Defensins and the dynamic genome: What we can learn from structural variation at human chromosome band 8p23.1
- 1 Department of Genetics, University of Leicester, Leicester LE1 7RH, United Kingdom;
- 2 Wessex Regional Genetics Laboratory, National Genetics Reference Laboratory (Wessex) and Human Genetics Division, School of Medicine, University of Southampton, Southampton SO17 1BJ, United Kingdom;
- 3 Institute of Genetics, University of Nottingham, Nottingham NG7 2RD, United Kingdom
Abstract
Over the past four years, genome-wide studies have uncovered numerous examples of structural variation in the human genome. This includes structural variation that changes copy number, such as deletion and duplication, and structural variation that does not change copy number, such as orientation and positional polymorphism. One region that contains all these types of variation spans the chromosome band 8p23.1. This region has been studied in some depth, and the focus of this review is to examine our current understanding of the variation of this region. We also consider whether this region is a good model for other structurally variable regions in the genome and what the implications of this variation are for clinical studies. Finally, we discuss the bioinformatics challenges raised, discuss the evolution of the region, and suggest some future priorities for structural variation research.
Structural variation and its importance
Polymorphic variation in genome structure has been appreciated for many years. Structural variation was apparent in early studies of human DNA polymorphisms, because of the ability to analyze duplications and deletions using Southern blots, and the phenotypic variation caused by copy number variation in genes of, for example, the Rh locus, the opsin locus, and the alpha-globin locus (Baine et al. 1976; Nathans et al. 1986; Colin et al. 1991; Wolf et al. 1999).
More recent studies have revealed the human genome to be considerably more variable than a single nucleotide polymorphism (SNP)-focused eye would have us believe. Initial studies found parts of the genome over- or under-represented compared to a reference genome and therefore variable in copy number, but the regions found in each study had minimal overlap with each other, suggesting that each study had sampled only a small proportion of the number of regions that are copy number variable (CNV) (Iafrate et al. 2004; Sebat et al. 2004; Sharp et al. 2005). A comprehensive study on 270 individuals used for the HapMap project confirmed this, and ∼12% of the genome showed an increase or decrease in DNA dosage in at least one of these individuals (Redon et al. 2006).
While array-CGH (comparative genomic hybridization) has driven a boom in copy number variation discovery, the characterization of balanced structural variation, in which there is no net gain or loss of sequence between individuals, has been limited by current technology (Baptista et al. 2008). Inversion polymorphisms have been discovered mostly from comparative sequence analysis of humans and chimpanzees (Feuk et al. 2005) and from paired-end mapping, where orientations of sequences from either end of a piece of DNA are inverted with respect to the reference sequence (Tuzun et al. 2005; Korbel et al. 2007). Current inversion genotyping methods rely on identification of the breakpoint and the assumption that all inverted alleles share the same breakpoint. This will be true only when the inversion event has happened just once, and in these cases, the inversion can also be genotyped by typing surrogate SNPs within the inversion and in complete association with it (Stefansson et al. 2005). Inversions with breakpoints within large inverted repeats can be genotyped using single-molecule haplotyping (Turner et al. 2006), but in complex repeat regions with many kilobases of inverted repeats or with multiple inversion breakpoints, even this method is fallible. This is because the location of the inverted repeat mediating the inversion must be known. At present, molecular cytogenetics and pulsed-field gel electrophoresis remain the only methods to genotype inversions with complex breakpoints or multiple origins. Polymorphic translocations, in which a sequence is polymorphically present at a different position, either within the chromosome or on a different chromosome, are common in subtelomeric regions, but, again, cytogenetics remains the most reliable way to type them (Wong et al. 1990; Martin-Gallardo et al. 1995; Linardopoulou et al. 2005). In short, current technologies are not sufficient for the analysis of medium- and large-scale structural variation in all its forms. Furthermore, small-scale structural variation is not yet fully characterized and requires more method development.
Such extensive variation in genomic structure suggests that at least some will have consequences on phenotype and disease susceptibility. These could be mediated in many ways, including direct DNA dosage or the unmasking of deleterious recessive mutations. Recent studies have implicated copy number variation as affecting gene expression, protein expression, and phenotypic variation in both genome-wide and locus-specific studies (Johansson et al. 1993; Stranger et al. 2007). For example, increased copy number of the salivary amylase gene cluster increases expression of the protein and ability to digest starch (Perry et al. 2007). Similarly, increased copy number of the beta-defensin region increases risk of the inflammatory skin disease psoriasis (Hollox et al. 2008). In both cases, a direct gene dosage effect is likely to link diplotype to phenotype. Structural variation can also act as a substrate for larger genomic rearrangements with profound clinical consequences (Giglio et al. 2001, 2002; Linardopoulou et al. 2005).
The application of array CGH to clinical patients is already revealing a rising number of CNVs, many of which are initially novel (Menten et al. 2006). Detailed clinical analysis of the phenotypes of such rare cases can give important insights into gene function, and there are ongoing efforts to relate specific chromosome gains and losses to dosage imbalance of individual genes. These are being facilitated by international databases such as DECIPHER (http://www.sanger.ac.uk/PostGenomics/decipher/) and ECARUCA (http://www.ecaruca.net).
There have been several excellent reviews recently that discuss the current genome-wide CNV studies and make general inferences from the pattern and nature of the CNVs found (Freeman et al. 2006; Beckmann et al. 2007; Cooper et al. 2007). In this review, we discuss the structural variation at the human beta-defensin region, what we know about it, and argue that this can inform studies on other structurally variable regions of human and other mammalian genomes.
Defensins—Renaissance molecules
Defensins are short cationic peptides that are expressed in epithelia and leukocytes and have an important role in the innate immune system. They are divided into three types: alpha, beta, and theta, depending on the pattern of disulfide-bridge formation between the six conserved cysteine residues that define defensins (Ganz 2003). They were initially characterized as effective antimicrobial peptides, killing both Gram-positive and Gram-negative bacteria and fungi (Klotman and Chang 2006; Pazgier et al. 2006), and are thought to kill microbes by depolarizing and permeabilizing the cell membrane (Lehrer et al. 1989; Kagan et al. 1990; Hill et al. 1991). Later studies showed that they have extensive cell-signaling activity, recruiting immature dendritic cells to the site of infection, which have been mirrored by other studies suggesting that chemokines, such as CCL20, also have potent antimicrobial activity (Yang et al. 1999, 2003; Biragyn et al. 2002). It has been suggested that these chemokines and defensins should be regarded as one functional group, termed “alarmins” (Oppenheim and Yang 2005). Alpha, beta, and theta defensins have also been shown to have potent antiviral activity, in particular against HIV-1 (Cole et al. 2002; Mackewicz et al. 2003; Quinones-Mateu et al. 2003; Chang and Klotman 2004). One beta-defensin signals through the melanocortin receptor in dogs and humans and controls coat color in dogs (Candille et al. 2007). Others have a function in reproduction; for example, DEFB126 coats the glycocalyx of sperm and is involved in attaching sperm to the oviduct epithelia (Yudin et al. 2005; Tollner et al. 2008). Beta-defensins have also evolved to become venom in the platypus (Whittington et al. 2008). It is likely that different defensins are much more than antimicrobial, and further functional work will uncover different roles for what is increasingly regarded as a multifunctional gene family.
Without analysis of synteny relationships, true orthologs of defensins can be difficult to identify because of the amino acid differences between defensins of different species. It is therefore difficult to determine whether “defensins” in insects and plants are true orthologs of vertebrate defensins. Nevertheless, analysis of the defensin family within mammals reveals a considerable amount about the evolution of these genes. Rapid duplication and divergence is a recurring theme, as would be expected in a gene family central to host–pathogen interaction (Hughes 1999; Lynn et al. 2004). There is evidence of positive selection in mammalian alpha-defensins and in rodent and primate beta-defensins (Morrison et al. 2003; Semple et al. 2005; Hollox and Armour 2008). Moreover, in rodents, there has been duplication and pseudogenization of the beta-defensin family, generating a distinct beta-defensin repertoire in the rat and mouse (Morrison et al. 2003).
The story is more complex than this may suggest. Despite evidence for duplication and pseudogenization in the rodents, evidence from primates and other mammalian groups suggests conservation of synteny (Patil et al. 2005). There has been positive selection of the amino acid sequence of some, but not all, beta-defensins in primates: the genes involved in the CNV region seem not to have changed much in the primate lineage (Hollox and Armour 2008). These genes are CNV in macaques as well as humans, suggesting persistence of copy number variation for more than 35 million years (Myr) (Lee et al. 2008). It is possible that this variation has prevented gene sequence diversification by facilitating gene conversion events between paralogs, although further investigation into the rates of gene conversion and selection pressure on these paralogs is required.
Defensin variation at 8p23.1
The alpha-defensin DEFA1A3 and the beta-defensins DEFB4, DEFB103, DEFB104, DEFB105, DEFB106, and DEFB107 map to 8p23.1 and vary in copy number independently: the alpha-defensin as a 19-kb tandemly repeated unit and the beta-defensins on a copy number variable unit that is at least 250 kb in size, but the exact size and breakpoints of the copy number variable region are not known (Fig. 1). This latter repeat unit also contains SPAG11, an alternatively spliced gene formed by a head-to-head fusion of two beta-defensins, which codes for a protein that shows potent antimicrobial activity and is present on spermatozoa, perhaps providing antimicrobial protection for its carrier (Yenugu et al. 2003; Zanich et al. 2003). The other alpha-defensins—DEFA4, DEFA5, and DEFA6—and the beta-defensin DEFB1 do not show CNV (Hollox et al. 2003; Aldred et al. 2005; Linzmeier and Ganz 2005).
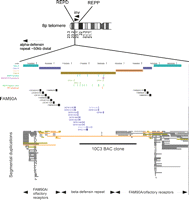
Genome assembly of the 8p23.1 region Highlighted is the REPD region containing one of two beta-defensin repeat units assembled, separated by a gap. The repeat classes II–V are from Taudien et al. (2004). Sites for assays for beta-defensin copy number are also shown, including PRT, MAPH, MLPA, and REDVR (Hollox et al. 2008). EPEV-1–EPEV-3 refer to simple tandem repeats used for copy number and segregation analysis in pedigrees. The segmental duplication track is based on the data from Bailey et al. (2001); (light to dark gray) 90%–98% similarity with its duplicate; (yellow) 98%–99% similarity; (orange) >99% similarity. Based on UCSC Genome Browser (http://genome.ucsc.edu) build hg18.
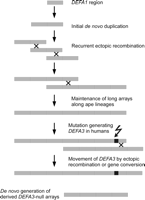
CNV of DEFA1 had been suspected from somatic cell hybrid mapping of chromosome 8 (Mars et al. 1995). We, and others, showed that the DEFA1 and DEFA3 genes were variants of the same gene that differed by one nucleotide (Aldred et al. 2005; Linzmeier and Ganz 2005). The renamed DEFA1A3 gene is encoded on a tandem repeat that has a diploid copy number between four and 11. The nucleotide change leading to DEFA3 is human-specific and is most frequently at the most proximal position in the tandem array. This suggests a relatively recent origin of this variant, although in 50% of arrays, the DEFA3 variant is not at the proximal position but elsewhere in the array, suggesting a recent history of non-allelic recombination (NAHR) or gene conversion within this array (Fig. 2) (Aldred et al. 2005). In native UK individuals, DEFA3 variant is absent in 10% of the population, and in sub-Saharan Africans (Yoruba), it is absent in 37% of the population. DEFA3 absence is associated with one SNP haplotype in Europeans but shows no association with any SNP haplotype in other populations (Ballana et al. 2007). The theta-defensin DEFT1 is encoded on the same 19-kb repeat as DEFA1A3. In humans, gorillas, and chimpanzees, this is an inactive pseudogene, but is active in other primate lineages tested (Tang et al. 1999; Nguyen et al. 2003). DEFA1 is present in multiple copies in other apes, making it likely that this has been variable in copy number since before the divergence of gibbons and great apes around 25 million years ago (Mya) (Aldred et al. 2005).
Evolution of the DEFA1A3 CNV region. Following an initial de novo duplication at least 25 Mya, recurrent ectopic recombination generates long arrays of DEFA1 repeats. A point mutation in humans generates the DEFA3 gene, which is shuffled through the array by continuing ectopic recombination.
The beta-defensin genes involved in CNV are on a large genomic repeat unit within a “sea” of more complex CNV involving retroviral elements and olfactory repeat (OR) regions, collectively known as “REPD” (for repeat distal) (Giglio et al. 2001). Another smaller OR region called “REPP” (repeat proximal) is 5 Mb proximal on 8p23.1 and shares a high level of identity with REPD (Fig. 1; Sugawara et al. 2003). The total diploid copy number of the beta-defensin region ranges from one to 12, and commonly between two and seven (Hollox et al. 2003). We have observed only one individual with a copy number of one in more than 1500 individuals with DNA from blood, suggesting that the null allele exists but is very rare (allele frequency ∼ 0.2%). Given the functional importance of these genes, the null homozygote may be strongly deleterious, and null alleles generated by recurrent mutation rapidly removed from the population by purifying selection. At the other end of the scale, exact definition of integer copy number becomes very difficult. Large diploid copy numbers are, at least in part, due to recent expansion of the beta-defensin copy number in independent lineages, and such expansions can be seen directly under G-band chromosome staining as a euchromatic variant (Hollox et al. 2003; Barber et al. 2005).
The repeat structure has been studied in detail, and four classes of repeat region have been identified (Classes II–V) (Fig. 1; Taudien et al. 2004). The main beta-defensin region appears to correspond to Class V, with more complex repeat-rich regions flanking it (Fig. 1). At the ends of the beta-defensin genomic repeat unit—perhaps at the shore of the sea of complex olfactory receptor repeats—are FAM90A gene clusters that are on Class III repeats. FAM90A genes are on several other chromosomes and are expressed in several different tissues, although their function is not known (Bosch et al. 2007).
Although the >250-kb repeat unit is too large to be captured by current paired-end mapping approaches, such methods can reveal information on the flanking regions surrounding the repeat unit (Korbel et al. 2007; Kidd et al. 2008). The fosmid paired-end mapping approach has the particular advantage that the structural variant is cloned as a fosmid and can be sequenced directly, providing sequence-level resolution (Iafrate et al. 2004). The disadvantage is that it is a time-consuming and expensive approach, with only a few genomes analyzed so far. Data presented so far in the Structural Variation Database (http://humanparalogy.gs.washington.edu/structuralvariation/) on four genomes is consistent with the whole beta-defensin repeat unit varying as a block with no internal small duplications or deletions. However, variation exists where the repeat unit meets the flanking sea of more complex repeats and sequencing of these particular fosmids may reveal, for the first time, some sequence-level information about CNV of the beta-defensin region. Hopefully, paired-end approaches combined with novel sequencing technologies may shed light on the small-scale variation at this locus, although novel approaches are needed to define the large-scale variation at a sequence-level resolution.
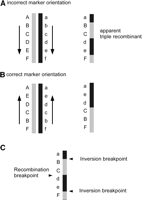
Sequence-based approaches and paired-end mapping cannot easily capture the large inversion between REPD and REPP. However, physical methods such as fluorescent in situ hybridization (FISH) and genetic analysis can type individuals for this inversion polymorphism. The region between REPP and REPD has been shown to be polymorphically inverted by FISH analysis with a frequency of 25% in Europeans (Giglio et al. 2001). This has been confirmed genetically and has since been used to confirm orientation of the region in certain CEPH individuals: recombination between inversion homozygotes results in an apparent triple recombinant when the genetic markers are arranged using a map based on a standard (non-inverted) assembly (Fig. 3) (Broman et al. 2003). As expected, the inversion breakpoints map to REPP and REPD, and analysis of informative markers within these repetitive regions may resolve the inversion breakpoints with greater accuracy. SNPs and other polymorphisms carried on an inverted region are expected to be isolated from recombination with non-inverted regions and to form a non-recombining haplotype clade. However, we and others (Broman et al. 2003) have shown that multiple alleles at many loci are found on both inverted and non-inverted haplotypes, suggesting a recurrent or frequently reverting process of inversion.
Apparent triple recombinants reflect inverted marker orientation. (A,B) A single recombination within an inversion between two inverted chromosomes is revealed as an apparent triple recombinant if the markers are assumed to be in the non-inverted orientation (Broman et al. 2003). (C) Analysis of the apparent triple recombinants can localize the site of recombination and the two breakpoints of the inversion.
Methods for typing defensin copy number
Accurately typing copy number is both more difficult than and different from SNP genotyping; it should measure a quantitative difference rather than a qualitative difference. It is clear that progress is limited by technology; just as discovery of CNVs was limited until the development of array-CGH technology, accurate CNV typing is limited by the technologies available. Nevertheless, progress has been made in both typing precision and accuracy (Clayton et al. 2005; McCarroll and Altshuler 2007). The accuracy is primarily a consequence of correct normalization and controls and is aided by the fact that we are confident that the underlying biological reality of germline copy number variation at any one sequence position is gain and loss of integer numbers of copies (2, 3, 4, etc., no intermediates), and where empirical data of sufficient accuracy have been examined in detail, this prediction has been borne out for somatic DNA (Armour et al. 2007). This has two important consequences: first, there should be adequate availability of well-characterized copy number reference controls to allow for comparison of results across laboratories, and, second, if any CNVs do show frequent somatic variation, they will require even more effort to type accurately, since the assumption of integer copy number variation may not be applicable.
Precision is a technical consequence of the method used for typing and can be regarded as the reproducibility of the result on repeated testing. Both precision and accuracy reflect on each other: for example, a highly precise method will produce clustering of results around integer copy number, and therefore calibrating each copy number to the center of these clusters will overcome any accuracy differences between experiments. Despite the importance of accuracy and precision in interpreting results correctly, very few published studies so far provide any analysis on the accuracy and precision of their data sets (McCarroll and Altshuler 2007; McCarroll 2008). Even with a precise method, difference in accuracy between cases and controls in an association study caused, for example, by inadequate normalization to known controls can result in a bias that is spuriously interpreted as a significant association. Where data have been shown, it is clear that, at present, no one method is quite good enough for large case-control studies and that power comes from combining methods and repeat typing (Hollox et al. 2008). We will briefly discuss various copy number typing methods that have been applied to the defensin loci.
Multiplex amplifiable probe hybridization (MAPH)
The first method used to measure beta-defensin copy number was multiplex amplifiable probe hybridization (Armour et al. 2000; Hollox et al. 2003). We constructed several probes across the beta-defensin region (Fig. 1) and showed that, with few exceptions, copy number reported from each probe was equivalent and therefore that the repeat unit varied as a whole repeat unit without substantial heterogeneity in structure (Hollox et al. 2005, 2008; Groth et al. 2008). The strength of the method is that any sequence, except high copy number repeats such as Alus and extremely GC-rich or GC-poor regions, can be used as a probe, and probes are straightforward to produce in a laboratory. The disadvantage is that each hybridization requires 1 μg of DNA, with duplicate hybridizations required for increased accuracy. This cannot be overcome by whole genome-amplification, which is known to introduce bias in relative copy number (Hosono et al. 2003). Without this, MAPH can seriously deplete DNA collections. The technique also requires manipulation of very small dry filters, which is time-consuming and can lead to sample loss.
Multiplex ligation-dependent probe amplification (MLPA)
This method, similar to MAPH, uses hybridization and ligation of two half-probes to specifically record the amount of sequence in a sample (Schouten et al. 2002). It is a single-tube assay and requires less DNA (100–250 ng), although that amount is still significant compared to PCR-based methods. A probe set for the beta-defensin region is commercially available, with probes for several defensin genes across the variable region (Fig. 1). When this method was used on 135 samples, it showed equivalent precision to a single paralog ratio test (see below) and MAPH (Armour et al. 2007).
Quantitative real-time PCR
This method, using fluorescent techniques such as TaqMan allowing the real-time measurement of PCR product accumulation, is increasingly popular in measuring copy number variation and has been applied to the beta-defensin region (Linzmeier and Ganz 2005; Chen et al. 2006; Fellermann et al. 2006). Its strength is that, in theory, it can measure the copy number of any sequence and requires DNA amounts sufficient for PCR, typically 5–10 ng. However, typical studies using this method to type genomic copy number typically do not present analysis of the error rate or a thorough test of the method by comparison from corroborating data from other methods. Quantitative real-time PCR has found its niche analyzing expression levels of genes, which often differs by greater than 10-fold (Bustin 2002). It is likely that reliably distinguishing four and five copies, for example, based on numbers inferred from a near-exponential curve, is beyond the resolution capabilities of this approach.
Paralog ratio test (PRT)
The paralog ratio test (PRT) is essentially a development of comparative PCR, but where test and reference loci are amplified by the same primer pair (Deutsch et al. 2004; Armour et al. 2007). This improves reproducibility by making the amplification kinetics of test and reference loci very similar. This requires careful design of primers, often using diverged dispersed repeat sequences so that the primers amplify the sequence on the region of interest, and one sequence elsewhere on the genome, preferably on another chromosome to minimize potential gene conversion between the test and reference sequences. For example, a PRT assay for the beta-defensin region has been designed to specifically amplify a heat-shock protein pseudogene on the copy number variable region and on chromosome 5. Figure 1 shows the location of the PRT assay relative to the genes on the repeat unit.
The sequence requirements for PRT assay development mean that it is limited to certain sequences, and an assay cannot necessarily be designed for a small sequence region, for example, one specific exon. However, the precision and accuracy are equivalent to those of MLPA and MAPH, and it has the advantage of potential high-throughput analysis and the small DNA requirement of PCR-based methods (Armour et al. 2007).
Simple tandem repeat (STR) analysis
Simple tandem repeats within CNV regions are likely to show multiple alleles, up to a maximum of the number of copies of CNV. These alleles, like single-copy STRs, can be amplified and resolved by length using electrophoresis. STR analysis is valuable for tracking individual copies of a repeat through a pedigree because of the high informativeness of these loci: each repeat has a high chance of carrying a different length allele (Hollox et al. 2003). There is also the possibility of simply counting the alleles as a discrete measure of copy number, and three STRs have been used to distinguish ambiguous copy number diplotypes (Fig. 1, shown as “EPEV”; Hollox et al. 2005).
Array-CGH
Although primarily intended for discovering CNV rather than typing it, analysis of the precision of the array-CGH technique suggests that it is at least equivalent to other methods. The main advantage is its multiplicity, in that a whole-genome bacterial artificial chromosome (BAC) array, for example, analyzes the whole genome for copy number variation and can, at least for the beta-defensin locus, genotype it reasonably well (Redon et al. 2006; Armour et al. 2007). It has several disadvantages, not least being the cost per test and amount of DNA required. Another issue is that while other genotyping methods measure copy number of a short segment, BAC array-CGH measures the copy number of a large segment of DNA (100 kb), and hybridization intensities can be influenced by variation in other sequences. For example, the clone RPCI-11 10C3 overlaps regions that are thought to be more highly variable than the beta-defensin “island” (Fig. 1), and this may account for differences in accuracy between, for example, PRT and BAC array-CGH, especially at higher copy numbers (Armour et al. 2007).
Hybridization intensity from SNP genotyping chips
Hybridization chips used for genotyping single nucleotide polymorphisms by comparative hybridization intensities of genomic DNA to the oligonucleotides representing each allele also give information on copy number from the raw intensity signal strength of the hybridization. This was the second platform used by Redon et al. (2006) for their genome-wide analysis of CNV. The principal disadvantage for most chips is that they were designed to analyze SNPs not CNVs, and CNV-rich segmental duplications, including the 8p23.1 region, have been deliberately under-represented in the array. New chips have probes designed to map to known CNV regions, and it will be important to assess how accurately and precisely they can genotype known multiallelic CNVs such as the beta-defensin region.
The alpha-defensin copy number polymorphism distal to the beta-defensin region has been the subject of less intensive investigation, primarily because of the difficulty in accurately diplotyping a locus with a copy number between four and 11. The small 19-kb repeat element, and surrounding single copy sequence, allows straightforward pulsed-field gel analysis of the region and accurate genotyping of control samples, but this is not a practical method for large sample numbers (Aldred et al. 2005). Because of the small repeat size, loci suitable for PRT are limited, and accurate diplotyping of this locus remains an area of active research.
Over the next few years, there is likely to be a plethora of association studies using genotyping methods and study designs of varying quality. Statistical methods to deal with CNV in population- and family-based data sets, especially those that incorporate a measure of the inaccuracy of the CNV genotyping method, will be an essential tool for such studies (Kosta et al. 2007; Ionita-Laza et al. 2008).
Disease studies and clinical relevance of structural variation at 8p23.1
Association with common multifactorial diseases
As discussed above, several copy number variable genes have been shown to affect susceptibility to disease. In addition, structural variations predispose to large structural rearrangements. Both these mechanisms have been shown to cause disease in the 8p23.1 region. The first positive study associated beta-defensin copy number with colonic Crohn’s disease, an inflammatory disease of the bowel (Fellermann et al. 2006). While noting the caveat that this was an unreplicated study of small sample size, the authors found an association between increased genomic copy number with increased expression of DEFB4 in the gut and protection against Crohn’s disease. They also provided a plausible mechanistic explanation for their results: low copy number predisposed to Crohn’s disease because of a lower anti-infection barrier. The next study genotyped beta-defensin copy number for two larger cohorts of psoriasis patients and controls from the Netherlands and Germany. Increasing beta-defensin copy number was shown to increase susceptibility to psoriasis in both cohorts. The data showed a linear relationship between copy number and disease relative risk, so that each additional copy increased the risk by about 34 percentage points (95% CI 25–43). This represents between a 2.2× and 3.1× difference in risk across the common copy number variation of two to seven copies (Hollox et al. 2008).
Association of beta-defensin region copy number with disease immediately suggests a mechanism for mediating its effect: an increase in gene product reflected in gene dosage. However, it does not suggest which gene on the repeat region is involved—there are at least seven, and a likely candidate must be inferred from other work. hBD2 protein, encoded by the DEFB4 gene, was initially discovered in extracts of psoriatic plaques (Harder et al. 1997) and is up-regulated in psoriatic skin, so this is the best candidate, but none of the other genes can be ruled out. Functional studies are required to test each candidate gene in turn. For DEFB4, there is a correlation with mRNA levels in lymphoblastoid cell lines and in gut mucosa (Hollox et al. 2003; Fellermann et al. 2006). The increase in risk in psoriasis increases linearly with an increase in dosage, supporting the idea that the risk factor is gene dosage itself and not some unknown sequence variant within the beta-defensin region (Hollox et al. 2008).
The relationship between expression level and genomic copy number may not always be so straightforward. For DEFA1A3, expression level is not simply correlated with copy number, yet the expression ratio of DEFA1:A3 is correlated with the genomic ratio of DEFA1:A3 (at least in the small sample size tested). We would predict that this ratio may be a risk factor for disease rather than total copy number.
Crohn’s disease has not been associated with variation at 8p23.1 in whole-genome association studies, and neither Crohn’s disease nor psoriasis has shown linkage to 8p23.1. The absence of an association can be explained by the low association of multiallelic CNV diplotype with neighboring SNP genotypes (Redon et al. 2006), which means that CNV will not be effectively interrogated by genotyping chips assaying tagSNPs flanking the CNV region. The absence of linkage can, at least in the psoriasis study, be attributed to the fact that the effect size is not strong enough to be detected. The common nature of the variation and the difficulty of interpreting segregation patterns may also cloud any potential linkage to 8p23.1.
The crucial role that defensins play in the innate immune response suggests that alpha- and beta-defensin genomic copy number variations may be excellent candidate loci for other inflammatory diseases. There have been several diseases linked to 8p23.1, including type II diabetes (Kim et al. 2004; Pezzolesi et al. 2004), asthma (Xu et al. 2001a; Dizier et al. 2003), and prostate cancer (Xu et al. 2001b; Wiklund et al. 2003). Given that linkage will only detect strong effects, the difficulty in interpreting segregation patterns and the high frequency of variation, absence of linkage signal does not preclude the CNV as a candidate locus for other diseases.
Predisposition to large-scale imbalances of 8p
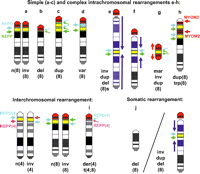
In addition to modifying the risk of common diseases, the 8p23.1 defensins are part of the repeat structures that predispose to an increasing number of recurrent genomic disorders with phenotypic consequences that range from mild to severe. The olfactory repeat regions REPD and REPP (Fig. 1) give rise to large rearrangements that range from simple deletions of the interval between REPP and REPD to complex rearrangements in which one or more large-scale imbalances may be generated (Fig. 4b–h).
Large-scale rearrangements mediated by 8p23.1 repeats. Schematic idiograms of the short arm of chromosome 8 simplified with color coding: (blue) REPD; (green) REPP; (yellow) the region between these repeats; (red) the region between REPD and the telomere. (a) The common inversion of 8p23.1. (b) deletion of the interval between the repeats with a residue of both repeats remaining (small light blue and green arrows). (c) Duplication of the interval between the repeats with three copies of REPP mixed with REPD repeats at two of the three repeat loci. (d) Copy number variation in which multiple copies of the DEFB4 repeat (light blue) can mimic the appearance of a duplication (c). (e,f) Classical inverted duplicated and deleted chromosome 8s with a proximal duplication breakpoint in the 8 centromere (e) or in band 8p21.2 (f). (Dark blue) The duplicated regions (with an arrow to indicate their orientation); (red) the deleted region. (g) The supernumerary neocentric marker chromosome 8 with the duplicated regions (red, with an arrow to indicate their orientation) flanking the region between the repeats (small light blue and green arrows). (h) The duplicated and triplicated chromosome 8: (yellow) the duplicated region; (red) the triplicated region; (red arrows) the approximate position of the MYOM2 repeats. (i) The unbalanced der(4)(t(4;8)(p16.1p23.1) translocation derived from mothers who are doubly heterozygous for the common inversions of chromosome 4 (left pair) and chromosome 8 (middle pair) producing an unbalanced der(4) chromosome with deletion of the distal (white) segment of 4p and duplication of the distal (red) section of 8p with a residue of REPD(8) (light blue) and REPP(4) (pink) repeats. (j) Terminal deletion of 8p resulting from post-zygotic breakage of a dicentric intermediate with the diagonal line indicating the distinct cell lines with the terminally deleted 8p (left side) and a classic inv dup del(8) chromosome (right side).
The 8p23.1 deletion syndrome (Fig. 4b) is associated with developmental delay, behavioral problems, and congenital heart disease (Devriendt et al. 1995) and is a result of ectopic recombination (also known as non-allelic homologous recombination [NAHR]) between REPD and REPP. Ectopic recombination is also thought to be the mechanism behind the reciprocal duplication syndrome (Fig. 4c) associated with mild dysmorphism and developmental and speech delay (Barber 2005; Barber et al. 2008). Accurate dosage imbalance techniques are essential to distinguish these genuine duplications from high copy beta-defensin CNVs that appear identical under the light microscope (Fig. 4d) (Barber et al. 2005). In a further recent example involving the corresponding paired repeats in band 4p16, five copies of a 750-kb band containing DEFB131 cosegregated with autosomal dominant microtia (Balikova et al. 2008).
Maternal heterozygosity for the common inversion (Fig. 4a) predisposes to recurrent intra- and interchromosomal rearrangements as a result of meiotic ectopic recombination (Floridia et al. 1996; Giglio et al. 2001, 2002). The intrachromosomal rearrangements include the classical inverted duplications and deletions of 8p in which the interval between REPP and REPD remains intact (Fig. 4e,f, yellow), regions proximal to this interval are duplicated and inverted (Fig. 4e,f, blue), and the region distal to this interval is deleted (Fig. 4e,f, red). In this case, ectopic recombination leads to the formation of an unstable intermediate dicentric chromosome at meiosis I, which, depending where it breaks at anaphase I, results in a duplicated region that can extend from REPP as far as the centromere (Fig. 4e) or only as far, for example, band 8p21.1 (Fig. 4f). The pathology of such large imbalances is significant, with most patients having developmental delay, mental retardation, facial dysmorphisms, agenesis of the corpus callosum, and other problems including congenital heart disease (Guo et al. 1995). Despite the high frequency of inversion heterozygotes, the prevalence of the recurrent inverted, duplicated, and deleted 8p has been estimated at only one in 20,000; thus, in the absence of any evidence for a high rate of in utero fetal loss, the predisposition is relatively weak given the high frequency of the common inversion. In addition, ectopic recombination may not be the exclusive mechanism, as non-homologous end joining (NHEJ) has been invoked in a case in which neither parent carried the heterozygous inversion (Cooke et al. 2008).
The reciprocal of the classical inverted, duplicated, and deleted 8p is a supernumerary marker chromosome consisting of two copies of the region distal to REPD inverted with respect to each other (Fig. 4g). This duplicated marker chromosome would normally be lost but for the formation of a neocentromere at the site of the remaining REPP and REPD repeats at its center. In addition, REPP and REPD can interact with the more distal myomesin 2 (MYOM2) repeats in 8p23.3 to produce a chromosome that is duplicated for the interval between REPP and REPD (Fig. 4h, yellow) and triplicated for the interval between REPD and the MYOM2 repeat in 8p23.3 (Fig. 4h, red arrow) (Giorda et al. 2007).
REPD and REPP also mediate interchromosomal rearrangements such as the recurrent de novo unbalanced translocation between chromosomes 4 and 8 that is one of the causes of Wolf-Hirschhorn syndrome (Fig. 4i) (Giglio et al. 2002; Maas et al. 2007). Remarkably, the parent of origin is consistently maternal and doubly heterozygous for the common 8p23.1 inversion as well as another common inversion between olfactory repeat regions at ∼4 Mb and ∼9 Mb from the telomere of chromosome 4. The breakpoint in all the original cases investigated was at REPD on chromosome 8 and at either REPD or REPP on the short arm of chromosome 4 (Giglio et al. 2002).
Somatic rearrangements involving 8p23.1 and cancer
Recently, it has become clearer that the dicentric intermediate that leads to the classic inverted duplication and deletion of 8p [inv dup del(8)] may persist into the zygote, where subsequent early post-zygotic mitotic events can lead to mosaicism for the inv dup del(8) and a second cell line with other breakage products of the dicentric chromosome (Vermeesch et al. 2003; Pramparo et al. 2004). These include terminal deletions with breakpoints from 8p21.1 (Fig. 4j) (Vermeesch et al. 2003) to 8p11.2 (Pramparo et al. 2004). Interestingly, the cell lines with deletions and duplications can complement for each other, resulting in a milder phenotype than might otherwise be expected. It is probable that this is a more common mechanism than previously suspected and that mosaicism may rescue conceptions that would otherwise be lost owing to in utero selection against large imbalances.
Given the number of constitutional rearrangements mediated by REPP and REPD at meiosis, it might be expected that the same repeats would predispose to somatic recombination events. Until, recently, there was little evidence for this. The REPP-to-REPD interval contains the malignant fibrous histiocytoma amplified sequence 1 gene (MFHAS1, formerly MASL1 OMIM *605352), but the amplicon in this cancer has been only crudely mapped and may not be related to the 8p23.1 repeats (Sakabe et al. 1999). An 8p23.1-to-8p22 amplicon has also been reported in esophageal cancer, but the minimum size of this amplicon in two gastric tumors was 2.6 Mb between 10.1 and 12.7 Mb from the 8p telomere (Vauhkonen et al. 2007), and is therefore unlikely to have a breakpoint in either REPP or REPD. However, the breakpoints in 10 carcinoma cell lines have recently been mapped in detail, and “tumor break-prone segmental duplications” that correspond to REPP and REPD on 8p23.1 have been identified (Darai-Ramqvist et al. 2008). These, in turn, coincide with evolutionary breakpoints and carcinoma-related chromosome rearrangement hotspots in the Mitelman Database of Chromosome Aberrations in Cancer (http://cgap.nci.nih.gov/Chromosomes/Mitelman).
Thus, the beta-defensin and olfactory receptor repeats in 8p23.1 have established a paradigm in which paired repeats and a common polymorphism predispose to a collection of simple and complex chromosomal imbalances with significant clinical consequences. These repeats also masquerade as cytogenetic duplications when amplified, may nucleate neocentromeres, and could yet play a role in the formation of oncogenic rearrangements.
Bioinformatics and structural variation
Bioinformatics of structural variation will allow incorporation of that variation into the genomic context, and analysis of that variation will allow further hypotheses to be generated and tested. However, the main problem is that the data produced by current laboratory methods are generally too ambiguous and incomplete for use in bioinformatics pipelines. For example, even the very latest genome build, which is based on sophisticated computational assembly of quite deep sets of trace file sequencing reads, still contains many gaps and regions of uncertainty, and these correlate significantly with the presence of complex and unstable segmental duplications and CNVs (Redon et al. 2006). To improve on this situation and to locate shorter, rarer, and less similar CNVs, we and others are exploring more “tunable” tools for alignment, assembly, and visualization of primary trace file data. Including data from other laboratory methods, such as array-CGH and SNP genotyping, can also improve the map, but these methods are far from perfect as they only estimate the nucleotide extent and copy number range for structurally variable regions. As mentioned previously in the context of the beta-defensin region at 8p23.1 (Taudien et al. 2004), all of this challenges the concept of a single reference genome assembly, and it almost certainly means that genome browsers will need to present many alternative genomes as a reference set upon which all possible structural variations can be displayed. As an example, there are alternative assemblies for the HLA region presented in genome browsers, and this approach should be extended and deepened to cover structurally variable regions, and possibly the whole genome (Traherne et al. 2006).
We are therefore at a stage that is both exciting and frustrating, in that we know that CNVs, inversions, and the like are very common and important to genome function, but we are unable to draw a truly high-resolution map of these elements. This forces bioinformatics groups like the Database of Genomic Variants (DGV) (http://projects.tcag.ca/variation/) to make compromises. For example, the DGV currently annotates each reported structural variant over its largest possible genome extent, and then considers all CNVs that seem to overlap as alternative discoveries of the same structurally variable element (Zhang et al. 2006). But, of course, in many cases, these structural variants will be distinct and non-overlapping, or will overlap but have different genomic end-points and evolutionary origins.
The above uncertainties raise major problems for computational and analytical handling of structural variation data in disease studies, and this situation is made even worse by additional limitations of method accuracy and precision (as discussed above). The accuracy and precision of each method should be estimated, ideally empirically from real data, and such error rates incorporated into bioinformatics pipelines. At present, the error rates remain so high that the power of bioinformatics for combining and analyzing large amounts of data is diluted by the associated error with each one of those pieces of data, and inference of anything but large effects is difficult because of experimental noise. Given these complications, advanced bioinformatics could help solve the problem by enabling better assay design, thereby improving the ability of assays to count CNV copies and to distinguish between copies at distinct locations. But this solution cannot be relied on until methods can first fully define all the sequence versions and locations of CNVs, as input data for the assay design software to operate on. The error rates of current laboratory methods are the major limiting factor in the development of bioinformatics for structural variation.
Evolution of the dynamic defensins: Genome driven, biology driven, or both?
Highly duplicated regions, or regions with a high frequency of retrotransposons that share high sequence similarity, provide an ideal environment for high rates of ectopic recombination. Indeed, high amounts of structural variation are correlated with duplication-rich regions of the genome, and molecular studies on specific loci provide direct evidence of the importance of ectopic recombination in generating structural variation (Lam and Jeffreys 2006, 2007; Turner et al. 2008). But why does structural variation exist? Is it a consequence of genome architecture, which itself is a consequence of other evolutionary processes? Or is there a particular biological reason why certain genes in certain regions are structurally variable? A well-characterized 900-kb inversion on chromosome 17 has been shown to be under selection, with higher fecundity in heterozygous females, which may suggest that other inversions are selectively maintained as well (Stefansson et al. 2005). The distribution of CNVs in the genome can argue both for their being mostly neutral or candidates for selection (Cooper et al. 2007). The association of CNVs with segmental duplications and other repeat-rich regions suggests that CNV formation is a mechanistically driven consequence of a particular genomic architecture, and therefore consistent with a neutralist perspective. The alternative view is that CNVs are biased to certain functional classes of genes, often involved in environment sensing and response, which suggests that CNV formation is not random and subject to selection (Redon et al. 2006).
What role does selection have in maintaining or removing diversity in CNVs? Studies on CNV in Drosophila suggest that most CNVs are in mutation–selection balance, with purifying selection removing deleterious mutations generated by a high mutation rate as a consequence of the genomic architecture (Dopman and Hartl 2007; Emerson et al. 2008). In humans, population differences in CNV allele frequency such as those observed for the CCL3L1 and amylase loci are indications that positive selection may be operating. In the amylase study, increase in copy number has been linked to a phenotypic effect (increase in production of salivary amylase) and a potential selective mechanism for increase in copy number (increase in starch in diets of certain populations) (Perry et al. 2007). Nevertheless, it is extremely difficult to prove conclusively that selection is responsible without an estimation of mutation rate and mutational model, and a justifiable model for the mechanism of selection itself.
The beta-defensin locus provides some tentative clues as to the processes involved. The very low frequency (∼0.2%) of null alleles suggests that the homozygote nulls may be lethal, and therefore purifying selection may be acting against this tail of the distribution, which may be in mutation–selection balance. As with the amylase locus, a gene dosage effect provides phenotypic variation for selection to act on. The correlation of copy number with disease suggests putative selective mechanisms acting against both sides of the allelic distribution: a double-edged sword of susceptibility to infection of low copy number and an inappropriate inflammatory response of high copy number. We suggest that such a model may be applied to other immune CNV loci, such as the CCL3L1 locus. Further disease-association studies will either reinforce this model or refute it.
In addition to structural variation, sequence variation between paralogous repeats is an extra level of variation that may show signs of selection. Two paralogous genes may have two different selective pressures acting on them—the pressure to diverge in sequence and the pressure to maintain gene dosage by concerted evolution driven by gene conversion between the two paralogs. Most analyses have focused on duplicate genes assuming no CNV, analyzing distribution patterns of nucleotide variation in the Rh locus (Innan 2003b) and the CMT1A repeats (Lindsay et al. 2006), for example.
All structural variation should be treated in a similar manner to other variation: neutral unless proven otherwise. For sequence variation within copy number variation, standard tests of neutrality based on the neutral coalescent process, such as Tajima’s D, are not applicable, although model-independent tests of neutrality such as the McDonald-Kreitman test can be applied, and have been for the beta-defensin genes and others (Nguyen et al. 2006; Hollox and Armour 2008). Attempts at modeling a neutral coalescent process at duplicate genes have been made (Innan 2003a), but none has incorporated variable copy number into such a model. Similarly, to our knowledge, a statistical framework testing for evidence for selection on copy number alone has not been developed.
Summary: Lessons to learn
We have provided a synopsis of research into a particular structurally variable region in the human genome in order to attempt to extract meaningful conclusions that can be used to guide research at other loci. This region contains simple (DEFA1A3) and more complex (beta-defensin) multiallelic CNVs, as well as an inversion polymorphism. The variation is characterized at the sequence level for the alpha-defensins, but not the beta-defensins. In both cases, we emphasize the power that segregation analysis can give in resolving the allelic architecture of structural variation, and the importance of reliable diplotyping of large numbers of samples. The technology for reliable diplotyping is still at an early stage, and we must resist the temptation to conduct association studies based on poor typing technology, which could dilute the literature with false-positive results.
Such structural variation can affect phenotypic variation, including disease susceptibility, perhaps best illustrated by the association of high beta-defensin copy number with psoriasis. It can also act as a substrate for gross structural rearrangements that are likely to have significant clinical consequences. Given the importance of structural variation, accurate recording and databasing of variation data are a must, as this will aid not only disease association studies but studies on the population genetics and evolution of these regions. However, the development of appropriate bioinformatics frameworks is dependent on improvement in laboratory methods for measuring and describing structural variation.
More generally, there remain two related challenges in structural variation research that should be a research focus in the future. The first is relating the sequence variation within structurally variable regions to the structural variation itself—for example, how and by how much do copy number variable repeats differ from each other? This will give insights into the evolutionary origins, population dynamics, and disease relevance of such regions. The second area, which needs to be developed in concert with the first, is an accurate quantitative descriptive framework for summarizing these data—for example, how do we modify Nei’s nucleotide diversity statistic (Li 1987) for application to regions where there are not necessarily two copies per diploid genome?
As each individual structurally variable locus is characterized, common themes will be reinforced, but novel genomic findings and unusual architecture will undoubtedly be revealed. We suggest that such a locus-by-locus approach will provide important insights into disease and evolution, and a much clearer picture of a dynamic genome.
Footnotes
-
↵4 Corresponding author.
↵4 E-mail ejh33{at}leicester.ac.uk; fax 44-116-252-3378.
-
Article is online at http://www.genome.org/cgi/doi/10.1101/gr.080945.108.
- Copyright © 2008, Cold Spring Harbor Laboratory Press