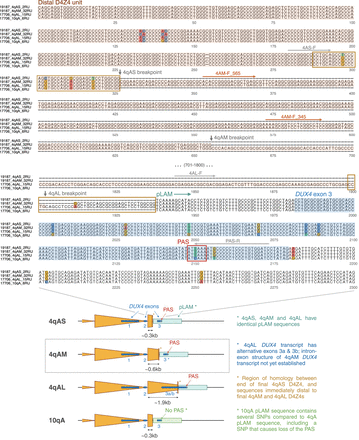
Base-level comparison of distal D4Z4 sequences from 4qAS, 4qAM, 4qAL and 10qA alleles. (Top) Alignment of distal D4Z4 sequences beginning from the proximal KpnI site for the final partial D4Z4 unit, and extending into the proximal portion of the pLAM sequence containing DUX4 exon 3. Sequences are extracted from consensus sequences generated using Racon from high-coverage Cas9-targeted ONT sequencing. Breakpoints for the final partial D4Z4 units (vertical arrows) were determined by alignment to a full-length D4Z4 sequence. The start of pLAM is marked as the first nucleotide following the final partial 4qAS D4Z4 unit. The 4qAM and 4qAL alleles contain an extra length of sequence between the end of their final partial D4Z4 units and the start of pLAM, which corresponds to the end of the 4qAS allele’s final partial D4Z4 unit (shown in orange boxes). Locations of primers used for PCR genotyping of 4qAS/4qAM/4qAL (Supplemental Fig. S6, Supplemental Table S5) are shown above their respective sequences; 4AS-F, 4AL-F, and PAS-R (gray) are from Lemmers et al. (2018b), whereas 4AM-F_565 and 4AM-F_345 (brown) were newly designed using the 4qAM consensus sequence. (Bottom) Schematic comparing the distal sequences of 4qAS, 4qAM (newly identified), 4qAL and 10qA alleles.











