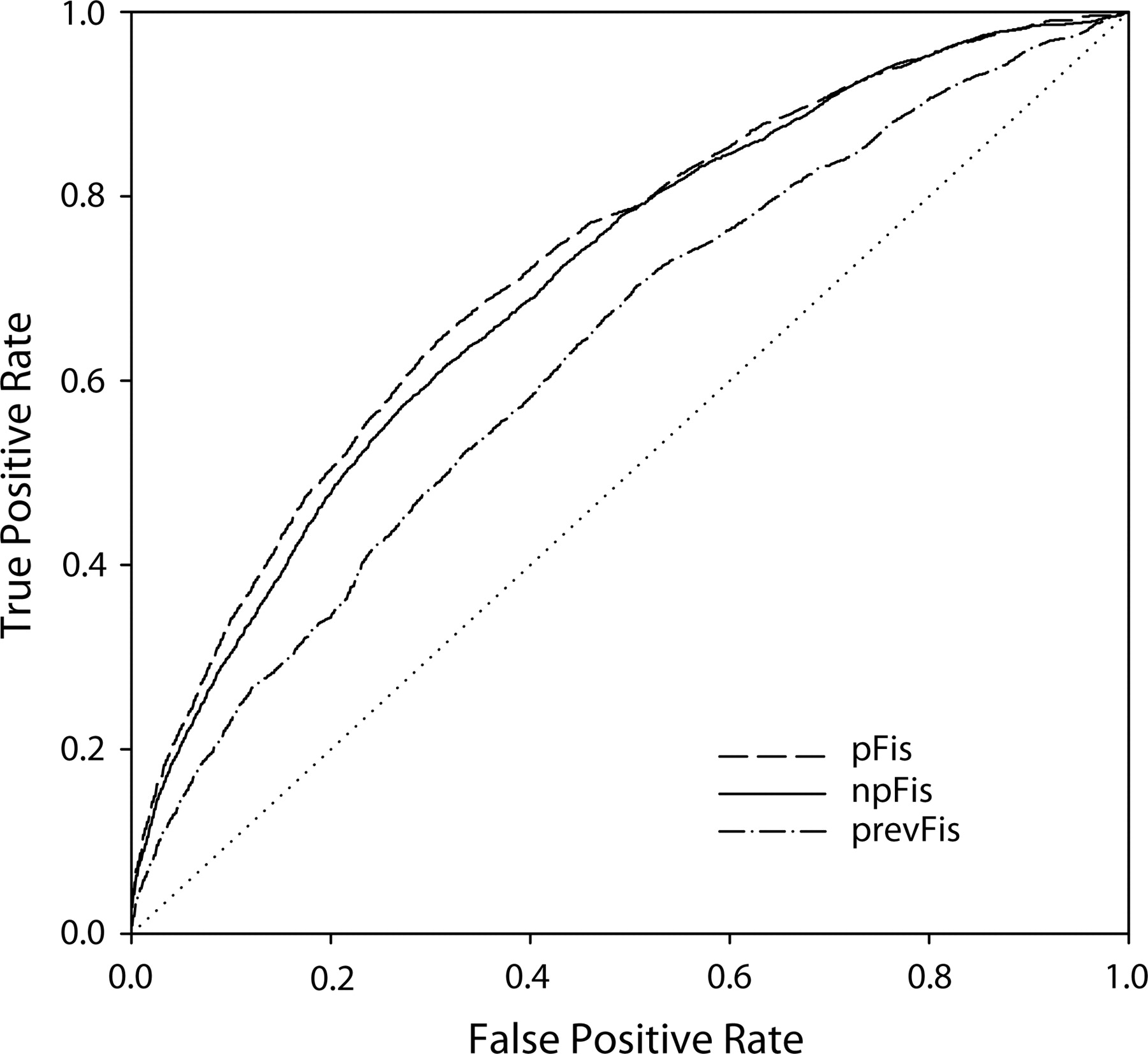
The result shown in Figure 6 presents a conundrum. The Fis protein binds DNA as a homodimer and the most recently estimated (Hengen et al. 1997) Fis motif (prevFis) is palindromic, yet the most informative and significant motif we found was the non-palindromic npFis motif. In order to resolve this conundrum, we performed experiments to determine which of the npFis, pFis, and prevFis motifs better discriminated Fis peak regions from randomly selected chromosomal regions not associated with Fis peaks. We used the motifs resulting from the log ratio ≥2 sets of sequences in Figure 6 to score all of the sequences corresponding to Fis peak regions with log ratio ≥1, and for each sequence assigned it a score based on the largest sum of individual information (Ri) values (Hengen et al. 1997) possible from non-overlapping motif match sites. Figure 7 is an ROC plot displaying the discriminative ability of the three different motifs, and contains two important results. First is that both of the npFis and pFis motifs derived in this work are better discriminators of chromosomal Fis-peak regions from non-Fis-peak regions than is the prevFis motif. Secondly, while the npFis and pFis motifs are basically very comparable in their discriminative ability, the pFis motif seems to have slightly better discriminative ability. This was not an expected result given their relative information content and significance values.
Receiver Operator Characteristic plots evaluating how well the npFis and pFis motifs from Figure 6 and the previously established Fis motif (Hengen et al. 1997) discriminated all (log ratio ≥ 1) Fis-peak-associated chromosomal sequences from random chromosomal sequences. The plotted curves are the average of 20 discrimination experiments that used different random chromosomal sequence sets.
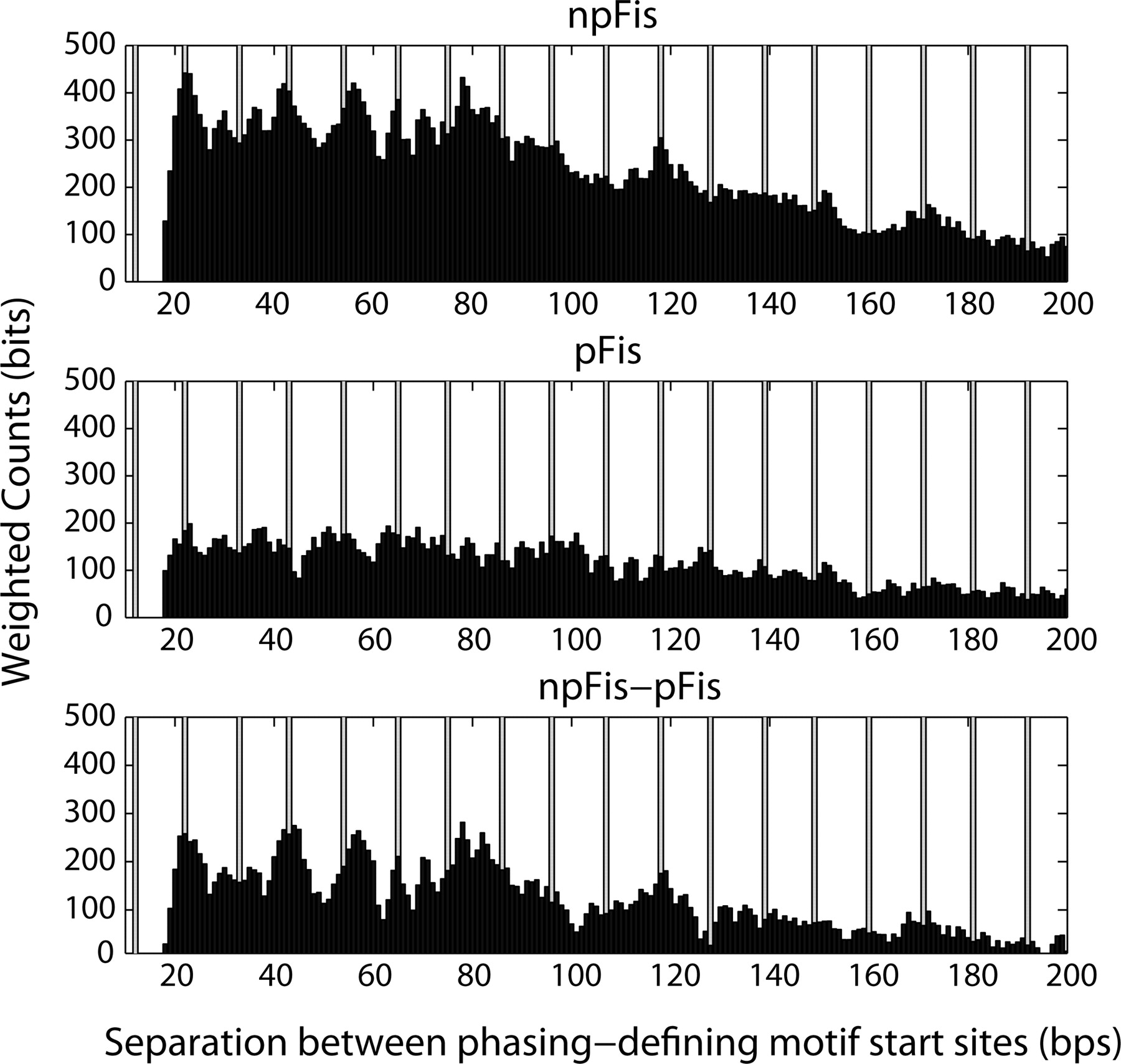
To better understand the relationship between the npFis and pFis motifs, we first identified the phasing-defining Fis (npFis or pFis) sites in the set of Fis peak region sequences. Both members of a pair of sites were considered phasing-defining sites if all intervening sites between the pair had lower Ri values. For each phasing-defining pair, we computed the separation distance between their start positions. We then created a histogram of the separation distances associated with npFis motifs and a histogram of the separation distances associated with pFis, and weighted each distance value by the Ri values of the site defining the separation distance. Since Ri values have been correlated with binding affinity for Fis (Shultzaberger et al. 2007), we interpret higher such weightings to be indicative of more physiologically likely Fis-binding configurations. Figure 8 (top and middle) shows these weighted histograms for the npFis and pFis motifs, and Figure 8 (bottom) is the subtractive difference of the pFis histogram from the npFis histogram. Figure 8 (bottom) shows the competitive difference of the two motifs in dictating Fis phasing in regions containing multiple potential Fis-binding sites. The pattern in the difference histogram of Figure 8 (bottom) shows the increased propensity for npFis to dictate helical or antihelical phasing of Fis molecules.
Histogram of the separation distances between match start sites in the Fis peak regions for the npFis (top) and pFis (middle) motifs. Distances are weighted by the motif match score (Ri value) for each instance when the motif defines a separation distance. The vertical bars indicate the motif separation distances that place the A-tracts in the core of the npFis and pFis motifs (beginning at position 6 in Fig. 6) in perfect helical register (assuming 10.6 bp/helical turn in B-DNA). (Bottom panel) The subtractive difference of the top and middle histograms.
Discussion
We have mapped genome-wide distribution of E. coli nucleoid associated protein Fis in exponentially growing cells using a high-resolution whole-genome tiling microarray. In addition, expression profiles of a wild type and a fis deletion mutant were generated to determine the effect Fis has on transcription. By integrating these two data sets, we were able to show that: (1) 894 Fis-binding sites were identified, ∼67% of which were located within non-coding regions, while the remaining ∼33% were found within coding regions; (2) Fis binding to the E. coli genome was insensitive to aerobicity; (3) expression profiles determined 1341 genes to be weakly affected by Fis, with only 30% containing Fis bound within the region; and (4) half of Fis-binding sites overlap with the binding regions of both RNA polymerase and σ70. In addition, computational analyses revealed that: (1) Fis-binding signal in the chromosome was found to be necessary but not sufficient to explain the preferred binding locations by Fis as revealed by ChIP-chip. (2) The average interval size between Fis-binding sites was 5 kb, which is half the average supercoiling domain size. Furthermore, the number of Fis peaks was almost double the estimated number of supercoiling domains, suggesting a stoichiometric relationship of two Fis-binding regions per supercoiling domain. (3) By utilizing a large number of the Fis-binding regions, a Fis-binding motif was generated and compared to the previously established binding motif.
Genome-wide distribution of E. coli nucleoid-associated protein Fis shows that Fis specifically binds ∼894 regions throughout the E. coli chromosome. The binding sites included 43 previously described regulatory targets and many novel-binding targets that have not been identified. Of the 894 binding regions identified, ∼67% were located within non-coding regions, while the remaining ∼33% were found within coding regions. The experiments were then repeated under anaerobic conditions, and it was found that oxygen had no detectable effect on the binding of Fis. The unusually high number of Fis-binding sites was quite surprising, given that no transcription factors in E. coli bind more than ∼200 sites (Martinez-Antonio and Collado-Vides 2003). A previous study on the genome-wide mapping of Fis binding identified only 224 target sites, with half of them found within non-coding regions and the other half within coding regions (Grainger et al. 2006). Differences between this study and the previous one (Grainger et al. 2006) may be due to the low-resolution array used in the previous study, since microarray resolution is a critical factor when performing ChIP-chip experiments. For example, the previous ChIP-chip study detected no Fis binding within the rRNA operon region, which is clearly activated by Fis bindings (Paul et al. 2004); however, the ChIP-chip result in this study shows genuine binding peaks on all of seven rRNA operons (Supplemental Fig. S1). These discrepancies are most likely due to the higher-resolution arrays’ increased ability to discern the actual binding from noise. When using high-resolution arrays for ChIP-chip, binding peaks appear as a normal Gaussian distribution, which are clearly illuminated when using a tiled array (Fig. 2); however, as the array resolution is decreased, so is the resolution of the peaks, thus making it difficult to discern between noise and the true signals.
The general concept of binding patterns of global transcription factors is that their target sites are located at promoter regions. Through interacting with RNA polymerase and/or other proteins, and/or hindering the binding of RNA polymerase at the promoter, it becomes able to activate or repress the transcription of the target genes. Our genome-wide analysis indicates that Fis binds numerous such regions (67%). On the other hand, our analysis suggests that the general concept for a global transcription factor in regulation may be partially incorrect for Fis, since Fis-binding regions were also found at the range of many different sites (33%) such as within ORF regions (Grainger et al. 2006). Note that only a certain proportion (30%) of bound Fis directly affects transcription. Thus, Fis should be considered as a genome-organizing protein like Crp, in addition to its function as a promoter-specific regulator (Grainger et al. 2005). The Fis protein showed the ability to bend DNA, indicating that the bending activity stabilizes DNA looping to enhance transcription as well as to promote DNA compaction (Travers and Muskhelishvili 1998; Skoko et al. 2006). The Fis binding within ORF regions may reflect the DNA bending activity to maintain chromosome structure and transcription regulation as well.
Genome-wide mapping of Fis-binding sites was then compared with expression profiles of a fis deletion mutant and its parental strain to determine the effect that Fis has on the transcription. The expression profiles determined 1341 genes to be affected by Fis, yet only 30% had Fis bound within the region. It is worthwhile to note that with 894 Fis-binding sites and the expression of 1341 affected by the removal of Fis, there inevitably will be some coincidental overlap, rendering it difficult to infer direct regulation by ChIP-chip and gene expression data alone. However, these experiments do put an upper limit on the number of promoters directly regulated by Fis, which is approximately 424. A surprising result from the expression profiling was the extremely small change a fis deletion has on expression. Although the expression of many genes was significantly affected by the deletion of fis, the median change in expression of those genes was only ∼0.37 log2 ratio. For comparison, the median change in expression when the global regulators fnr and arcA are deleted is ∼0.88 and ∼0.89 log2 ratio, respectively (Covert et al. 2004). The small effect that Fis seems to have on transcriptional expression could explain the minimal growth rate difference between the wild-type strain and a fis deletion mutant, during log-phase growth (Zhi et al. 2003). Recently, using high-resolution atomic force microscopy (AFM), a ternary complex of Fis, RNAP, and σ70 was visualized at tyrT promoter (Maurer et al. 2006). Visualization of the ternary complex showed that Fis forms a discrete assembly by positioning in close proximity to an RNAP molecule. Owing to the fact that there was weak interaction between Fis and the RNAP, that result may explain the weak regulation observed in this study.
When compared with ChIP-chip data of RNA polymerase and σ70, it was found that half of the Fis-binding sites overlap with the binding regions of both RNA polymerase and σ70. Interestingly, 54 Fis-binding sites within coding regions and intergenic regions between convergently transcribed genes were also occupied by RNAP and σ70 (Fig. 4; Supplemental Table 1). A recent study on the E. coli transcriptome using high-density tiling microarrays has also suggested the existence of many novel transcripts within the gene coding region (Reppas et al. 2006). This observation could also be found in the ChIP-chip analysis of σ70 and σ32, indicating that a significant portion of the binding sites of σ70 and σ32 are not associated with the 5′-ends of current annotated genes (Wade et al. 2006). Therefore, the Fis-binding sites within gene region may be regulatory cis-elements of Fis for modulating transcription of the currently unknown transcripts in the E. coli genome. As another view of this issue, we speculate that Fis regulates the transcription by the formation of DNA microloops, which form a separate topological domain (Postow et al. 2004). In those regions, the RNAP may be trapped to repress the transcription or may recycle to efficiently activate the gene transcription process.
Computational analyses in this work resulted in a refinement to the Fis DNA binding signal and subsequently to new insights into the functional behavior of the Fis protein. Fis has a previously documented (Skoko et al. 2006) dual behavior, which is that while it can bind nonspecifically to completely coat long stretches of duplex DNA, it also has preferred binding sites to which it binds and sets the phasing of the stretches of nonspecifically bound Fis. The two npFis and pFis motifs we identify in Figure 6 are quite similar when one realizes that their core signals are an A-tract and an AT-tract, respectively, and that A-tracts and AT-tracts >4 nt have very similar DNA bending characteristics (Hagerman 1990; Hud et al. 1998; Hud and Plavec 2003; Stefl et al. 2004). There are differences, though, for while the less significant and less informative pFis motif contains a more generic and palindromic AT-tract (reminiscent of the previously estimated Fis motif) (Hengen et al. 1997), the more highly significant and more informative npFis motif contains an A6-tract. Because selectivity of Fis binding is thought to reflect the intrinsic bent nature of particular DNA sequences (Betermier et al. 1994) and because A6-tracts induce the largest intrinsic curvature to segments of DNA (Koo et al. 1986), our results imply that the npFis motif represents the highest-affinity DNA sequence signal for Fis. DNA segments that more resemble the pFis motif, then, would be lower-affinity sites (that are still preferred over random DNA). While this preferential hierarchy is likely modulated by the influence of flanking nucleotides on binding affinity (Pan et al. 1996; Perkins-Balding et al. 1997), the 15-bp core is enough to specify high-affinity binding sites (Bruist et al. 1987). The result that A-/AT-tracts constitute a critical component of high-affinity Fis-binding sites is supported by numerous previous experiments. For instance, 39 of the 60 confirmed Fis binding sites used to construct the prevFis motif (Hengen et al. 1997) contain A-/AT-tract cores, and many of the known high-affinity Fis binding sites contain A-tract cores (Pan et al. 1996).
The identification of a preferred DNA sequence signal for Fis binding (npFis) and the dominating helical and anti-helical phasing signal of the npFis motif over the pFis motif (Fig. 8, bottom) together have important implications in supercoiled DNA. Fis bends DNA when it binds (Thompson et al. 1988), and helically phased Fis binding induces and stabilizes curved DNA (Hubner et al. 1989; Lazarus and Travers 1993; Muskhelishvili et al. 1995; Perkins-Balding et al. 1997). Curved segments of supercoiled DNA are most thermodynamically favorably located at apices of plectonemes, which aside from uniquely orienting a supercoiling domain (Laundon and Griffith 1988) greatly enhance a local region’s exposure to transcription machinery (ten Heggeler-Bordier et al. 1992; Lazarus and Travers 1993; Rochman et al. 2002; Muskhelishvili and Travers 2003). Fis-bound stretches of DNA that are not curved overall—which would be ensured by high-affinity Fis binding sites that are not helically phased—would not have a propensity to occur at apices, but would be associated with duplex crossovers and branch points (Schneider et al. 2001). This inferred mechanism for structuring supercoiled DNA complements the observed Fis peak interval distribution (Fig. 5). We interpret the discoveries that the Fis peak interval distribution and previously inferred supercoiling domain size distribution were both exponentially distributed, that the average Fis peak interval (5 kb) was half of the average domain size (10 kb), and that the number of Fis peaks (894) was almost double the estimated number of supercoiling domains (450) to be strong evidence for an average of two Fis-binding regions per supercoiling domain. The roles that these regions would play in structuring supercoiling domains through the stabilization of crossovers, loops, bends, or apices would be largely influenced by the phasing of those DNA sequences that most resemble the high-affinity binding site motif npFis.
In a broader context, our results imply that A-tracts flanked by appropriately positioned C/G residues are preferred Fis-binding sites, and, in particular, A6-tracts provide the strongest Fis-binding signal. The E. coli chromosome contains an over-representation of (83,358) A-/AT-tracts that demonstrate a 10–12-bp periodicity and are grouped in clusters (Tolstorukov et al. 2005) in roughly 150-bp regions. As discussed in previous work (Laundon and Griffith 1988; Rippe et al. 1995), such A-/AT-tract clusters would have a higher propensity to be intrinsically curved and thus to induce branches in superhelical plectonemes and to position promoters at the apices of superhelices. These are the same topological roles ascribed to the Fis protein. Our results, then, support the supposition (Tolstorukov et al. 2005) that A-/AT-tracts constitute a sequence-directed structuring code for the E. coli chromosome by in part serving as binding sites for the nucleoid-associated protein Fis. In summary, our genome-wide approach using ChIP-chip analysis not only provides a comprehensive assessment of the genomic distribution of the bound Fis and its role in transcription regulation, but also suggests directions for furthering our understanding of the structure, function, and evolution of the E. coli nucleoid.
Methods
Bacterial strains and growth conditions
E. coli strain MG1655 was used to generate the deletion mutant and the BOP608 strain harboring Fis-8myc (Cho et al. 2006a). Deletion mutant (MG1655 Δfis) was constructed by a λ Red and FLP-mediated site-specific recombination system (Datsenko and Wanner 2000). Glycerol stocks of E. coli strains were inoculated into M9 minimal medium containing 2 g/L glucose as a carbon source and cultured overnight at 37°C with constant agitation. The cultures were inoculated into 100 mL of fresh M9 medium containing 2 g/L glucose and cultured at 37°C with constant agitation to an appropriate cell density (Covert et al. 2004). In the case of anaerobic cultures, after the medium (250 mL) was flushed with a nitrogen/carbon dioxide (9:1) mixture gas for 30 min to assure anaerobic conditions, the strains were grown at 37°C with continuous sparging with the gas mixture, and agitation in the minimal medium (Cho et al. 2006b).
Chromatin immunoprecipitation (ChIP)
E. coli strain BOP608 was used to perform all ChIP-chip experiments. BOP608 cultures at mid-log growth phase aerobically (OD A600 ≈ 0.6) or anaerobically (OD A600 ≈ 0.2) were cross-linked by 1% formaldehyde (37% solution; Fisher Scientific) at room temperature for 25 min. Following quenching the unused formaldehyde with 125 mM glycine for an additional 5 min of incubation at room temperature, the cross-linked cells were harvested and washed three times with 50 mL of ice-cold TBS. The washed cells were resuspended in 0.5 mL of lysis buffer composed of 50 mM Tris-HCl (pH 7.5), 100 mM NaCl, 1 mM EDTA, protease inhibitor cocktail (Sigma), and 1 kU of Ready-Lyse lysozyme (Epicentre). The cells were incubated for 30 min at 37°C and then treated with 0.5 mL of 2× IP buffer composed of 100 mM Tris-HCl (pH 7.5), 200 mM NaCl, 1 mM EDTA, and 2% (v/v) Triton X-100. The lysate was then sonicated four times for 20 sec each in an ice bath to fragment the chromatin complexes using Misonix Sonicator 3000 (output level = 2.5). The range of the DNA size resulting from the sonication procedure was 300–1000 bp, and the average DNA size was 500 bp. Cell debris was removed by centrifugation at 37,000g for 10 min at 4°C, and the resulting supernatant was used as cell extract for the immunoprecipitation. To immunoprecipitate the Fis–DNA, σ70–DNA, or RNAP–DNA complexes, 3 μg of anti-c-myc antibody (9E10; Santa Cruz Biotech), 6 μL of anti-σ70 antibody (2G10; Neoclone) or 6 μL of anti-RNAP β subunit antibody (NT63; Neoclone) were then added into the cell extract, respectively. For the control (mock-IP), 2 μg of normal mouse IgG (Upstate) was added into the cell extract. They were then incubated overnight at 4°C, and 50 μL of the Dynabeads Pan Mouse IgG (for c-myc) or protein A (for σ70 and RNAP β subunit) magnetic beads (Invitrogen) was added into the mixture. After 5 h of incubation at 4°C, the beads were washed twice with the IP buffer (50 mM Tris-HCl at pH 7.5, 140 mM NaCl, 1 mM EDTA, and 1% [v/v] Triton X-100), once with the wash buffer I (50 mM Tris-HCl at pH 7.5, 500 mM NaCl, 1% [v/v] Triton X-100, and 1 mM EDTA), once with wash buffer II (10 mM Tris-HCl buffer at pH 8.0, 250 mM LiCl, 1% [v/v] Triton X-100, and 1 mM EDTA), and once with TE buffer (10 mM Tris-HCl at pH 8.0, 1 mM EDTA) in order. After removing the TE buffer, the beads were resuspended in 200 μL of elution buffer (50 mM Tris-HCl at pH 8.0, 10 mM EDTA, and 1% SDS) and incubated overnight at 65°C for reverse cross-linking. After reversal of the cross-links, RNAs were removed by incubation with 200 μL of TE buffer with 1 μL of RNaseA (QIAGEN) for 2 h at 37°C. Proteins in the DNA sample were then removed by incubation with 4 μL of proteinase K solution (Invitrogen) for 2 h at 55°C. The sample was then purified with a PCR purification kit (QIAGEN). Prior to the microarray experiments, the gene-specific quantitative PCR was carried out using the DNA samples.
Real-time qPCR
To measure the enrichment of the Fis-binding targets in the DNA samples, 1 μL of IP or mock-IP DNA was used to carry out gene-specific real-time qPCR with the specific primers to the promoter regions (primer sequences are available upon request). The real-time qPCR conditions were as follows: 25 μL SYBR mix (QIAGEN), 1 μL of each primer (10 pM), 1 μL of IP or mock-IP DNA, and 22 μL of ddH2O. All real-time qPCR reactions were done in triplicate. The samples were cycled for 15 sec to 94°C, for 30 sec to 52°C, and for 30 sec to 72°C (total 40 cycles) in iCycler (Bio-Rad). Three independent biological replicates were prepared and subject to be analyzed by three independent technical replicates for the real-time qPCR.
Amplification of DNA
To amplify the DNA samples, 7 μL of the IP or mock-IP DNA, 2 μL of 5× Sequenase buffer, and 1 μL of 40 μM Rand 9-Ns primer (5′-TGGAAATCCGAGTGAGTNNNNNNNNN) were mixed in a PCR tube. The mixture was heated for 2 min to 94°C and then cooled to 10°C in a PCR machine (Bio-Rad). One microliter of 5× Sequenase buffer, 1.5 μL of dNTP mix (2.5 mM each), 1.5 μL of BSA (0.5 mg/mL), 0.75 μL of DTT (0.1 M), and 0.3 μL of Sequenase (13 U/μL) were added to the mixture. The mixture was ramped from 10°C to 37°C over 8 min, held for 8 min at 37°C, heated for 2 min to 94°C, and then cooled to 10°C. 0.9 μL of Sequenase dilution buffer and 0.3 μL of Sequenase (13 U/μL) were added to the samples and ramped from 10°C to 37°C over 8 min, held for 8 min at 37°C, and then cooled to 4°C. The samples were diluted by addition of 45 μL of ddH2O. A reaction mixture (100 μL) of 15 μL of the diluted DNA, 10 μL of 10× pfu reaction buffer, 10 μL of dNTP mix (2.5 mM each), 1 μL of 100 μM Rand-univ primer (5′-TGGAAATCCGAGTGAGT), 1 μL of pfu polymerase (5 U/μL), and 63 μL of ddH2O was prepared on ice. Four tubes per sample were prepared to achieve enough DNA quantity for microarray hybridization. The samples were cycled for 30 sec to 94°C, for 30 sec to 40°C, for 30 sec to 50°C, and for 2 min to 72°C (total 25 cycles). The amplified samples were then purified by using a PCR purification kit (QIAGEN). The amplified DNA samples were then ethanol-precipitated and dissolved in 9 μL (IP DNA) and 7 μL (mock-IP DNA) of ddH2O, respectively. DNA yields ranged from 5 to ∼10 μg, and A260/280 was between 1.8 and 2.0. The enrichment of the Fis-binding targets in the amplified DNA samples was measured using gene-specific real-time qPCR.
Whole-genome-tiled microarray analysis
We used a custom-tiled NimbleGen microarray for the ChIP-chip assay. The microarray includes all the E. coli MG1655 genome sequence spaced on average 25 bp apart, resulting in 371,034 oligonucleotide probes that randomly distributed on the array. Detailed methods used for microarray process are described in Supplemental Methods.
Transcriptional analysis
Affymetrix E. coli Antisense Genome Arrays were used for all transcriptional analyses. Cultures were grown to mid-exponential growth phase aerobically (OD A600 ≈ 0.6) or anaerobically (OD A600 ≈ 0.2). Cultures (3 mL for aerobic and 9 mL for anaerobic) were added to 2 volumes of RNAprotect Bacteria Reagent (QIAGEN), and total RNA was then isolated using RNeasy columns (QIAGEN) with DNase I treatment. Total RNA yields were measured using a spectrophotometer (A260), and quality was checked by visualization on agarose gels and by measuring the sample A260/A280 ratio (>1.8). cDNA synthesis, fragmentation, end-terminus biotin labeling, and array hybridization were performed as recommended by Affymetrix standard protocol. Raw CEL files were analyzed using a robust multi-array average for normalization and calculation of probe intensities. The processed probe signals derived from each microarray were averaged for both the wild-type and fis deletion mutant strains. To assess statistically significant differential expression, the probe signals were tested using pairwise t-test comparisons between wild-type and fis deletion mutant strains. Genes meeting a 1% FDR (false discovery rate)-adjusted P-value cutoff (0.0001) were chosen as significant changes in gene expression. The filtered genes were then ascribed to genes directly or indirectly affected by Fis protein.
Refinement of Fis peak chromosomal regions
After manually defining Fis peaks, we wrote a greedy algorithm to identify the chromosomal sequence region associated with 70% of the (log ratio) area under each peak. The algorithm worked by first identifying the three consecutive probes whose associated peak area was greatest, and then expanding the consecutive set of probes in either the 5′ or 3′ direction depending on which neighboring probe had a higher value. This process ceased when 70% of the peak area had been accumulated in a set of consecutive probes. The chromosomal start position of the first probe and the chromosomal end position of the last probe were used to define the “refined chromosomal peak region.”
Motif searching
To find the Fis-binding site position weight matrix (PWM), we first constructed sets of refined chromosomal peak region sequences reflecting different levels of conservativeness. The most conservative set consisted of sequences for only Fis peaks with associated log ratios ≥ 4. Less conservative sets were constructed for log ratios of 3, 2, and 1. The rationale for such sets was that sequences associated with high log ratios were more likely to contain more and/or stronger Fis-binding DNA sequences. We then used Meme (Bailey and Elkan 1994) to search for the most significant motif in each set of sequences. Since Fis binds as a dimer and since the previously estimated Fis motif (Hengen et al. 1997) is palindromic, we also searched for the most significant palindromic motif in each set of sequences (accomplished by using the “−pal” option to Meme). In all searches, the reverse complement of each sequence was allowed to contain sites. Supplemental Figure 3 shows the results of the motif searches.
Motif discrimination ability
We tested the ability of the npFis, pFis, and prevFis motifs to discriminate Fis peak sequences from non-Fis-peak chromosomal sequences by first constructing 20 sets of randomly selected chromosomal sequences. Each such set contained the same number of sequences with the same length distribution as the log ratio ≥ 1 set of refined Fis peak sequences. In scoring a single DNA sequence, we used the position weight matrix (PWM) for the appropriate motif to identify all sites in the sequence (including its reverse complement) with an individual information (Ri) > 0.0 bits (Schneider 1997). Using dynamic programming, we then computed the set of non-overlapping sites with the greatest sum of Ri values. The score for a sequence was defined as this sum of Ri values. A discrimination experiment, then, consisted of scoring the log ratio ≥ 1 set of refined Fis peak sequences and a set of randomly selected chromosomal sequences and creating a receiver operating characteristic (ROC) plot from the combined results. We performed 20 such discrimination experiments for each motif using the 20 sets of random chromosomal sequences and reported the average ROC plot in Figure 7.
Sequence positioning relationship between npFis and pFis motifs
To understand how the npFis and pFis sequence signals interact in Fis peak regions, we scored the log ratio ≥1 set of refined Fis peak sequences with both of the npFis and pFis PWMs and computed the set of nonoverlapping sites with the greatest sum of Ri values—irrespective of the identity (npFis or pFis) of each associated site. In this way, each sequence had an optimal patterning of npFis and pFis sites. Any pair of these sites was associated with a distance between their respective start sites, defined by the number of intervening nucleotide positions. In each sequence, we identified all pairs of sites such that for each pair composed of site1 with Ri = R1 and site2 with Ri = R2, any sitej between site1 and site2 had Rj < R1 and Rj < R2. Since the individual information Ri of a site has been shown to be correlated to the binding energy of Fis (Shultzaberger et al. 2007), we reasoned that the higher Ri sites would be more strongly bound by Fis protein and would dictate the positioning of any intervening bound Fis molecules. Both site1 and site2 can be npFis or pFis motifs. To quantify how the npFis and pFis motifs contribute to Fis positioning, and thus to different distances between all pairs of sites site1 and site2, we created a separate distance histogram for both npFis motif sites (Fig. 8, top) and pFis motif sites (Fig. 8, middle)—using as a distance “count” the Ri value of a site. Thus for each pair of sites site1 and site2 (with R1 and R2, respectively) separated by d nucleotides, a “weighted” count R1 for distance d was added to the histogram for either npFis or pFis, and similarly for R2. To assess how the npFis and pFis motifs differently contribute to different motif site separation distances, we subtracted the pFis distance histogram from the npFis distance histogram (see Fig. 8, bottom). All distance histograms were smoothened using an averaging window of 3 bp.
Raw ChIP-chip data
The data file for all raw data can be downloaded from the following web site: http://systemsbiology.ucsd.edu/publications/.
Acknowledgments
We thank Mark Abrams for insightful discussions regarding manuscript writing. This work was supported by National Institutes of Health Grant GM062791.
References
- ↵L.B. Augustin,B.A. Jacobson,J.A. Fuchs,(1994) Escherichia coli Fis and DnaA proteins bind specifically to the nrd promoter region and affect expression of an nrd-lac fusion. J. Bacteriol. 176:378–387.
- ↵T.A. Azam,A. Iwata,A. Nishimura,S. Ueda,A. Ishihama,(1999) Growth phase-dependent variation in protein composition of the Escherichia coli nucleoid. J. Bacteriol. 181:6361–6370.
- ↵T.A. Azam,S. Hiraga,A. Ishihama,(2000) Two types of localization of the DNA-binding proteins within the Escherichia coli nucleoid. Genes Cells 5:613–626.
- ↵T.L. Bailey,C. Elkan,(1994) Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2:28–36.
- ↵C.A. Ball,R. Osuna,K.C. Ferguson,R.C. Johnson,(1992) Dramatic changes in Fis levels upon nutrient upshift in Escherichia coli. J. Bacteriol. 174:8043–8056.
- ↵S.M. Bearson,J.A. Albrecht,R.P. Gunsalus,(2002) Oxygen and nitrate-dependent regulation of dmsABC operon expression in Escherichia coli: Sites for Fnr and NarL protein interactions. BMC Microbiol. 2:13, 10.1186/1471-2180-2-13.
- ↵Y. Benjamini,Y. Hochberg,(1995) Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. [Ser. B] 57:289–300.
- ↵M. Betermier,D.J. Galas,M. Chandler,(1994) Interaction of Fis protein with DNA: Bending and specificity of binding. Biochimie 76:958–967.
- ↵N. Blot,R. Mavathur,M. Geertz,A. Travers,G. Muskhelishvili,(2006) Homeostatic regulation of supercoiling sensitivity coordinates transcription of the bacterial genome. EMBO Rep. 7:710–715.
- ↵A.J. Bokal,W. Ross,T. Gaal,R.C. Johnson,R.L. Gourse,(1997) Molecular anatomy of a transcription activation patch: FIS-RNA polymerase interactions at the Escherichia colirrnB P1 promoter. EMBO J. 16:154–162.
- ↵D.F. Browning,C.M. Beatty,A.J. Wolfe,J.A. Cole,S.J. Busby,(2002) Independent regulation of the divergent Escherichia coli nrfA and acsP1 promoters by a nucleoprotein assembly at a shared regulatory region. Mol. Microbiol. 43:687–701.
- ↵D.F. Browning,C.M. Beatty,E.A. Sanstad,K.E. Gunn,S.J. Busby,A.J. Wolfe,(2004) Modulation of CRP-dependent transcription at the Escherichia coli acsP2 promoter by nucleoprotein complexes: Anti-activation by the nucleoid proteins FIS and IHF. Mol. Microbiol. 51:241–254.
- ↵D.F. Browning,D.C. Grainger,C.M. Beatty,A.J. Wolfe,J.A. Cole,S.J. Busby,(2005) Integration of three signals at the Escherichia coli nrf promoter: A role for Fis protein in catabolite repression. Mol. Microbiol. 57:496–510.
- ↵M.F. Bruist,A.C. Glasgow,R.C. Johnson,M.I. Simon,(1987) Fis binding to the recombinational enhancer of the Hin DNA inversion system. Genes & Dev. 1:762–772.
- ↵B.K. Cho,E.M. Knight,B.O. Palsson,(2006a) PCR-based tandem epitope tagging system for Escherichia coligenome engineering. Biotechniques 40:67–72.
- ↵B.K. Cho,E.M. Knight,B.O. Palsson,(2006b) Transcriptional regulation of the fad regulon genes of Escherichia coli by ArcA. Microbiology 152:2207–2219.
- ↵M.W. Covert,E.M. Knight,J.L. Reed,M.J. Herrgard,B.O. Palsson,(2004) Integrating high-throughput and computational data elucidates bacterial networks. Nature 429:92–96.
- ↵O.T. Croinin,R.K. Carroll,A. Kelly,C.J. Dorman,(2006) Roles for DNA supercoiling and the Fis protein in modulating expression of virulence genes during intracellular growth of Salmonella enterica serovar Typhimurium. Mol. Microbiol. 62:869–882.
- ↵R.T. Dame,(2005) The role of nucleoid-associated proteins in the organization and compaction of bacterial chromatin. Mol. Microbiol. 56:858–870.
- ↵K.A. Datsenko,B.L. Wanner,(2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. 97:6640–6645.
- ↵C.J. Dorman,P. Deighan,(2003) Regulation of gene expression by histone-like proteins in bacteria. Curr. Opin. Genet. Dev. 13:179–184.
- ↵S.E. Finkel,R.C. Johnson,(1992) The Fis protein: It’s not just for DNA inversion anymore. Mol. Microbiol. 6:3257–3265.
- ↵D.C. Grainger,D. Hurd,M. Harrison,J. Holdstock,S.J. Busby,(2005) Studies of the distribution of Escherichia colicAMP-receptor protein and RNA polymerase along the E. colichromosome. Proc. Natl. Acad. Sci. 102:17693–17698.
- ↵D.C. Grainger,D. Hurd,M.D. Goldberg,S.J. Busby,(2006) Association of nucleoid proteins with coding and non-coding segments of the Escherichia coli genome. Nucleic Acids Res. 34:4642–4652.
- ↵P.J. Hagerman,(1990) Sequence-directed curvature of DNA. Annu. Rev. Biochem. 59:755–781.
- ↵N.D. Heintzman,R.K. Stuart,G. Hon,Y. Fu,C.W. Ching,R.D. Hawkins,L.O. Barrera,S. Van Calcar,C. Qu,K.A. Ching,(2007) Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39:311–318.
- ↵P.N. Hengen,S.L. Bartram,L.E. Stewart,T.D. Schneider,(1997) Information analysis of Fis binding sites. Nucleic Acids Res. 25:4994–5002.
- ↵C.D. Herring,M. Raffaelle,T.E. Allen,E.I. Kanin,R. Landick,A.Z. Ansari,B.O. Palsson,(2005) Immobilization of Escherichia coli RNA polymerase and location of binding sites by use of chromatin immunoprecipitation and microarrays. J. Bacteriol. 187:6166–6174.
- ↵P. Hubner,P. Haffter,S. Iida,W. Arber,(1989) Bent DNA is needed for recombinational enhancer activity in the site-specific recombination system Cin of bacteriophage P1. The role of FIS protein. J. Mol. Biol. 205:493–500.
- ↵N.V. Hud,J. Plavec,(2003) A unified model for the origin of DNA sequence-directed curvature. Biopolymers 69:144–158.
- ↵N.V. Hud,P. Schultze,J. Feigon,(1998) Ammonium ion as an NMR probe for monovalent cation coordination sites of DNA quadruplexes. J. Am. Chem. Soc. 120:6403–6404.
- ↵R.C. Johnson,M.F. Bruist,M.I. Simon,(1986) Host protein requirements for in vitro site-specific DNA inversion. Cell 46:531–539.
- ↵A. Kelly,M.D. Goldberg,R.K. Carroll,V. Danino,J.C. Hinton,C.J. Dorman,(2004) A global role for Fis in the transcriptional control of metabolism and type III secretion in Salmonella enterica serovar Typhimurium. Microbiology 150:2037–2053.
- ↵I.M. Keseler,J. Collado-Vides,S. Gama-Castro,J. Ingraham,S. Paley,I.T. Paulsen,M. Peralta-Gil,P.D. Karp,(2005) EcoCyc: A comprehensive database resource for Escherichia coli. Nucleic Acids Res. 33:D334–D337.
- ↵T.H. Kim,L.O. Barrera,M. Zheng,C. Qu,M.A. Singer,T.A. Richmond,Y. Wu,R.D. Green,B. Ren,(2005) A high-resolution map of active promoters in the human genome. Nature 436:876–880.
- ↵H.S. Koo,H.M. Wu,D.M. Crothers,(1986) DNA bending at adenine · thymine tracts. Nature 320:501–506.
- ↵R.D. Kornberg,(1974) Chromatin structure: A repeating unit of histones and DNA. Science 184:868–871.
- ↵D. Kostrewa,J. Granzin,C. Koch,H.W. Choe,S. Raghunathan,W. Wolf,J. Labahn,R. Kahmann,W. Saenger,(1991) Three-dimensional structure of the E. coliDNA-binding protein FIS. Nature 349:178–180.
- ↵A. Landy,(1989) Dynamic, structural, and regulatory aspects of lambda site-specific recombination. Annu. Rev. Biochem. 58:913–949.
- ↵C.H. Laundon,J.D. Griffith,(1988) Curved helix segments can uniquely orient the topology of supertwisted DNA. Cell 52:545–549.
- ↵L.R. Lazarus,A.A. Travers,(1993) The Escherichia coliFIS protein is not required for the activation of tyrT transcription on entry into exponential growth. EMBO J. 12:2483–2494.
- ↵A. Martinez-Antonio,J. Collado-Vides,(2003) Identifying global regulators in transcriptional regulatory networks in bacteria. Curr. Opin. Microbiol. 6:482–489.
- ↵S. Maurer,J. Fritz,G. Muskhelishvili,A. Travers,(2006) RNA polymerase and an activator form discrete subcomplexes in a transcription initiation complex. EMBO J. 25:3784–3790.
- ↵S.M. McLeod,S.E. Aiyar,R.L. Gourse,R.C. Johnson,(2002) The C-terminal domains of the RNA polymerase alpha subunits: Contact site with Fis and localization during co-activation with CRP at the Escherichia coli proP P2 promoter. J. Mol. Biol. 316:517–529.
- ↵L.D. Murphy,S.B. Zimmerman,(1997) Isolation and characterization of spermidine nucleoids from Escherichia coli. J. Struct. Biol. 119:321–335.
- ↵G. Muskhelishvili,A. Travers,(2003) Transcription factor as a topological homeostat. Front. Biosci. 8:D279–D285.
- ↵G. Muskhelishvili,A.A. Travers,H. Heumann,R. Kahmann,(1995) FIS and RNA polymerase holoenzyme form a specific nucleoprotein complex at a stable RNA promoter. EMBO J. 14:1446–1452.
- ↵O. Ninnemann,C. Koch,R. Kahmann,(1992) The E. coli fis promoter is subject to stringent control and autoregulation. EMBO J. 11:1075–1083.
- ↵C.Q. Pan,S.E. Finkel,S.E. Cramton,J.A. Feng,D.S. Sigman,R.C. Johnson,(1996) Variable structures of Fis-DNA complexes determined by flanking DNA-protein contacts. J. Mol. Biol. 264:675–695.
- ↵S.J. Park,G. Chao,R.P. Gunsalus,(1997) Aerobic regulation of the sucABCD genes of Escherichia coli, which encode alpha-ketoglutarate dehydrogenase and succinyl coenzyme A synthetase: Roles of ArcA, Fnr, and the upstream sdhCDAB promoter. J. Bacteriol. 179:4138–4142.
- ↵B.J. Paul,W. Ross,T. Gaal,R.L. Gourse,(2004) rRNA transcription in Escherichia coli. Annu. Rev. Genet. 38:749–770.
- ↵D. Perkins-Balding,D.P. Dias,A.C. Glasgow,(1997) Location, degree, and direction of DNA bending associated with the Hin recombinational enhancer sequence and Fis-enhancer complex. J. Bacteriol. 179:4747–4753.
- ↵L. Postow,C.D. Hardy,J. Arsuaga,N.R. Cozzarelli,(2004) Topological domain structure of the Escherichia coli chromosome. Genes & Dev. 18:1766–1779.
- ↵B. Ren,F. Robert,J.J. Wyrick,O. Aparicio,E.G. Jennings,I. Simon,J. Zeitlinger,J. Schreiber,N. Hannett,E. Kanin,(2000) Genome-wide location and function of DNA binding proteins. Science 290:2306–2309.
- ↵N.B. Reppas,J.T. Wade,G.M. Church,K. Struhl,(2006) The transition between transcriptional initiation and elongation in E. coli is highly variable and often rate limiting. Mol. Cell 24:747–757.
- ↵K. Rippe,P.H. von Hippel,J. Langowski,(1995) Action at a distance: DNA-looping and initiation of transcription. Trends Biochem. Sci. 20:500–506.
- ↵C. Robinow,E. Kellenberger,(1994) The bacterial nucleoid revisited. Microbiol. Rev. 58:211–232.
- ↵K. Robison,A.M. McGuire,G.M. Church,(1998) A comprehensive library of DNA-binding site matrices for 55 proteins applied to the complete Escherichia coli K-12 genome. J. Mol. Biol. 284:241–254.
- ↵M. Rochman,M. Aviv,G. Glaser,G. Muskhelishvili,(2002) Promoter protection by a transcription factor acting as a local topological homeostat. EMBO Rep. 3:355–360.
- ↵T.D. Schneider,(1997) Information content of individual genetic sequences. J. Theor. Biol. 189:427–441.
- ↵T.D. Schneider,R.M. Stephens,(1990) Sequence logos: A new way to display consensus sequences. Nucleic Acids Res. 18:6097–6100.
- ↵R. Schneider,R. Lurz,G. Luder,C. Tolksdorf,A. Travers,G. Muskhelishvili,(2001) An architectural role of the Escherichia coli chromatin protein FIS in organising DNA. Nucleic Acids Res. 29:5107–5114.
- ↵R.K. Shultzaberger,L.R. Roberts,I.G. Lyakhov,I.A. Sidorov,A.G. Stephen,R.J. Fisher,T.D. Schneider,(2007) Correlation between binding rate constants and individual information of E. coli Fis binding sites. Nucleic Acids Res. 35:5275–5283.
- ↵D. Skoko,D. Yoo,H. Bai,B. Schnurr,J. Yan,S.M. McLeod,J.F. Marko,R.C. Johnson,(2006) Mechanism of chromosome compaction and looping by the Escherichia colinucleoid protein Fis. J. Mol. Biol. 364:777–798.
- ↵R. Stefl,H. Wu,S. Ravindranathan,V. Sklenar,J. Feigon,(2004) DNA A-tract bending in three dimensions: Solving the dA4T4 vs. dT4A4 conundrum. Proc. Natl. Acad. Sci. 101:1177–1182.
- ↵R.A. Stein,S. Deng,N.P. Higgins,(2005) Measuring chromosome dynamics on different time scales using resolvases with varying half-lives. Mol. Microbiol. 56:1049–1061.
- ↵B. ten Heggeler-Bordier,W. Wahli,M. Adrian,A. Stasiak,J. Dubochet,(1992) The apical localization of transcribing RNA polymerases on supercoiled DNA prevents their rotation around the template. EMBO J. 11:667–672.
- ↵J.F. Thompson,U.K. Snyder,A. Landy,(1988) Helical-repeat dependence of integrative recombination of bacteriophage lambda: Role of the P1 and H1 protein binding sites. Proc. Natl. Acad. Sci. 85:6323–6327.
- ↵M.Y. Tolstorukov,K.M. Virnik,S. Adhya,V.B. Zhurkin,(2005) A-tract clusters may facilitate DNA packaging in bacterial nucleoid. Nucleic Acids Res. 33:3907–3918.
- ↵A. Travers,G. Muskhelishvili,(1998) DNA microloops and microdomains: A general mechanism for transcription activation by torsional transmission. J. Mol. Biol. 279:1027–1043.
- ↵D. Ussery,T.S. Larsen,K.T. Wilkes,C. Friis,P. Worning,A. Krogh,S. Brunak,(2001) Genome organisation and chromatin structure in Escherichia coli. Biochimie 83:201–212.
- ↵J.T. Wade,D.C. Roa,D.C. Grainger,D. Hurd,S.J. Busby,K. Struhl,E. Nudler,(2006) Extensive functional overlap between sigma factors in Escherichia coli. Nat. Struct. Mol. Biol. 13:806–814.
- ↵H. Wu,K.L. Tyson,J.A. Cole,S.J. Busby,(1998) Regulation of transcription initiation at the Escherichia coli nir operon promoter: A new mechanism to account for co-dependence on two transcription factors. Mol. Microbiol. 27:493–505.
- ↵J. Xu,R.C. Johnson,(1995) aldB, an RpoS-dependent gene in Escherichia coli encoding an aldehyde dehydrogenase that is repressed by Fis and activated by Crp. J. Bacteriol. 177:3166–3175.
- ↵J. Zhang,Y. Zeuner,A. Kleefeld,G. Unden,A. Janshoff,(2004) Multiple site-specific binding of Fis protein to Escherichia coli nuoA-N promoter DNA and its impact on DNA topology visualised by means of scanning force microscopy. ChemBioChem 5:1286–1289.
- ↵H. Zhi,X. Wang,J.E. Cabrera,R.C. Johnson,D.J. Jin,(2003) Fis stabilizes the interaction between RNA polymerase and the ribosomal promoter rrnB P1, leading to transcriptional activation. J. Biol. Chem. 278:47340–47349.
Notes
[1] [Supplemental material is available online at www.genome.org.]
[2] Article published online before print. Article and publication date are at http://www.genome.org/cgi/doi/10.1101/gr.070276.107.

