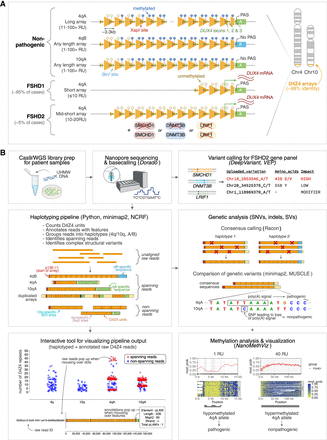
D4Z4End2End: an all-in-one, long-read-based workflow for the genetic and epigenetic analysis of FSHD. (A) Schematic of the pathogenesis of FSHD. The causative locus of FSHD is the D4Z4 macrosatellite array at the Chromosome 4q subtelomere, which comprises 1 to >100 ∼3.3 kb tandem repeat units. FSHD occurs in the presence of (i) a PAS-containing D4Z4 haplotype (4qA), which enables stable DUX4 expression and (ii) epigenetic dysregulation of the array, which can occur via array contraction (FSHD1) or pathogenic variants in chromatin modifiers (FSHD2). Both FSHD1 and FSHD2 are associated with D4Z4 hypomethylation and ectopic DUX4 expression in skeletal muscle. A nonpathogenic, homologous D4Z4 array is located at the Chromosome 10q subtelomere. A 4q-specific XapI site and 10q-specific BlnI site is used to distinguish 4q and 10q alleles in Southern blot assays. (B) Overview of the all-in-one, long-read-based workflow for FSHD. Ultra-high-molecular-weight (UHMW) DNA is used to perform Oxford Nanopore Technologies (ONT) Cas9-targeted or whole-genome sequencing (WGS). Canonical and 5mCG basecalling is performed using Dorado. Variants in reads overlapping FSHD2-associated genes are called using DeepVariant (Poplin et al. 2018), and the potential pathogenicity of these variants is assessed using Variant Effect Predictor (VEP) (McLaren et al. 2016). For assessment of the D4Z4 array, raw reads are input into a custom script which annotates and haplotypes D4Z4 reads (Supplemental Code). The output of the haplotyping pipeline can be explored using an interactive visualization tool, which displays raw reads as individual dots grouped into their haplotypes and plotted based on their number of D4Z4 repeat units (y-axis), and highlights 4q and 10q spanning reads. Hovering over individual dots displays the annotated raw read. Accurate consensus sequences for 4q and 10q alleles can be generated from spanning reads using Racon (Vaser et al. 2017), enabling analysis of allele-specific D4Z4 genetic variants. Allele-specific, array-wide methylation analysis is performed using NanoMethViz (Su et al. 2021, 2025), allowing detection and visualization of FSHD1- and FSHD2-specific methylation patterns.











