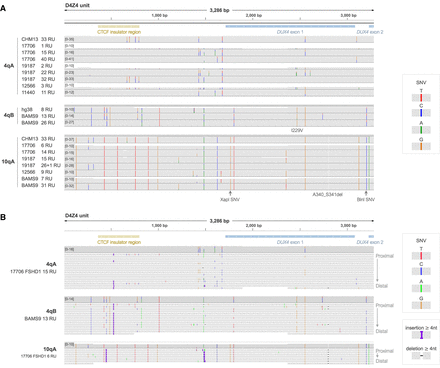
Array-wide and base-level analysis of D4Z4 genetic variants using consensus sequences from high-coverage long-read sequencing. (A) Integrative Genomics Viewer (IGV) coverage plots for alignments of D4Z4 units from the consensus sequences for patient 4q and 10q alleles, showing 4qA-, 4qB- and 10qA-specific genetic variants consistent with CHM13v2.0 and GRCh38. (B) IGV alignments of D4Z4 units from the consensus sequences for representative 4qA, 4qB and 10qA alleles from samples 17,706 and BAMS3. Within each alignment track, full-length D4Z4 units are arranged from top to bottom in order of proximal to distal position within the array. Indels <4 nt which were not consistent between repeat units were hidden to aid in visualization (Supplemental Fig. S3). Consensus sequences were generated using Racon using spanning reads from high-coverage Cas9-targeted ONT sequencing. SNV, single nucleotide variant.











