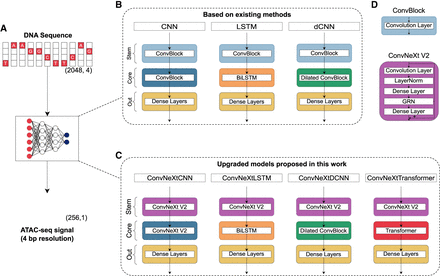
Overview of the benchmarking setup. (A) Models are compared based on their ability to predict experimental ATAC-seq at 4-bp resolution from an input of 2048-bp long DNA sequences. (B) Models based on existing methods include CNN, LSTM, and dCNN-based architectures. Each model has a convolution-based stem to extract genomic features. (C) Models proposed in this work, including a transformer-based architecture. The new models use ConvNeXt V2 blocks to effectively extract features from the input DNA sequence. (D) ConvBlock uses a single convolutional layer whereas ConvNeXt V2 block additionally uses dense layers, layer normalization and Global Response Normalization (GRN).











