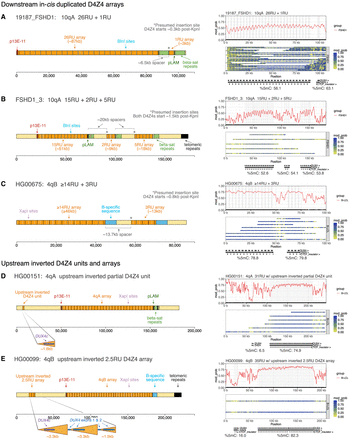
Analysis of the structure and methylation of duplicated D4Z4 arrays and upstream inverted D4Z4 units. (A), (B), (C) Structure of 4q and 10q alleles identified by the pipeline to have in-cis duplications downstream from the D4Z4 array, alongside smoothed methylation plots showing that these duplicated arrays are hypermethylated. The overall %5mC for each D4Z4 array is shown below the annotation track for the methylation plot. The locations of DUX4 exons are shaded in gray on the smoothed methylation plot. (D) Representative example of the inverted D4S2463 unit located ∼42 kb upstream of the 4q D4Z4 array, which is found to be hypomethylated in FSHD and control fibroblasts and B-LCL samples. (E) The HG00675 cell line from the 1000 Genomes Project (Gustafson et al. 2024) has a 4qB allele with an upstream inverted D4Z4 array of 2.5 repeat units in place of the D4S2463 unit. This upstream inverted array is also hypomethylated.











