Abstract
Allelic variation within the mouse androgen-binding protein (ABP) α subunit gene (Abpa) has been suggested to promote assortative mating and thus prezygotic isolation. This is consistent with the elevated evolutionary rates observed for the Abpa gene, and the Abpb and Abpg genes whose products (ABPβ and ABPγ) form heterodimers with ABPα. We have investigated the mouse sequence that contains the three Abpa/b/g genes, and orthologous regions in rat, human, and chimpanzee genomes. Our studies reveal extensive “remodeling” of this region: Duplication rates of Abpa-like and Abpbg-like genes in mouse are >2 orders of magnitude higher than the average rate for all mouse genes; synonymous nucleotide substitution rates are twofold higher; and the Abpabg genomic region has expanded nearly threefold since divergence of the rodents. During this time, one in six amino acid sites in ABPβγ-like proteins appear to have been subject to positive selection; these may constitute a site of interaction with receptors or ligands. Greater adaptive variation among Abpbg-like sequences than among Abpa-like sequences suggests that assortative mating preferences are more influenced by variation in Abpbg-like genes. We propose a role for ABPα/β/γ proteins as pheromones, or in modulating odorant detection. This would account for the extraordinary adaptive evolution of these genes, and surrounding genomic regions, in murid rodents.
Mammalian genome sequences are a boon for investigating evolutionary processes and their rates (Copley et al. 2003; Wolfe and Li 2003). Their analysis shows that >90% of genes that were present in the common ancestor of primates and rodents 65 to 110 million years ago are still present as single copies in the genome sequences of humans, mice, and rats (International Human Genome Sequencing Consortium [IHGSC] 2001; Mouse Genome Sequencing Consortium [MGSC] 2002; Rat Genome Sequencing Project Consortium [RGSPC] 2004). Those genes not preserved as single copies in both primate and rodent lineages have been duplicated or deleted or have formed pseudogenes. Conserved genes are likely to possess functions that are held in common by primates, rodents, and, in all likelihood, by most mammals. In contrast, frequently duplicated genes are likely to be associated with adaptation and functional innovation (Ohno 1970; Hughes 1999; Emes et al. 2003). Gene deletion and pseudogene formation events are rare, except among genes that have also been subject to duplication (MGSC 2002; RGSPC 2004).
Adaptive evolution can be inferred from estimating rates of nucleotide substitution between gene homolog pairs. Estimating the ratio of synonymous (“silent”) substitutions per synonymous site (KS) in protein-coding gene regions provides a useful metric for neutral evolution. When the ratio of nonsynonymous, amino-acid-changing substitutions per nonsynonymous site (KA) exceeds KS, then one or both of these genes are likely to have been subject to adaptive evolution (Hurst 2002). However, such cases are rare, because most amino acid sites are functionally conserved (Golding and Dean 1998). Detecting the effect of positive selection requires methods that account for variable selective pressures among sites. Prominent among these is the method described in Nielsen and Yang (1998) and Yang et al. (2000), which uses maximum likelihood and Bayes methods to predict the probabilities that sites have been subject to positive selection.
Nucleotide substitutions, in common with other mutations, are more likely to be fixed within a population when reproductive advantage is achieved in competitive situations (Hughes 1999). From genome-wide studies, it is clear that the majority of genes that have been subject to strong adaptive evolution participate in sexual reproduction, in immunity, and in chemosensation (MGSC 2002; RGSPC 2004). Elevated gene duplication rates and KA/KS ratios among genes thus might indicate their involvement in these functional categories.
Previously we noted both in the discussion of the mouse genome sequence (MGSC 2002), and elsewhere (Laukaitis et al. 2003), that mouse salivary androgen-binding protein (ABP) subunit genes are clustered together with numerous paralogous genes on Chromosome 7, and that these genes have no counterparts in humans. Thus, they appear to have proliferated rapidly in the time period since the last common ancestor of primates and rodents. ABP is a heterodimeric protein containing an α-subunit (Abpa), together with one of two sequence-similar subunits termed β or γ, encoded by genes Abpb and Abpg that are closely linked on Chromosome 7 (Dlouhy et al. 1987; Laukaitis et al. 2003). The KA/KS ratios for these three genes in Mus musculus subspecies and other species of Mus are considerably elevated (Hwang et al. 1997; Karn and Nachman 1999; Karn et al. 2002; MGSC 2002; Karn and Laukaitis 2003), suggesting that they have been subject to adaptive evolution and thus might function in reproduction, immunity, or chemosensation.
The ABP α, β, and γ subunits are representatives of the secretoglobin protein family. These are small, globular, secreted proteins whose phyletic distribution is currently restricted to the mammals and whose functions all remain elusive (Karn 1994; Klug et al. 2000; Reynolds et al. 2002; Laukaitis and Karn 2004). In laboratory experiments, female mice have demonstrated a significant mating preference for males with a comparable Abpa genotype (Laukaitis et al. 1997; Talley et al. 2001). As a consequence, and because of its rapid evolution and androgen-binding function (Dlouhy and Karn 1983; Karn and Nachman 1999; Karn et al. 2002), ABP has been proposed to act as a signal in a prezygotic isolation mechanism. Studies of wild mouse populations across a transect of the European hybrid zone in Bohemia partially support this idea. Mice captured in populations in the tails of the domesticus-musculus Abpa allele cline show a weak assortative preference in Y-maze studies, and this pattern of preference across the transect is consistent with incipient reinforcement (Bimova et al. 2004). It is not clear, however, that this effect is strong enough to contribute significantly to the maintenance of the zone. This is because Abpa shows a cline of introgression across a Danish transect of the European hybrid zone that is little different from the clines of markers thought not to be under significant selection (Dod et al. 2004).
The availability of human, mouse, and rat genome sequences provided us with an opportunity to investigate several issues that might clarify the evolution and function of these enigmatic molecules. We sought to understand whether Abpa, Abpb, and Abpg genes and their paralogs arose in the rodent lineage, particularly subsequent to the divergence of rat and mouse lineages, and whether primate genomes contain evidence of Abpa, Abpb, and Abpg genes. In addition, we were interested in whether adaptive evolution has acted preferentially at single sites within Abpa, Abpb, and Abpg gene sequences. Our comparative genomics findings demonstrate how strong adaptive evolution has remodeled a megabase genomic region, its genes, and their codons over a period of less than 12–24 million years.
RESULTS
Our first goal was to predict genes encoding secretoglobin family members, and paralogous pseudogenes, within an orthologous region of mouse, rat, and human genomes. This allowed us to compare coding and noncoding sequences. We were then able to address additional goals, including the reconstruction of evolutionary events, and the prediction of evolutionary rates for genes and codons.
Gene Predictions and Nomenclature
Using previously described mouse Abpa-like and Abpbg-like gene sequences as templates (Dlouhy et al. 1987; Karn and Laukaitis 2003), we identified nine and five Abpa-like genes and pseudogenes, respectively, and eight and five Abpbg-like genes and pseudogenes, respectively, in the mouse genome (Table 1; Supplemental material). Pseudogenes were predicted on the basis of in-frame stop codons or frame shifts; we recognize that pseudogenes with full-length open reading frames are misassigned as functional genes by this method.
Positions of Genes in the Abpa/b/g Orthologous Genomic Region Between Scn1b and Uble1b in the Three Species Mus musculus, Rattus norvegicus, and Homo sapiens
Gene name | Start | Finish | Stranda | Pseudogeneb | Median KA/KSc | Celera comparisond |
|---|---|---|---|---|---|---|
| Mouse Chromosome 7e | ||||||
| Scn1b | 22722387 | 22733263 | - | — | — | |
| Abpa1 | 22864062 | 22865250 | - | 1.261 | Cg_a.ac1, Identical | |
| Abpbg1 | 22872730 | 22874741 | + | 0.925 | Cg_bg.ac1, sense | |
| Abpa2 | 22899391 | 22900549 | - | 0.749 | Cg_a.ab1, Identical | |
| Abpbg2 | 22911541 | 22913573 | + | 0.513 | Cg_bg.ab2, Identical | |
| Abpbg12 | 22968351 | 22971550 | - | 0.894 | Cg_bg.ab1, Missense | |
| Abpbg3 | 23059588 | 23060737 | - | ψ | 1.103 | Cg_bg.aa1, Identical, ψ |
| Abpa3 | 23067244 | 23068432 | + | ψ | 0.795 | Cg_a.aa1, Identical, ψ |
| Abpa12 | 23102839 | 23104016 | + | 0.888 | Cg_a.ai1, Missense | |
| Abpbg4 | 23151387 | 23151716 | - | ψ | 1.158 | Cg_bg.ah1, Identical, ψ |
| Abpa4 | 23157659 | 23158824 | + | ψ | 1.188 | Cg_a.ah1, Missense, ψ |
| Abpa13 | 23225803 | 23226988 | + | 0.687 | Cg_a.aj2, Missense | |
| Abpbg13 | 23296589 | 23297381 | - | ψ | 0.855 | Cg_bg.an1, Identical, ψ |
| Abpbg5 | 23371211 | 23373004 | - | 0.867 | Cg_bg.am1, Identical | |
| Abpa5 | 23379940 | 23381122 | + | 1.131 | Cg_a.am1, Identical | |
| Abpbg6 | 23448140 | 23448924 | - | ψ | 1.052 | Cg_bg.a12, Missense, ψ |
| Abpa6 | 23457498 | 23458686 | + | ψ | 0.657 | Cg_a.al1, Missense, ψ |
| Abpbg7 | 23468193 | 23469980 | - | ψ | 0.828 | Cg_bg.al1, Identical, ψ |
| Abpa7 | 23478045 | 23479219 | + | ψ | 0.973 | Cg_a.a12, Identical, ψ |
| Abpbg8 | 23582205 | 23584143 | - | 0.530 | Cg_bg.ak1, Missense | |
| Abpa8 | 23588659 | 23589846 | + | 0.539 | Cg_a.ak1, Identical | |
| Abpbg9 | 23694727 | 23696519 | - | 1.156 | Cg_bg.aj2, Missense | |
| Abpa9 | 23708810 | 23709989 | + | ψ | 1.696 | Cg_a.aj1, Identical, ψ |
| Abpbg10 | 23784269 | 23786064 | - | 0.823 | Cg_bg.ar2, Identical | |
| Abpa10 | 23798099 | 23799277 | + | 0.876 | Cg_a.ar1, Identical | |
| Abpbg11 | 23852520 | 23854299 | - | 0.750 | Cg_bg.aq1, Identical | |
| Abpa11 | 23862002 | 23863183 | + | 0.989 | Cg_a.aq1, Missense | |
| Abpa14 | 23935733 | 23940927 | + | 1.122 | Cg_a.aq2, Identical | |
| Uble1b | 23981976 | 24009260 | - | — | — | |
| Singletons | Cg_a.ag_1, ψ | |||||
| Cg_a.ab_2 | ||||||
| Cg_a.af_1, ψ | ||||||
| Cg_bg.ag1 | ||||||
| Cg_bg.ae1, ψ | ||||||
| Cg_bg.af1, ψ | ||||||
| Cg_bg.aj1 | ||||||
| Rat Chromosome 1f | ||||||
| Scn1b | 86240364 | 86250239 | - | — | — | |
| Abpa1 | 86406893 | 86408040 | - | 1.075 | — | |
| Abpbg1 | 86413577 | 86415531 | + | 0.908 | — | |
| Abpbg2 | 86471441 | 86472350 | - | 0.778 | — | |
| Abpa2 | 86479128 | 86480275 | + | 1.075 | — | |
| Abpbg3 | 86615057 | 86616828 | - | ψ | 0.709 | — |
| Abpa3 | 86624816 | 86625956 | + | 1.018 | — | |
| Uble1b | 86684151 | 86711255 | - | — | — | |
| Human Chromosome 19g | ||||||
| UBA2 (Uble1b) | 39611152 | 39652635 | + | — | — | |
| SCGB4A1P | 39759479 | 39760446 | - | — | — | |
| SCGB4A2 | 39776303 | 39777308 | - | — | — | |
| SCGB4A3P | 39828407 | 39828797 | + | ψ | — | — |
| SCGB4A4 | 39848385 | 39849843 | - | — | — | |
| Abpbg1 | 39890294 | 39890500 | - | ψ | — | — |
| Abpa1 | 39893447 | 39893862 | + | ψ | — | — |
| Scn1b | 40213542 | 40223192 | + | — | — |
a Positions and orientation of transcription were determined using the BLAT server at UCSC (http://genome.cse.ucsc.edu).
b Pseudogenes were predicted on the basis of in-frame stop codons or frame shifts, and/or missing exons.
c Median pairwise KA/KS values were calculated using codeml (Yang and Nielsen 2000).
d Comparison of genes predicted from C57BL/6J and corresponding genomic DNA from mouse strains 129X1/SvJ, DBA/2J, and A/J from the Celera Genomics subscription database. Gene predictions were compared at the nucleotide level to those of C57B16/J; missense and sense mutations were predicted by comparing translated cDNA sequences.
e Mouse gene coordinates correspond to NCBI build 30 (USCS February 2003 mm3).
f Rat gene coordinates correspond to Baylor RGSC v3.1 (UCSC June 2003 rn3).
g Human gene coordinates correspond to NCBI build 34 (UCSC July 2003 hg16).
The genes for the ABP subunits α, β, and γ expressed in mouse submaxillary gland were originally designated Abpa, Abpb, and Abpg, respectively (Dlouhy et al. 1987). To describe these sequences, we have extended the original nomenclature by numbering the series of α-like and β/γ-like paralogs as Abpa1, Abpa2... Abpa14, and Abpbg1, Abpbg2... Abpbg13, respectively. Numbering is sequential according to the order in which paired genes (see below) are found in the mouse or rat genome. In this system, the original Abpa is referred to as Abpa11, Abpb as Abpbg11, and Abpg as Abpbg10, but we are not proposing to change the previous designations of Abpa, Abpb, and Abpg. In the appropriate places in this paper, we use both designations (e.g., Abpa11 [Abpa]) to facilitate reference to previous and future papers involving these three genes.
Genomic Arrangements of Abpa, Abpb, and Abpg Gene Homologs
Abpa- and Abpbg-like genes and pseudogenes were found to be ordered nonrandomly on the mouse genome (Fig. 1):
Of the 13 Abpa-like genes or pseudogenes, 11 were found immediately adjacent to an Abpbg-like gene or pseudogene.
These 11 Abpa-like/Abpbg-like pairs are oriented on opposing strands in a head-to-head (5′-to-5′; bidirectional) manner.
The pairs' coding regions are separated by only short sequences (median 7.6 kb).
The pairs are arranged in two clusters of Abpa-like/Abpbg-like gene pairs with opposite transcriptional orientations: two Abpa-like/Abpbg-like pairs, followed by nine inverted Abpbg-like/Abpa-like pairs.
Six, of a possible seven, pairs appear to contain both full-length genes, rather than containing pseudogenes.
A graphical representation of the relative position and transcriptional orientation of the Abpabg-like genes and pseudogenes located on Rattus norvegicus Chromosome 1, Mus musculus Chromosome 7, and Homo sapiens Chromosome 19. Coordinates are taken from genome releases rn3 (Baylor RGSC v3.1), mm3 (NCBI build 30), and hg16 (NCBI build 34), respectively. The 5′-to-3′ orientations of the genes are shown by the direction of the arrowheads. Scn1b and Uble1b genes, which lie in orthologous genomic regions in all three species, are numbered 1 and 2, respectively. Abpa-like genes are shown in blue, Abpbg-like genes in red, and primate SCGB4A1–4(P) genes in green. Filled arrowheads represent predicted functional genes whereas open arrowheads denote predicted pseudogenes. The sequence of duplication events among rodent genes inferred from phylogenetic trees (see text) is implied from the dendrogram, shown in black. Blue and red lines in this dendrogram represent branches that are not supported by the phylogenetic trees shown in Figure 3 (see text). The scale bar shows approximate genomic distance in megabases. Gaps (>5 kb) in the genomic assembly of each species are shown as black boxes.
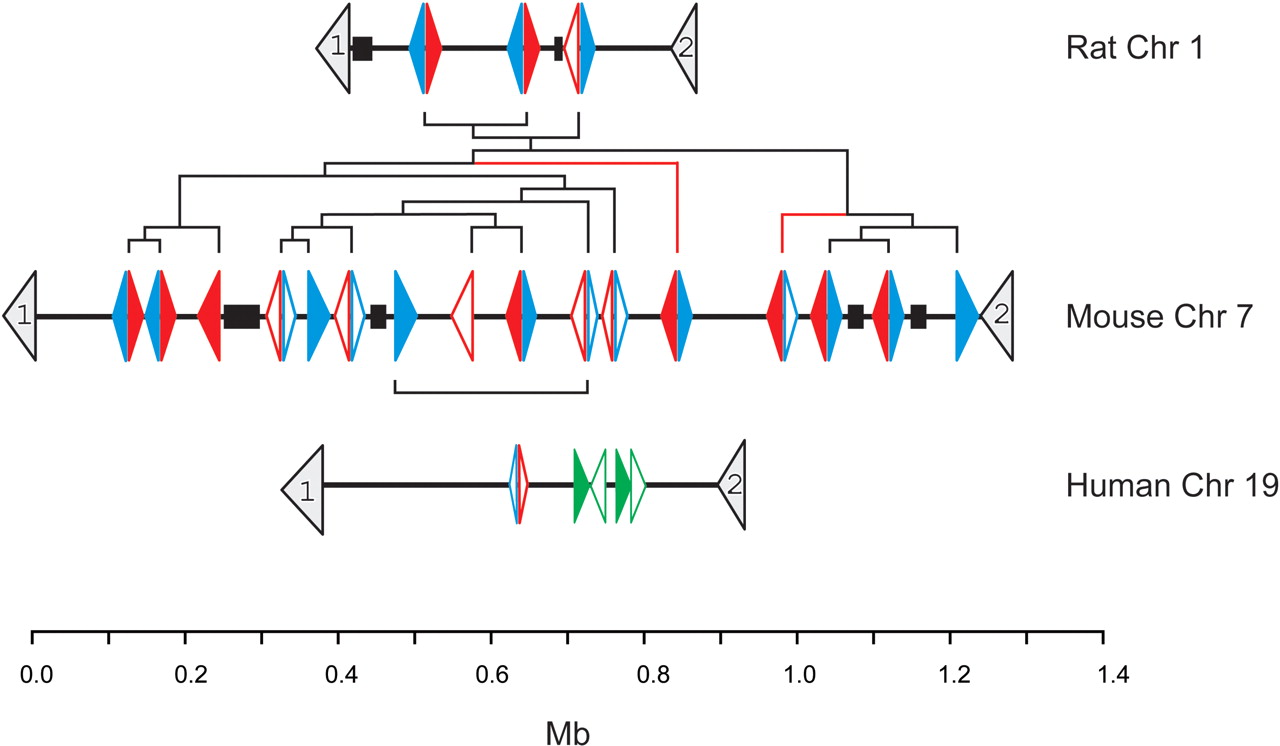
These observations immediately suggest that transcription, on complementary strands, of Abpa-like and Abpbg-like gene pairs may be coregulated using common, or else overlapping, regulatory elements (Trinklein et al. 2004), thereby facilitating coexpression, or antagonistic expression, of both Abpa-like and Abpbg-like genes. This is consistent with the head-to-head orientation of Abpa (Abpa11) and Abpb (Abpbg11) genes whose products function cooperatively in a heterodimer. Thus we expect that expression of each of the four remaining Abpa-like/Abpbg-like homolog pairs, including that which includes Abpg (Abpbg10) and Abpa10, are also coregulated. Indeed, Abpa2 and Abpbg2 represent paired genes that, according to GenBank entries AAB67069 and AAQ72534, are both expressed in the same tissue, the lacrimal gland.
Using identical methods, we were able to predict only three Abpa-like genes, two Abpbg-like genes, and one Abpbg-like pseudogene in the rat genome. Gaps in the rat genome assembly are too small to account for the greatly diminished number of Abpa and Abpbg genes and pseudogenes relative to mouse. Similar to the mouse, these six sequences are arranged in bidirectional Abpa-like/Abpbg-like pairs (Fig. 1).
Finally, as previously (MGSC 2002), we found no apparently functional Abpa or Abpbg genes in the human genome. However, single Abpa-like and Abpbg-like pseudogenes are present in the orthologous genomic region on human Chromosome 19 and chimpanzee Chromosome 20 (Fig. 1). The human and chimpanzee Abpa-like pseudogene contains a single stop codon (Fig. 2), and its 3′-exon could not be identified, and thus may have been deleted from the primate genome. Only the 5′-exon of the human and chimpanzee Abpbg-like pseudogene could be found (Fig. 2). As expected, this is in close proximity (2.9 kb) to the 5′ of the Abpa-like pseudogene, and retains the head-to-head orientation of the gene pairs seen in the two rodents (Fig. 1).
Amino acid sequence multiple alignment of rodent and primate ABPα and ABPβγ homologs, and primate secretoglobin homologs. Conceptual translations of both genes and pseudogenes (denoted by Ψ) are shown, with stop codons replaced by “X” symbols in white on black; codons containing a frame shift are also shown in white on black. Pseudo-exon sequences that could not be identified in genomic sequence are replaced by “.” characters; “-” represents a gap position. Multiple sequence alignments were produced by CLUSTAL W, manually adjusted, and colored using Chroma (Goodstadt and Ponting 2001) and an 80% consensus; gap positions were ignored in the calculation of a consensus sequence. Exons are shown as horizontal bars, and intron and exon boundaries were determined from alignments of the gene predictions and genomic DNA sequence using the UCSC genome browser. Protein secondary structure predicted by PHDSec (Rost and Sander 1993; PHD) shows four α-helices (H) in α-like chains, as has been demonstrated for cat Fel dI, rabbit uteroglobin, and human uteroglobin, and five α-helices in βγ-like chains. Codons predicted to be subject to positive selection by Codeml with a posterior probability of p > 0.9 in one model, and of at least p > 0.5 in one other model, are termed ω+ sites and are marked with an asterisk above the alignment. Positions of rabbit uteroglobin (UTG) residues predicted to interact with a bound ligand are each indicated by “X” above the aligned rabbit UTG sequence. (Mm) Mus musculus; (Hs) Homo sapiens; (Rn) Rattus norvegicus; (Pt) Pan troglodytes; (As) Apodemus sylvaticus; (Af) Apodemus flavicollis; (Aa) Apodemus agarius; and (Oc) Oryctolagus cuniculus.

The larger number of Abpa-like and Abpbg-like homologs in mouse, relative to rat and human, is also reflected in a larger size of its orthologous region (Fig. 1): mouse ∼1.3 Mb, rat ∼0.5 Mb, and human ∼0.6 Mb. (The large size of the human region, despite its single Abpa-like and Abpbg-like homologs, is likely due to the presence of multiple KRAB-box zinc finger genes that appear to have infiltrated this region during primate evolution.)
Evolution of Mouse and Rat Abpa-Like and Abpbg-Like Genes
Phylogenetic “5′ trees” were constructed from multiple alignments of nucleotide sequences that lie 5′ upstream to either Abpa-like, or Abpbg-like, sequences (see Methods). These extend far beyond the very short 5′-UTR sequences of these genes and their TATA boxes and transcriptional start sites (Fig. 3A; Laukaitis et al. 2003). Because nongenic regions are less encumbered by selection, as compared with genes (MGSC 2002), these sequences, on average, are more likely to have been subject to neutral, rather than selective, evolution. Phylogenetic trees constructed from them are thus more likely to recapitulate evolutionary events than are trees derived from coding sequences.
(A) Schematic representation of gene structures and repeat elements between the Abpa11 (Abpa) and Abpbg11 (Abpb) gene pair. The position and size of genes and repeat elements are shown to scale. Coordinates were obtained from the genome browser at UCSC (Kent 2002). The upper portion of the figure shows the genomic architecture of Abpa11 and Abpbg11 genes in greater detail. Highlighted regulatory elements correspond to those described previously (Laukaitis et al. 2003). The dashed lines represent the upstream regions used to generate the 5′ phylogenetic trees shown in B. (B) 5′ trees: phylogenetic relationships of rodent Abpa-like and Abpbg-like genes. Repeat-masked genomic DNA sequences 5′ upstream of Abpa-like genes and Abpbg-like genes were aligned (see Methods). For Abpa-like and Abpbg-like genes, 300 bp and 1 kb, respectively, 5′ to the translational start site were used for generation of the trees. Trees were generated using the neighbor joining method. The lineages containing the proposed roots of the trees are shown by black dots. Bootstrap values >50% are shown.
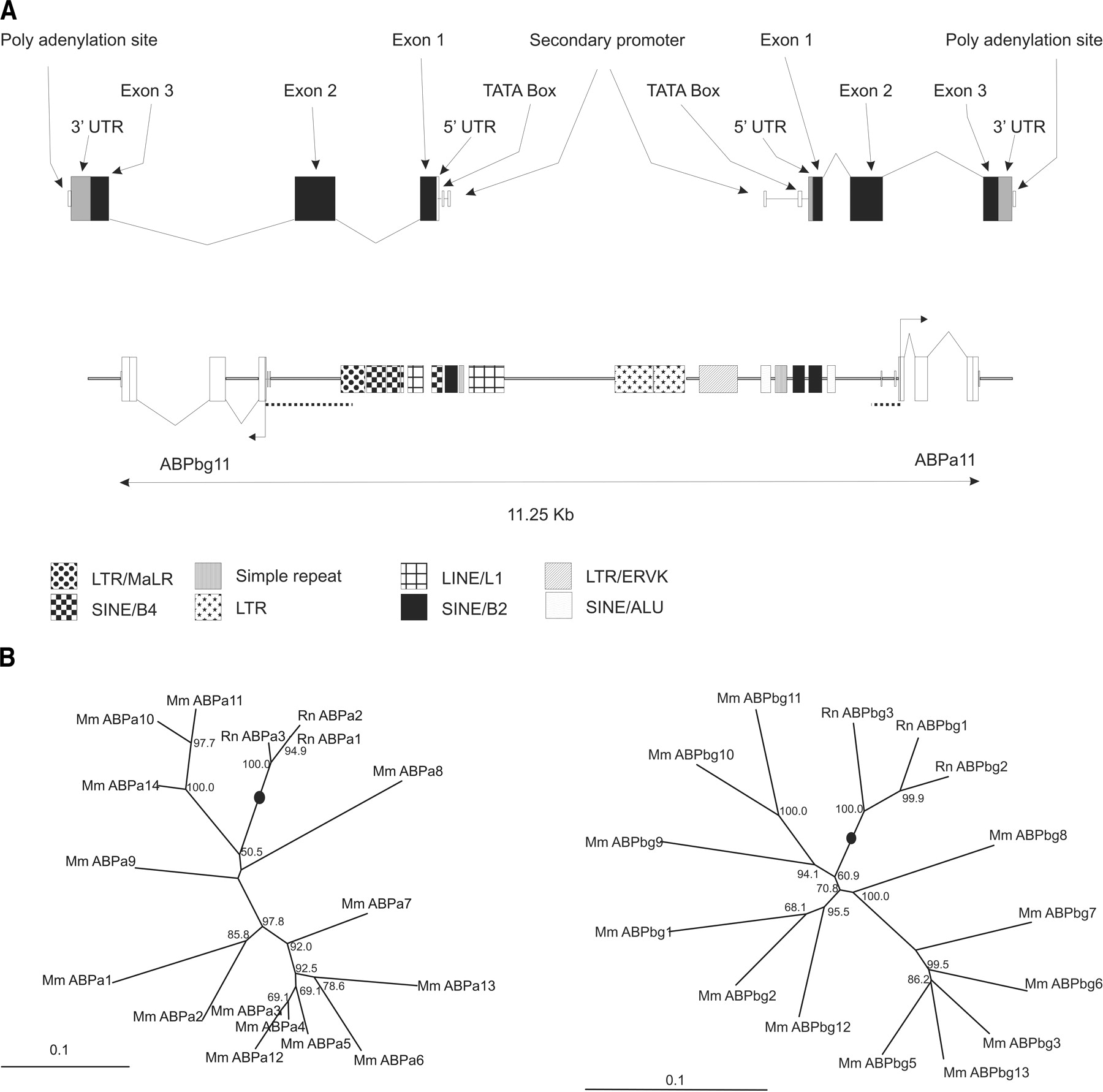
The 5′ trees provide detailed insights into the temporal sequence of gene duplications within these mammalian genomic regions. The most parsimonious explanation of the 5′ trees suggests the following sequence of events:
The common ancestor of primates and rodents possessed a single gene pair of sequences, one of which was Abpa-like and the other Abpbg-like. This can be inferred from the single (pseudogenic) versions of these genes in human and chimpanzee, and a prediction (see below) that the common ancestor of mouse and rat also only possessed a single gene pair.
The mouse Abpa-like and Abpbg-like gene repertoires arose independently from the rat gene repertoires. In the 5′ trees, rat Abpa-like or Abpbg-like sequences form monophyletic groups that exclude mouse sequences (Fig. 3B). Assuming no excision of genes from the rat lineage, this is most parsimoniously explained by single Abpa-like and Abpbg-like genes in the common ancestor of mouse and rat.
Each pair of Abpa-like and Abpbg-like genes arose from a single duplication event. Putting aside a single branch in each tree that lacks bootstrap support, the 5′ tree of Abpa-like genes and the 5′ tree of Abpbg-like sequences share a common topology (Fig. 3B), when genes paired together on the genomes are considered (see Fig. 1): the two trees share 48,386 out of 49,140 quartets (i.e., 98.5% similarity in quartet distance; Estabrook et al. 1985). Duplication of gene pairs as a single unit appears to have been the dominant mode of duplication in mouse and rat, although five unpaired “singleton” Abpa-like or Abpbg-like sequences are also present in the mouse genome. Some of these may yet be found to be paired when gaps in the genome assembly are filled (Fig. 1).
Duplication of a “parent” pair of Abpa- and Abpbg-like genes always resulted in two “daughter” pairs that are juxtaposed on the resulting genome. When the common topology of the phylogenetic trees (Fig. 3B) is superimposed on the gene orders along the two rodent genomes, this never results in a crossing-over of branches (Fig. 1). This implies that “daughter” gene pairs were never interspersed among other gene pairs, but were duplicated side-by-side.
Only a single inversion of mouse genes occurred in this region. Abpa1/Abpbg1 and Abpa2/Abpbg2 gene pairs are inverted with respect to all other pairs, yet these form a single monophyletic group. Thus either this single ancestral gene pair was subjected to an inversion event, or else the genomic region encompassing both extant pairs was inverted. (It is also possible that the inversion represents an artifact of the mouse genome assembly.)
Evolution of Other Rodent Abpa-Like and Abpbg-Like Genes
We also constructed trees from amino acid alignments, and from KS values, including Abpa-like sequences from species of Apodemus (Wickliffe et al. 2002), which are more closely related to the mouse than they are to the rat (Lundrigan et al. 2002). These trees all show that the Apodemus sequences cluster separately from both the mouse and rat clusters of Abpa-like sequences (data not shown). Thus, expansions of Abpa-like genes occurred independently for each of the three Mus, Apodemus, and Rattus genera.
Genome sequence data were also available from several mouse strains other than C57BL/6J (129X1/SvJ, DBA/2J, and A/J mouse strains; Mural et al. 2002). These data were investigated to determine whether Abpa-like and Abpbg-like genes are heterogeneous in number and sequence among different laboratory strains. Using identical methods, we identified each of the C57BL/6J 14 Abpa-like and 13 Abpbg-like (pseudo)genes in these other strains. In no case did we find a C57BL/6J gene to be a pseudogene in the other strains (and vice versa). These genes only differ by single-nucleotide substitutions in seven genes (six missense and one sense substitutions). The protein-coding sequences of the remaining 20 genes were identical at the nucleotide level in these strains. The six missense mutations are at sites distinct from predicted ω+ sites (see below).
One Abpa-like and two Abpbg-like genes (and two Abpa-like and two Abpbg-like pseudogenes) were additionally identified that are absent from the C57BL/6J genome assembly. However, these may yet be found to lie within gaps in the current C57BL/6J assembly (Fig. 1). Indeed, identical sequences to six of these seven genes or pseudogenes are present in unassembled C57BL/6J sequence (data not shown).
Primate Abpa-Like and Abpbg-Like Pseudogenes and Secretoglobin Evolution
5′ to the human Abpa- and Abpbg-like pseudogenes on human Chromosome 19, we identified three additional genes and one pseudogene; these lie within the orthologous Scn1b–Uble1b region (Fig. 1). These are secretoglobin homologs: for example, a search of the Pfam database (Bateman et al. 2004) with the conceptual translation of the 5′-most of these genes reveals significant sequence similarity to uteroglobins (E = 2.5 × 10–3). We name these genes SCGB4A1–4(P), following the proposals of the Secretoglobin Nomenclature Committee (Klug et al. 2000). SCGB4A1P and SCGB4A3P appear to be pseudogenes: the former's exon 1 has been translocated to lie 3′ to exons 2 and 3, whereas the latter contains an in-frame stop codon (Fig. 2). These four sequences appear to have arisen from two recent duplications because SCGB4A1P and SCGB4A4 are highly sequence-similar, as are SCGB4A2 and SCGB4A3P. Thus, it is possible that the sequence-dissimilar gene pair SCGB4A2 and SCGB4A4 encodes protein subunits of a heterodimer, similar to that seen for mouse ABPα and β or γ, and for cat Fel dI. All but SCGB4A1P are present in the current chimpanzee genome assembly (Fig. 1).
From EST data, it appears that SCGB4A2 is expressed in the eye and ovarian cancerous tissue (GenBank accessions BU738523, BM716941, AI073890, AI472323, AI821523, AI821558, BX105421, AA290868, AA481857, AA290985, and AA291047). Interestingly, SCGB4A1P, a likely pseudogene, has corresponding ESTs isolated from lung, multiple sclerosis lesions, infant brain, and in head and neck tissues (GenBank accessions BM984683, CD173361, N57568, T16687, and BE142511, respectively). In addition to these data, we assayed cDNAs prepared from a variety of human tissues. Only SCGB4A2 was successfully amplified and was isolated from pancreas and spleen cDNAs (data not shown).
Abpa-Like and Abpbg-Like Pseudogenes
The set of mouse Abpa, Abpb, and Abpg homologs contains a surprisingly high proportion (37%) of predicted pseudogenes. Pseudogene formation appears to have occurred following acquisition of frame shifts, exon deletions, or stop codons, or the substitution of critical residues such as an initiating methionine (in mouse Abpa9), or structurally important cysteine residues. In all 10 of these cases, disruptions to the open reading frames of mouse Abpa-like or Abpbg-like proteins appear to have occurred independently, because no two disruptions coincide in the alignment (Fig. 2). This implies that when duplication within the mouse genome occurred, it always involved functional Abpa-like and Abpbg-like genes, rather than pseudogenes. This would not have been expected if an unusually high duplication rate within this region was responsible for the elevated number of observed gene duplicates. Rather, it is likely that fixation of functional paired gene duplicates is the outcome of recurrent episodes of positive selection where selective advantage has accrued to individuals with relatively rare gene duplications.
Evolutionary Rates
Using codeml, we calculated the pairwise KA/KS values among mouse and rat Abpa-like or among Abpbg-like genes (Table 1; Supplemental Table 1). The medians of these values are unusually high: 0.99 for Abpa-like homologs, and 0.86 for Abpbg-like homologs. These values are similar to those calculated previously (Karn and Nachman 1999; Karn et al. 2002; MGSC 2002; Karn and Laukaitis 2003) and indicate that these genes have experienced relaxed selective constraints and/or adaptive evolution.
Median values of KS calculated by codeml for mouse Abpa-like and Abpbg-like pairwise comparisons were 0.27 and 0.53, respectively. These values are significantly higher than the median KS value (0.197) between mouse and rat ortholog pairs used in the initial analysis of the rat genome (RGSPC 2004). Only 16.4% and 0.3% of mouse–rat ortholog pairs are characterized by KS values >0.27 and >0.53, respectively.
Because we believe that these mouse genes arose by duplication after the mouse–rat divergence, the elevated neutral substitution rates imply that mouse Abpa-like and Abpbg-like genes have suffered an unusually high mutation rate relative to other genes. Similar conclusions were reached for rat Abpa-like and Abpbg-like genes (data not shown).
We were able to exclude the possibility that these KS increases are simply due to abnormal G+C content or values of κ, the ratio of transitions to transversions (data not shown). Moreover, these are not due to alignment inaccuracies because the low frequencies of insertion/deletion positions (Fig. 2) provide little margin of error in the alignment process. Instead, a combination of two evolutionary processes may have led to the elevation of these KS values. First, the genomic region in which the Abpa-like and Abpbg-like genes lie may have been unusually hypermutable, with inflated mutation rates relative to the rest of the genome. Second, substitutions at synonymous sites may have been elevated due to mutational correlation of adjacent bases (Bains 1992) such as methylated CG dinucleotides.
We attempted to quantitate the contributions of these two evolutionary processes by examining the neutral substitution rate within the nucleotide sequences lying 5′ upstream of either Abpa-like or Abpbg-like sequences; these may be assumed to be free from natural selection. The median number of substitutions per site between pairs of Abpa-like sequences was 0.251, and that for Abpbg-like sequences was 0.216; these calculations only compared a mouse with a rat sequence to provide nucleotide distances for orthologous sequences. These values are only slightly elevated when compared with an average of 0.174 substitutions per neutral site found in a rat–mouse genome-wide comparison (Fig. 5B, below; 0.083 + 0.091; RGSPC 2004). This appears to rule out hyper-mutability of this genomic region as an explanation of the significant elevations of KS values.
Site-specific KA/KS analysis of Abpa-like and Abpbg-like genes mapped to the tertiary structure of cat Fel dI. ω+ codons that are predicted to be under positive selection for mouse paralogs are mapped to a ribbon representation of the tertiary structure of feline major allergen Fel dI (PDB 1POU; Kaiser et al. 2003). (A) Ribbon diagram of the Fel dI dimer. Side chains of mapped predicted ω+ sites are highlighted in red. A single ligand molecule (2-methyl-2,4-pentanediol) present in the crystal structure is colored orange. (B) A representation of the molecular surface of the Fel dI chain 1 and 2 heterodimer. Mapped ABPα ω+ sites are shown in blue, and ABPβγ ω+ sites are shown in red. (C) Mapped ω+ sites predicted for the Abpa-like mouse paralogs. (D) Mapped ω+ sites predicted for the Abpbg-like mouse paralogs. (E) Prediction of the ABPβγ terminal extension and the mapped location of identified ω+ sites. The ABPβγ teminal extension is predicted to form a helical structure of ∼24 residues and is shown to scale. Swiss-PDBviewer (http://www.expasy.org/spdbv/; Guex et al. 1999) was used for all structural manipulations, and POVRAY (http://www.povray.org) was used to generate images.
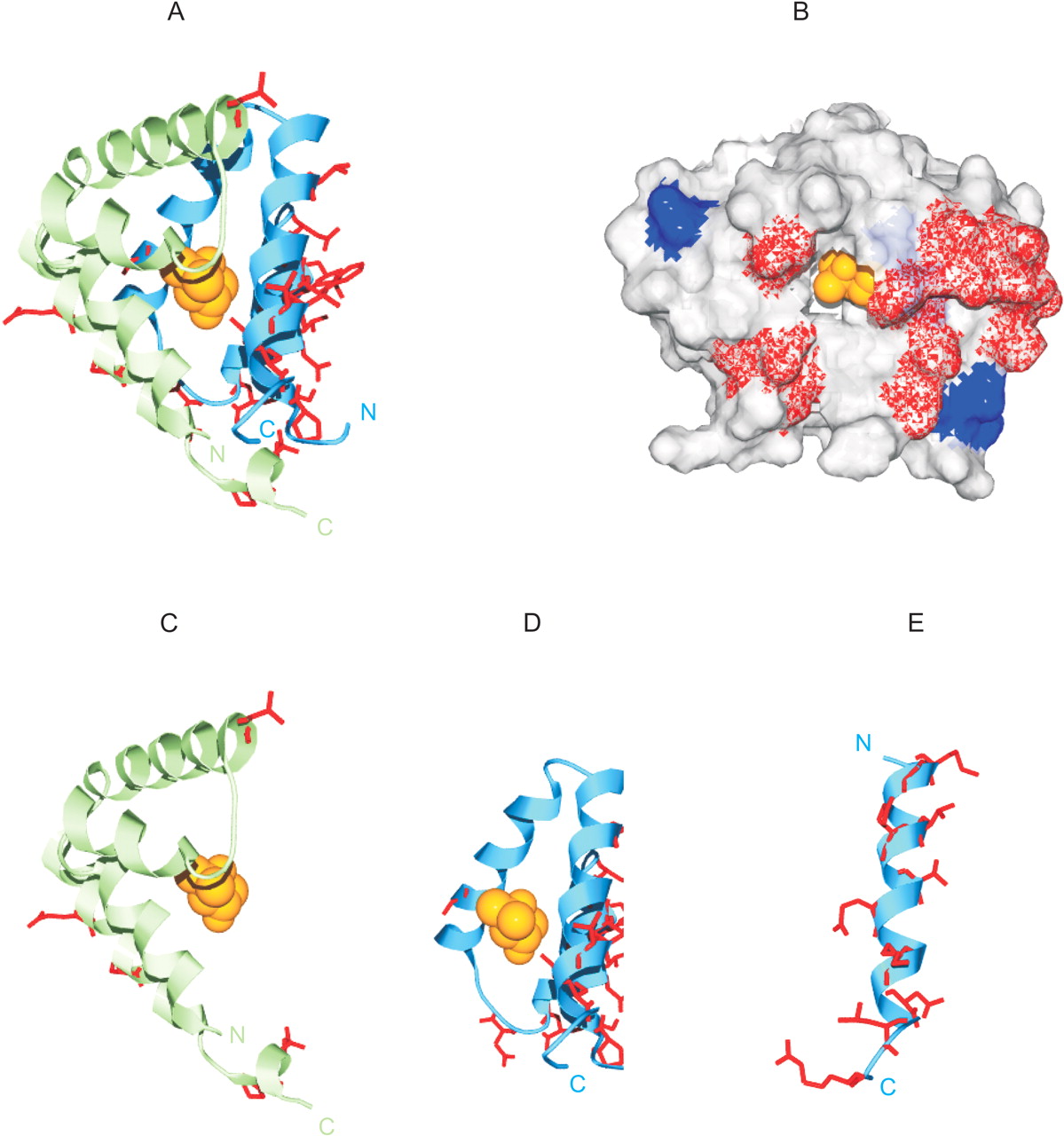
Concerted Evolution
We considered whether the apparent monophyly of mouse Abpa-like or Abpbg-like genes (Fig. 3B) might instead be caused by the effects of concerted evolution (Li 1997). On one hand, the unusually high KS values observed between mouse and rat orthologs argue against concerted evolution. This is because concerted evolution acts to homogenize sequences, and low, rather than high, KS values would thus be expected. Furthermore, concerted evolution usually is observed only in coding regions (e.g., Shibata and Yamazaki 1995), and the monophyly of mouse Abpa-like or Abpbg-like genes was inferred from 5′-flanking, rather than coding, sequences (Fig. 3). On the other hand, the equivalence of inactivating frameshift mutations in mouse Abpbg6 and rat Abpbg3 pseudogenes (Fig. 2) suggests that concerted evolution might be obscuring the orthology relationship between these two sequences. The true evolutionary history of these genes thus appears to have involved widespread gene duplication and coding sequence diversification, with perhaps more limited episodes of concerted evolution within coding sequence.
Secretoglobin Protein Structure
The detection of multiple Abpa-like and Abpbg-like genes in rodents yielded an opportunity to compare their sequences with the homologous chains of the cat allergen Fel dI (chains 1 and 2, respectively), whose protein tertiary structure has been determined (Kaiser et al. 2003). ABPα-like sequences are closely related to cat Fel dI chain 1, whereas ABPβγ-like sequences are closely related to Fel dI chain 2. Thus, the heterodimeric structure of ABPαβ and ABPαγ is recapitulated by the sequence-similar Fel dI chains 1 and 2. This conservation of primary and quaternary structure indicates that the genome of the eutherian common ancestor of cats, rodents, and primates contained a similar gene pair.
A multiple alignment of ABPαβγ-like conceptual amino acid sequences from mouse, rat, human, and chimpanzee genes (Fig. 2) demonstrates that these share the same four α-helix secondary structure that has been observed in structures of rabbit uteroglobin and cat Fel dI (Mornon et al. 1980; Kaiser et al. 2003). They also share the three cysteine residues that form single intrachain and interchain disulfide bonds in the Fel dI heterodimer. In addition, the Abpbg-like genes encode C-terminal extensions that are predicted to form a fifth α-helical structure, which is present in the ABPβγ-like Fel dI chain 2 but absent from other secretoglobins, such as Fel dI chain 1 and uteroglobin (Karn and Laukaitis 2003). These structural observations assisted the interpretation of evolutionary rate findings described below.
Positive Selection of Abpa-Like and Abpbg-Like Genes
By comparing the gene sequences of mouse and rat Abpa-like or Abpbg-like homologs, it is evident that their exons are changing more rapidly than are their introns and 3′-untranslated regions (UTRs; Fig. 4). This is a hallmark of positive selection, rather than neutral or purifying evolution. We compared the conservation of nucleotide sequence within exon 3 with that within 100 alignment positions 5′ upstream in intron 2, and 100 positions down-stream in the 3′-UTR for Abpbg-like homologs (Fig. 4); a similar comparison has been recently reported for Abpb and Abpg (Laukaitis et al. 2003). The intron and 3′-UTR sequences were highly conserved between mouse and rat homologs: 81.3% and 92.6% of sites, respectively, contain a nucleotide that is conserved in at least 70% of sequences. These are significantly higher levels than conservation levels in exon 3, where the corresponding figure is only 50.5% (Fig. 4).
Multiple nucleotide sequence alignment of mouse and rat Abpbg-like exons 3 and surrounding genomic DNA. Genomic DNA corresponding to exon 3 (98 positions) and 100 nucleotide positions of both flanking intronic and 3′-UTR sequence was aligned with HMMER, and manually adjusted. We found that 81.3%, 50.5%, and 92.6% of the sites in the intron, exon, and 3′-UTR, respectively, exhibited ≥70% consensus. In these calculations, positions with fewer than 50% gaps were considered. The 14 codons of exon 3 corresponding to predicted ω+ sites are shown by horizontal bars.

Next we used codeml to predict codons (“ω+ sites”) that have been subjected to positive selection. Five and 28 such ω+ sites were identified for mouse Abpa-like or Abpbg-like homologs, respectively (Fig. 2). These results demonstrate that adaptation has been the dominant evolutionary force during recent divergence of these rodent homologs. We mapped these ω+ sites to the crystal structure of the cat Fel dI heterodimer (Fig. 5). Although predicted ω+ sites appear to be relatively uniformly distributed along the protein sequence, a clear pattern is revealed when the sites are examined in relation to the predicted three-dimensional structure: they are overrepresented among exposed amino acids (Table 2), and they form a pronounced cluster on one face of the heterodimer (Fig. 5). Thus, the majority of these ω+ sites are available to participate in binding interactions. Moreover, they are entirely absent from the dimer's internal cavity (Fig. 5) and are distinct from sites in uteroglobin thought to bind internal ligands (Callebaut et al. 2000; Kaiser et al. 2003; Karn and Laukaitis 2003).
The Relative Solvent Accessibility of ABPαβγ ω+ Sites That Map to the Recombinant Cat Allergen Fel DI (PDB 1PUO)
1PUO chain 1 | ABPα | 1PUO chain 2 | ABPβγ | |||||
|---|---|---|---|---|---|---|---|---|
| All sites | Mouse and rat ω+ | Mouse only ω+ | All sites | Mouse and rat ω+ | Mouse only ω+ | |||
| No. of residues | 68 | 4 | 5 | 67 | 29 | 17 | ||
| No. buried | 5 | 0 | 0 | 6 | 3 | 1 | ||
| No. intermediate | 22 | 0 | 0 | 19 | 6 | 2 | ||
| No. exposed | 41 | 4 | 5 | 42 | 20 | 14 | ||
| Buried (%) | 7.4 | 0.0 | 0.0 | 9.0 | 10.3 | 5.9 | ||
| Intermediate (%) | 32.4 | 0.0 | 0.0 | 28.4 | 20.7 | 11.8 | ||
| Exposed (%) | 60.3 | 100.0 | 100.0 | 62.7 | 69.0 | 82.4 | ||
[i] The solvent accessibility values of cat Fel dl residues equivalent to ABPαβγ ω+ residues were calculated from the structure coordinates using the DSSP algorithm (Kabsch and Sander 1983). Relative accessibility scores were partitioned into three states: buried (<9% relative accessibility), intermediate (9%-35% relative accessibility), and exposed (≥36% relative accessibility), as described previously (Rost and Sander 1994).
Several secretoglobins are known to bind hydrophobic ligands within an interior cavity formed at the interface of two homologous chains (Beato and Baier 1975; Parker et al. 1982; Dlouhy and Karn 1983; Callebaut et al. 2000), and the cat Fel dI crystal structure contains strong electron density within this cavity, indicating the presence of a bound ligand of unknown type (Kaiser et al. 2003). Our results suggest that positive selection has not acted strongly on sites that confer specificity on binding internal ligands. Rather, it has constantly remodeled a single face of the heterodimer that consists of all but one of the five α-helices of ABPβγ-like homologs.
DISCUSSION
Our comparative genomics findings demonstrate that the mouse genomic sequence containing Abpa-like and Abpbg-like homologs has been subject to an exceptional degree of change over the relatively short interval of time (12–24 million years; Adkins et al. 2001; Springer et al. 2003) since the divergence of mouse and rat lineages. Rates of gene duplication, pseudogene formation, and synonymous and nonsynonymous nucleotide substitution are all significantly higher than for most other rodent genes. Our findings show that this genomic region has been subject to extensive and sustained incidents of positive selection.
Phylogenetic tree (Fig. 3B) and gene order (Fig. 1) data strongly indicate that the last common ancestor of rats and mice possessed only a single Abpa- and Abpbg-like gene pair. Thus the past 12–24 million years has experienced 14-fold and 13-fold expansions of the Abpa- and Abpbg-like mouse gene (and pseudogene) repertoires, respectively. Concomitantly, the mouse Scn1b–Uble1b orthology region has been enlarged several fold. Duplications are likely to have been of genes rather than pseudogenes, often with a subsequent loss of function and acquisition of reading frame disruptions and truncations. In rat, expansions have been more modest, resulting in only three genes and pseudogenes of each type.
We note that the rapidity of gene duplication and pseudogene formation inferred for these families is exceedingly rare. On average, only 3% of extant mouse genes arose from duplication since the last common ancestor with rat, and the rat and mouse genome sizes differ by only ∼6% (RGSPC 2004). Although the Scn1b–Uble1b regions in mouse and rat share a common evolutionary origin, they have been subject to a rare and extensive remodeling in relatively recent times.
Rapid evolutionary change was also observed within coding sequences. Per-gene median KA/KS values for Abpa-like and Abpbg-like genes are close to 1, which might indicate either neutral or adaptive evolution. In contrast, site-specific evolutionary rate estimates indicated the past action of positive selection among both Abpa-like and Abpbg-like genes: five and 28 codons, respectively, were identified as having experienced positive selection. The low number of such ω+ sites in Abpa-like, relative to Abpbg-like sequences may indicate that the former may include several nonfunctional pseudogenes, albeit with full-length open reading frames, that are evolving neutrally.
Loss of selective constraints on a gene eventually leads to pseudogene formation. Relatively few rodent genes have single pseudogenic counterparts in the human (C.P. Ponting and C. Webber, unpubl.). Those that do, such as genes for olfactory receptors and odorant-binding proteins (Emes et al. 2004), predominantly have functions in chemosensation, perhaps reflecting a diminution of the sense of smell in humans (Glusman et al. 2001). The rapid expansion of gene families independently in mouse and rat has also been observed for rodent pheromone genes (Emes et al. 2004). These analogies suggest that the Abpa-like and Abpbg-like genes of the primate/rodent common ancestor may have possessed chemosensation-related functions and that ABP paralogs may act as pheromones.
An intriguing picture of the functional regions of ABP is developing from this and other studies. ABP subunits are probably four- or five-helix bundles with the unusual boomerang shape of uteroglobin/clara proteins (Callebaut et al. 2000; Karn and Laukaitis 2003). As in the case of uteroglobin/clara, the subunits form a hydrophobic cavity at the interface of the two dimers (Callebaut et al. 2000) wherein a ligand having the A-ring structure of testosterone and progesterone (Karn 1998) can be bound. Secretoglobins have six conserved residues (residues F6, L13, Y21, F28, M41, and I63 in uteroglobin/clara) oriented toward the interior of the cavity. Of these, one (F28) functions in maintenance of the dimer interface, and the other five coordinate with the ligand. The alternative combinations of αβ and αγ subunits provide for sequence variation at three positions (corresponding to uteroglobin/clara L13, M42, and I63): this limited variation has been offered as the explanation of different binding affinities for testosterone and DHT (Karn and Nachman 1999; Karn and Laukaitis 2003). Overall, however, these internal residues are highly conserved, in contrast with the exterior residues we have identified as evolving under strong positive selection (Fig. 5).
The clustering of ω+ sites on one face of the ABPαβ/γ heterodimer implies that functional innovation at this interface has frequently conferred reproductive advantage to individual rodents. One possibility is that this represents the binding surface to an, as yet unknown, ABPαβ/γ receptor; their interaction contributes to assortative mating in mouse populations. If so, the greater adaptive variation among Abpbg-like, compared with Abpa-like, sequences suggests that assortative mating preferences are more influenced by allelic variation in ABPβγ-like than in ABPα-like molecules. This emerging picture is consistent with a protein that can act both as a molecule capable of communicating information on the basis of its own structure and as a carrier for a ligand that may participate in the information transfer.
Our comparative genomic studies have produced a picture of adaptive evolution shaping the number and sequences of Abpa and Abpbg subunit-like genes. The possibility of multiple combinations of subunits encoded in the highly duplicated region of Abp genes we describe here provides the potential for numerous signals in a variety of tissues. Future studies should aim to elucidate the breadth of expression of the many Abp paralogs in the tissues and secretions of M. musculus. This will undoubtedly help to provide clues to the function or functions of these paralogs and may help answer the broader question of the functional commonalities of secretoglobins in general.
METHODS
Gene Prediction and Annotation
A large-scale study of the mouse genome previously identified nine candidate Abpa-like genes lying in close proximity on Chromosome 7 (MGSC 2002; Emes et al. 2003). We used the amino acid conceptual translations of these candidate genes, and gene identification and gene building techniques, to identify orthologous genomic regions and homologous mouse, rat, and primate genes.
Prediction of Rodent Abp-Like Genes
The amino acid sequences of known Abpa, Abpb, and Abpg genes (Karn and Laukaitis 2003; Laukaitis et al. 2003) and predicted Abpa-like genes were used to search the genomes of M. musculus strain C57BL/6J (NCBI build 30, mm3), Rattus norvegicus (rn3), and Homo sapiens (NCBI build 34, hg16) using the BLAT server at UCSC (Kent 2002). These searches identified orthologous genomic segments in mouse, rat, and human that contain Abpa and Abpbg homologs. These segments are flanked by Scn1b and Uble1b genes in all three genomes. To obtain gene predictions, the genomic sequences that intervene between these flanking genes were used as templates for gene prediction (on both strands) using both Genewise (Birney and Durbin 2000) and hmmsearch (Eddy 1998), each of which uses hidden Markov models (HMMs; Eddy 1995, 1996). Genewise searches used HMMs constructed from amino acid multiple sequence alignments; hmmsearch used HMMs constructed from nucleotide multiple sequence alignments of known Abpa-like or Abpbg-like gene sequences extracted from the genome.
Prediction of Primate Secretoglobin Genes
In the human genome, the Scn1b–Uble1b intervening region contains a region aligned, by BLAT at the UCSC genome browser, to cat major allergen Fel dI mRNA (accession no. M77341), a known secretoglobin. This indicated that secretoglobin genes or pseudogenes are present within this orthologous genomic region. tBLASTn searches of human ESTs using the conceptual translations of cat Fel dI as a query were used to identify several homologous human ESTs that are mapped to the Scn1b–Uble1b orthologous segment. Predicted genes were mapped to the H. sapiens genome using BLAT. Introns and exons were predicted by requiring consistency between a multiple alignment of genomic sequences and transcript sequences mapped by BLAT to the genome; splice-site consensus sequences were required for genes. BLASTn searches of the November 2003 Pan troglodytes assembly obtained from Ensembl (http://www.ensembl.org/) were used to identify potential chimpanzee orthologs of these candidate human genes.
Nucleotide Sequence Multiple Alignments
Genomic DNAs corresponding to each of the Abpa-like and Abpbg-like genes were obtained from the genome browser at UCSC using positions obtained using BLAT. Among the Abpa-like genes, Abpa11 (Abpa) and Abpa10 exhibited greatest sequence similarity; their genomic DNA sequences were aligned using CLUSTAL W (Thompson et al. 1994). This alignment was used to generate an HMM using HMMer (Eddy 1998), which was then used to align all Abpa genomic DNA sequences. The same procedure was used to align the Abpbg-like genes using Abpbg10 (Abpg) and Abpbg11 (Abpb) genes for the starting alignment. The corresponding genomic sequences of 129X1/SvJ, DBA/2J, and A/J mouse strains from the Celera Genomics subscription database were also obtained and analyzed using these HMMs.
The positions of introns and exons were predicted using evidence from BLAT searches using the predicted protein sequences and also by maintaining conserved intron donor and acceptor splice sites (Zhang 1998).
Amino Acid Sequence Multiple Alignments
Conceptual amino acid sequences were translated from these gene predictions and aligned using CLUSTAL W (Thompson et al. 1994). Conceptual amino acid sequences of pseudogenes were prepared by applying frame shifts, where required, and ignoring any stop codons. All pairs of these gene sequences were extracted from the multiple alignments and their pairwise values of KA and KS calculated using codeml, an application from the PAML package (v1.31) of phylogenetic software (Yang 1997) with ambiguity positions ignored.
Predictions of Evolutionary Relationships
To overcome problems associated with building trees from rapidly evolving gene sequences, genomic DNA sequences 5′ to the translational start site of each gene were aligned. We assume that these DNA regions have evolved neutrally and thus trees (so-called 5′ trees) constructed from these alignments, may be used to accurately infer evolutionary relationships among Abp paralogs. To align the genomic DNA, the repeat elements present in the intervening genomic DNA of the Abpa and Abpbg gene pairs were first masked using the RepeatMasker Web server (http://ftp.genome.washington.edu/cgi-bin/RepeatMasker). The intervening sequences between Abpa11 and Abpbg11, or Abpa10 and Abpbg10, were then aligned by CLUSTAL W and manually edited to minimize gaps; HMMs were generated from these alignments. All repeat-masked genomic DNA sequences were then aligned using these HMMs and the program hmmalign (Eddy 1998), and the aligned DNA then edited to reduce gap positions.
The 5′ tree for Abpa-like genes used 300-base regions containing no repeat elements. Similarly, the 1-kb regions 5′ to the translational start site of Abpbg-like genes was used. Trees were generated using the neighbor joining (NJ) method (Saitou and Nei 1987), where distances correspond to percentage sequence divergence. Bootstrap values were calculated from 1000 trials with gap positions excluded; trees were visualized using Treeview (Page 1996). NJ trees were also generated from protein sequence alignments. Phylogenetic trees with contemporaneous tips based on KS values from codeml were generated using the Fitch-Margoliash algorithm (Fitch and Margoliash 1967) as modified from the “kitsch” program in Phylip (Felsenstein 1989) and 20 iterations. Topological comparisons of the phylogenetic 5′ trees for Abpa-like and Abpbg-like genes were performed by counting gene quartets common to both trees as a proportion of the total number of possible quartets (Estabrook et al. 1985); for this calculation, all paired genes' branches were considered. The Qdist program (Mailund and Pedersen 2004) was used for this purpose.
Nucleotide Substitution Levels in Noncoding Genomic DNA
The calculation of nucleotide distances between aligned 5′ genomic DNA sequences for mouse and rat Abpa-like and Abpbg-like genes was performed using baseml (Yang 1994). We used the parametric TN93 substitution model (Tamura and Nei 1993), which takes into account different rates of transversions and transitions. For both Abpa-like and Abpbg-like genes, the medians of all distances between mouse sequences and rat sequences were calculated as an indication of the substitution rates since the divergence of the mouse and rat lineages.
Site-Specific KA/KS Analysis
Site-specific KA/KS analyses were conducted using codeml (Yang et al. 2000). For each analysis, a cDNA sequence alignment was prepared that corresponded to its amino acid alignment. To ensure that the site-specific KA/KS analysis was not influenced by sites with a large number of gaps, all columns with gaps in >25% of the sequences were removed from the cDNA alignment. Although the 5′ trees were used as input files for codeml to generate results described in this report, other tree topologies, such as those generated from KS estimates, gave essentially identical results.
Codeml uses maximum likelihood to predict sites in a group of cDNA sequences that have been subject to positive selection (Zanotto et al. 1999; Bishop et al. 2000; Yang and Bielawski 2000; Yang et al. 2000; Peek et al. 2001; Schaner et al. 2001; Jansa et al. 2003). Log likelihood values (l) are calculated for each model by maximum likelihood allowing the comparison by a Likelihood Ratio Test (LRT) between (1) simple models where sites are predicted to have a KA/KS or ω ratio between 0 and 1, and (2) more complex models that also allow for ratios that are >1. If the complex model indicates an estimated ω ratio that is >1, and the test statistic (2Δl) is greater than critical values of the Chi square (χ2) distribution with the appropriate degree of freedom (Yang et al. 1998), then positive selection can be inferred. Bayesian probabilities are used to predict which sites (codons) from the original data have most likely been subjected to positive selection.
We used three pairs of simple and complex models: M0 (one-ratio; Goldman and Yang 1994) versus M3 (discrete; Yang et al. 2000); M1 (neutral) versus M2 (selection; Nielsen and Yang 1998); and M7 (β) versus M8 (β + ω; Yang et al. 2000). As described previously (Emes et al. 2004), only nonconserved alignment positions predicted to be under positive selection with a posterior probability >0.90 by one codeml model, and >0.50 by at least one other model, were highlighted. We have termed these alignment positions “ω+ sites.” ω+ sites were mapped to the crystal structure of the major cat allergen Fel dI (PDB 1PUO; Kaiser et al. 2003), an engineered heterodimer of two secretoglobin molecules. Fel dI amino acid sequences were aligned to the family alignment using HMMer, and manually adjusted to minimize insertions and deletions within secondary structure elements. Swiss-PDBviewer (http://www.expasy.org/spdbv/; Guex et al. 1999) was used for structural manipulations, and POVRAY (http://www.povray.org) was used to render images.
Solvent accessibility values of Fel dI amino acids that align to ABPα, β, and γ ω+ residues were calculated using structural information from PDB (http://www.rcsb.org/pdb/) and the DSSP algorithm (Kabsch and Sander 1983); ligands were removed prior to accessibility score estimation. Relative accessibility scores were produced by normalizing these scores by amino-acid-specific maximal accessibility values. These relative scores were partitioned into three states: buried (<9% relative accessibility), intermediate (9%–35% relative accessibility), and exposed (≥36% relative accessibility), as previously (Rost and Sander 1994).
Primate-Specific Secretoglobin Gene Expression
The expression profiles of human secretoglobin homologs were obtained using the EST table from the genome browser at UCSC (http://genome.cse.ucsc.edu/). To obtain further expression information, gene-specific primers were designed and used to assay a human multiple tissue cDNA panel (Clontech multiple tissue cDNA panels 1 and 2, product numbers K1420-1 and K1421-1) by PCR. Gene-specific primers for each of the genes were used to amplify 1 μL of a 2 × diluted stock of each cDNA under the following cycle conditions: 30 sec at 95°C; five cycles of 3 min at 72°C; 30 sec at 95°C; five cycles of 3 min at 70°C; 30 sec at 95°C; and 25 cycles of 3 min at 68°C. Amplified products were separated by electrophoresis in 2% agarose gels and purified (Minelute, gel extraction; QIAGEN). Then 1 μL of eluted cDNA was ligated to PST-1blue vector and sequenced on both strands using vector primers.
Notes
[10] Article and publication are at http://www.genome.org/cgi/doi/10.1101/gr.2540304. Article published online ahead of print in July 2004.
[11] Present address: The Wellcome Trust Sanger Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge CB10 1SA, United Kingdom.
[12] Corresponding author. E-MAIL [email protected]; FAX 44 (0)1865 282651.
We thank Peter Oliver and Ponting group members for helpful discussions. This work was funded by the UK Medical Research Council, and by the Holcomb Research Institute, Butler Univer-sity. M.C.R. was supported for his work at Oxford University by a Seitz Scholarship, Butler University. That support is gratefully acknowledged.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
References
- ↵Adkins, R.M., Gelke, E.L., Rowe, D., and Honeycutt, R.L. 2001. Molecular phylogeny and divergence time estimates for major rodent groups: Evidence from multiple genes. Mol. Biol. Evol. 18: 777–791.
- ↵Bains, W. 1992. Local sequence dependence of rate of base replacement in mammals. Mutat. Res. 267: 43–54.
- ↵Bateman, A., Coin, L., Durbin, R., Finn, R.D., Hollich, V., Griffiths-Jones, S., Khanna, A., Marshall, M., Moxon, S., Sonnhammer, E.L., et al. 2004. The Pfam protein families database. Nucleic Acids Res. 32: D138–D141.
- ↵Beato, M. and Baier, R. 1975. Binding of progesterone to the proteins of the uterine luminal fluid. Identification of uteroglobin as the binding protein. Biochim. Biophys. Acta 392: 346–354.
- ↵Bimova, B., Karn, R.C., and Pialek, J. 2004. The role of salivary androgen-binding protein in reproductive isolation between two subspecies of house mouse: Mus musculus musculus and Mus musculus domesticus. Biol. J. Linn. Soc. (in press).
- ↵Birney, E. and Durbin, R. 2000. Using GeneWise in the Drosophila annotation experiment. Genome Res. 10: 547–548.
- ↵Bishop, J.G., Dean, A.M., and Mitchell-Olds, T. 2000. Rapid evolution in plant chitinases: Molecular targets of selection in plant–pathogen coevolution. Proc. Natl. Acad. Sci. 97: 5322–5327.
- ↵Callebaut, I., Poupon, A., Bally, R., Demaret, J.P., Housset, D., Delettre, J., Hossenlopp, P., and Mornon, J.P. 2000. The uteroglobin fold. Ann. NY Acad. Sci. 923: 90–112.
- ↵Copley, R.R., Goodstadt, L., and Ponting, C.P. 2003. Eukaryotic domain evolution inferred from genome comparisons. Curr. Opin. Genet. Dev. 13: 623–628.
- ↵Dlouhy, S.R. and Karn, R.C. 1983. The tissue source and cellular control of the apparent size of androgen binding protein (Abp), a mouse salivary protein whose electrophoretic mobility is under the control of sex-limited saliva pattern (Ssp). Biochem. Genet. 21: 1057–1070.
- ↵Dlouhy, S.R., Taylor, B.A., and Karn, R.C. 1987. The genes for mouse salivary androgen-binding protein (ABP) subunits α and γ are located on Chromosome 7. Genetics 115: 535–543.
- ↵Dod, B., Smadja, C., Karn, R.C., and Boursot, P. 2004. Testing for selection on the androgen-binding protein in the Danish mouse hybrid zone. Biol. J. Linn. Soc. (in press).
- ↵Eddy, S.R. 1995. Multiple alignment using hidden Markov models. Proc. Int. Conf. Intell. Syst. Mol. Biol. 3: 114–120.
- ↵Eddy, S.R. 1996. Hidden Markov models. Curr. Opin. Struct. Biol. 6: 361–365.
- ↵Eddy, S.R. 1998. Profile hidden Markov models. Bioinformatics 14: 755–763.
- ↵Emes, R.D., Goodstadt, L., Winter, E.E., and Ponting, C.P. 2003. Comparison of the genomes of human and mouse lays the foundation of genome zoology. Hum. Mol. Genet. 12: 701–709.
- ↵Emes, R.D., Beatson, S.A., Ponting, C.P., and Goodstadt, L. 2004. Evolution and comparative genomics of odorant- and pheromone-associated genes in rodents. Genome Res. 14: 591–602.
- ↵Estabrook, G.F., McMorris, F.R., and Meacham, C.A. 1985. Comparison of undirected phylogenetic trees based on subtrees of four evolutionary units. System. Zool. 34: 193–200.
- ↵Felsenstein, J. 1989. PHYLIP—Phylogeny inference package. Cladistics 5: 164–166.
- ↵Fitch, W.M. and Margoliash, E. 1967. Construction of phylogenetic trees. Science 155: 279–284.
- ↵Glusman, G., Yanai, I., Rubin, I., and Lancet, D. 2001. The complete human olfactory subgenome. Genome Res. 11: 685–702.
- ↵Golding, G.B. and Dean, A.M. 1998. The structural basis of molecular adaptation. Mol. Biol. Evol. 15: 355–369.
- ↵Goldman, N. and Yang, Z. 1994. A codon-based model of nucleotide substitution for protein-coding DNA sequences. Mol. Biol. Evol. 11: 725–736.
- ↵Goodstadt, L. and Ponting, C.P. 2001. CHROMA: Consensus-based colouring of multiple alignments for publication. Bioinformatics 17: 845–846.
- ↵Guex, N., Diemand, A., and Peitsch, M.C. 1999. Protein modelling for all. Trends Biochem. Sci. 24: 364–367.
- ↵Hughes, A.L. 1999. Adaptive evolution of genes and genomes. Oxford University Press, New York.
- ↵Hurst, L.D. 2002. The KA/KS ratio: Diagnosing the form of sequence evolution. Trends Genet. 18: 486.
- ↵Hwang, J.M., Hofstetter, J.R., Bonhomme, F., and Karn, R.C. 1997. The microevolution of mouse salivary androgen-binding protein (ABP) paralleled subspeciation of Mus musculus. J. Hered. 88: 93–97.
- ↵International Human Genome Sequencing Consortium (IHGSC). 2001. Initial sequencing and analysis of the human genome. Nature 409: 860–921.
- ↵Jansa, S.A., Lundrigan, B.L., and Tucker, P.K. 2003. Tests for positive selection on immune and reproductive genes in closely related species of the murine genus Mus. J. Mol. Evol. 56: 294–307.
- ↵Kabsch, W. and Sander, C. 1983. Dictionary of protein secondary structure: Pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22: 2577–2637.
- ↵Kaiser, L., Gronlund, H., Sandalova, T., Ljunggren, H.G., van Hage-Hamsten, M., Achour, A., and Schneider, G. 2003. The crystal structure of the major cat allergen Fel dI, a member of the secretoglobin family. J. Biol. Chem. 278: 37730–37735.
- ↵Karn, R. 1994. The mouse salivary androgen-binding protein (ABP) α subunit closely resembles chain 1 of the cat allergen Fel dI. Biochem. Genet. 32: 271–277.
- ↵Karn, R. 1998. Steroid binding by mouse salivary proteins. Biochem. Genet. 36: 105–117.
- ↵Karn, R.C. and Laukaitis, C.M. 2003. Characterization of two forms of mouse salivary androgen-binding protein (ABP): Implications for evolutionary relationships and ligand-binding function. Biochemistry 42: 7162–7170.
- ↵Karn, R.C. and Nachman, M.W. 1999. Reduced nucleotide variability at an androgen-binding protein locus (Abpa) in house mice: Evidence for positive natural selection. Mol. Biol. Evol. 16: 1192–1197.
- ↵Karn, R.C., Orth, A., Bonhomme, F., and Boursot, P. 2002. The complex history of a gene proposed to participate in a sexual isolation mechanism in house mice. Mol. Biol. Evol. 19: 462–471.
- ↵Kent, W.J. 2002. BLAT—The BLAST-like alignment tool. Genome Res. 12: 656–664.
- ↵Klug, J., Beier, H.M., Bernard, A., Chilton, B.S., Fleming, T.P., Lehrer, R.I., Miele, L., Pattabiraman, N., and Singh, G. 2000. Uteroglobin/Clara cell 10-kDa family of proteins: Nomenclature committee report. Ann. NY Acad. Sci. 923: 348–354.
- ↵Laukaitis, C.M. and Karn, R.C. 2004. Evolution of the secretoglobins: A genomic and proteomic view. Biol. J. Linn. Soc. (in press).
- ↵Laukaitis, C.M., Critser, E.S., and Karn, R.C. 1997. Salivary androgen-binding protein (ABP) mediates sexual isolation in Mus musculus. Evolution 51: 2000–2005.
- ↵Laukaitis, C.M., Dlouhy, S.R., and Karn, R.C. 2003. The mouse salivary androgen-binding protein (ABP) gene cluster on Chromosome 7: Characterization and evolutionary relationships. Mamm. Genome 14: 679–691.
- ↵Li, W.-H. 1997. Molecular Evolution. Sinauer Associates, Sunderland, MA.
- ↵Lundrigan, B.L., Jansa, S.A., and Tucker, P.K. 2002. Phylogenetic relationships in the genus Mus, based on paternally, maternally, and biparentally inherited characters. Syst. Biol. 51: 410–431.
- ↵Mailund, T. and Pedersen, C.N. 2004. QDist—Quartet distance between evolutionary trees. Bioinformatics Feb 12 [Epub ahead of print].
- ↵Mornon, J.P., Fridlansky, F., Bally, R., and Milgrom, E. 1980. X-Ray crystallographic analysis of a progesterone-binding protein. The C222(1) crystal form of oxidized uteroglobin at 2–2 Å resolution. J. Mol. Biol. 137: 415–422.
- ↵Mouse Genome Sequencing Consortium (MGSC). 2002. Initial sequencing and comparative analysis of the mouse genome. Nature 420: 520–562.
- ↵Mural, R.J., Adams, M.D., Myers, E.W., Smith, H.O., Miklos, G.L., Wides, R., Halpern, A., Li, P.W., Sutton, G.G., Nadeau, J., et al. 2002. A comparison of whole-genome shotgun-derived mouse chromosome 16 and the human genome. Science 296: 1661–1671.
- ↵Nielsen, R. and Yang, Z. 1998. Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 148: 929–936.
- ↵Ohno, S. 1970. Evolution by gene duplication. Springer-Verlag, New York.
- ↵Page, R.D. 1996. TreeView: An application to display phylogenetic trees on personal computers. Comput. Appl. Biosci. 12: 357–358.
- ↵Parker, M., Needham, M., and White, R. 1982. Prostatic steroid binding protein: Gene duplication and steroid binding. Nature 298: 92–94.
- ↵Peek, A.S., Souza, V., Eguiarte, L.E., and Gaut, B.S. 2001. The interaction of protein structure, selection, and recombination on the evolution of the type-1 fimbrial major subunit (fimA) from Escherichia coli. J. Mol. Evol. 52: 193–204.
- ↵Rat Genome Sequencing Project Consortium (RGSPC). 2004. Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature 428: 493–521.
- ↵Reynolds, S.D., Reynolds, P.R., Pryhuber, G.S., Finder, J.D., and Stripp, B.R. 2002. Secretoglobins SCGB3A1 and SCGB3A2 define secretory cell subsets in mouse and human airways. Am. J. Respir. Crit. Care Med. 166: 1498–1509.
- ↵Rost, B. and Sander, C. 1993. Prediction of protein secondary structure at better than 70% accuracy. J. Mol. Biol. 232: 584–599.
- ↵Rost, B. and Sander, C. 1994. Conservation and prediction of solvent accessibility in protein families. Proteins 20: 216–226.
- ↵Saitou, N. and Nei, M. 1987. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4: 406–425.
- ↵Schaner, P., Richards, N., Wadhwa, A., Aksentijevich, I., Kastner, D., Tucker, P., and Gumucio, D. 2001. Episodic evolution of pyrin in primates: Human mutations recapitulate ancestral amino acid states. Nat. Genet. 27: 318–321.
- ↵Shibata, H. and Yamazaki, T. 1995. Molecular evolution of the duplicated Amy locus in the Drosophila melanogaster species subgroup: Concerted evolution only in the coding region and an excess of nonsynonymous substitutions in speciation. Genetics. 141: 223–236.
- ↵Springer, M.S., Murphy, W.J., Eizirik, E., and O'Brien, S.J. 2003. Placental mammal diversification and the Cretaceous–Tertiary boundary. Proc. Natl. Acad. Sci. 100: 1056–1061.
- ↵Talley, H., Laukaitis, C., and Karn, R. 2001. Female preference for male saliva: Implications for sexual isolation of Mus musculus subspecies. Evolution 55: 631–634.
- ↵Tamura, K. and Nei, M. 1993. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol. 10: 512–526.
- ↵Thompson, J.D., Higgins, D.G., and Gibson, T.J. 1994. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22: 4673–4680.
- ↵Trinklein, N.D., Aldred, S.F., Hartman, S.J., Schroeder, D.I., Otillar, R.P., and Myers, R.M. 2004. An abundance of bidirectional promoters in the human genome. Genome Res. 14: 62–66.
- ↵Wickliffe, J., Lee, V., Smith, E., Tandler, B., and Phillips, C. 2002. Gene expression, cell localization, and evolution of rodent submandibular gland androgen-binding protein. Eur. J. Morphology 40: 257–260.
- ↵Wolfe, K.H. and Li, W.H. 2003. Molecular evolution meets the genomics revolution. Nat. Genet. 33: 255–265.
- ↵Yang, Z. 1994. Estimating the pattern of nucleotide substitution. J. Mol. Evol. 39: 105–111.
- ↵Yang, Z. 1997. PAML: A program package for phylogenetic analysis by maximum likelihood. Comput. Appl. Biosci. 13: 555–556.
- ↵Yang, Z. and Bielawski, J.P. 2000. Statistical methods for detecting molecular adaptation. Trends Ecol. Evol. 15: 496–503.
- ↵Yang, Z. and Nielsen, R. 2000. Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol. Biol. Evol. 17: 32–43.
- ↵Yang, Z., Nielsen, R., and Hasegawa, M. 1998. Models of amino acid substitution and applications to mitochondrial protein evolution. Mol. Biol. Evol. 15: 1600–1611.
- ↵Yang, Z., Nielsen, R., Goldman, N., and Pedersen, A.M. 2000. Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics 155: 431–449.
- ↵Zanotto, P.M., Kallas, E.G., de Souza, R.F., and Holmes, E.C. 1999. Genealogical evidence for positive selection in the nef gene of HIV-1. Genetics 153: 1077–1089.
- ↵Zhang, M.Q. 1998. Statistical features of human exons and their flanking regions. Hum. Mol. Genet. 7: 919–932.
WEB SITE REFERENCES
- http://genome.cse.ucsc.edu/; UCSC genome browser.
- http://www.ensembl.org/; Ensembl genome browser.
- http://www.expasy.org/spdbv/; Swiss-PDBviewer.
- http://www.povray.org/; POVRAY, graphical representation programs.
- http://www.rcsb.org/pdb/; the protein data bank.

