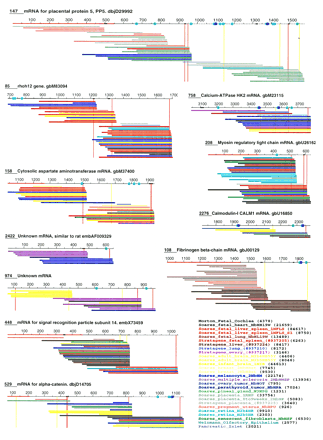
Clusters of 3′ ESTs aligned with their respective contigs (top line of each cluster). Contigs annotated with a GenBank entry name can be considered as identical to the corresponding mRNA (BLAST2 score ⩾ 2000, 97%–99% identity over highest scoring segment). (Unknown mRNA) Contigs do not show any significant resemblance (BLAST score ⩾150 or P ⩾ 0.02) to a human non-EST sequence in GenBank release 104. mRNA extensions are shown with broken lines. Contigs that do not extend corresponding mRNAs are numbered from the mRNA 5′ end; other contigs are numbered from position 1. Thicker segments in contigs indicate possible internal priming sites (see Methods). Potential destabilization signals are shown with blue and dark blue dots, corresponding to sequences AUUUA and UUAUUUA(U/A)(U/A), respectively. ESTs are colored according to their source library, as indicated at bottom right. Numbers in parentheses indicate the total number of ESTs in each library. Red lines on contigs indicate coding sequences. Vertical red and yellow lines give the positions of all AAUAAA and AUUAAA sequences, among which are the actual polyadenylation signals (see text). Only ESTs that fully match their respective contig are shown. Clusters are numbered according to the number of ESTs they contain (1 is largest).











