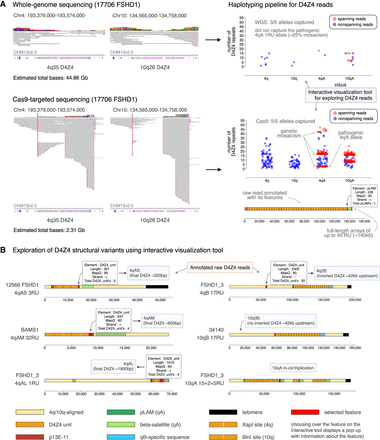
An interactive tool for exploring annotated and haplotyped D4Z4 raw reads from long-read sequencing data. (A) Sequencing data obtained for the 4q35 and 10q26 D4Z4 regions using ultra-long whole-genome sequencing (WGS) and Cas9-targeted sequencing, for sample 17,706 (FSHD1). (Left) Alignment of sequencing data for the D4Z4 region against a single reference genome, such as CHM13v2.0 (32 repeat units for both the 4q35 and 10q26 D4Z4 arrays), leads to large gaps in alignment and potential mis-mapping of reads, making interpretation difficult. (Right) Our pipeline processes the sequencing data to annotate raw reads with their D4Z4 features, enabling accurate repeat unit counting and assignment to 4q/10q and A/B haplotypes. (Top) WGS generates a high total sequencing output (44.86 Gb for sample 17,706) but yields low coverage of the 4q35 and 10q26 regions, which can lead to pathogenic alleles being missed. (Bottom) Cas9-targeted sequencing leads to much higher coverage of the 4q35 and 10q26 regions with a much lower total sequencing output (2.31 Gb for sample 17,706), which enables the capture of low-frequency pathogenic alleles, genetic mosaicism, long full-length arrays, and high numbers of spanning reads. (B) The pipeline is accompanied by an interactive visualization tool that enables exploration of the structure of raw D4Z4 reads, revealing D4Z4 variants such as (left) 4qAS, 4qAL and a newly identified 4qAM allele, as well as (right) 10qB alleles from 4q/10q translocations, and in-cis triplication alleles.











