Revisiting models of enhancer–promoter communication in gene regulation
Abstract
Enhancer–promoter communication is fundamental to gene regulation in metazoans, yet the mechanisms underlying these interactions remain debated. Two primary models have been proposed: the structural bridge model, in which enhancers and promoters come into close proximity through stable, protein-mediated interactions, and the hub model, in which dynamic clusters of transcription-associated proteins facilitate communication over variable distances. Emerging evidence suggests that although enhancer–promoter pairs do come into close proximity during transcriptional activation, these interactions are highly transient, and the precise distances remain challenging to measure. Moving forward, resolving the distinctions between these models will require novel techniques to more precisely measure the spatial and temporal dynamics of enhancer–promoter interactions. Understanding how enhancers interact with promoters will deepen our understanding of the regulation of gene expression and the molecular underpinnings of transcriptional control.
Gene expression programs are encoded by cis-regulatory elements, specialized DNA sequences that control the timing of transcriptional activation. Two primary types of cis-regulatory element impact transcription in Metazoan species: promoters, which facilitate transcription initiation from an embedded transcription start site, and enhancers, which regulate the rate of mRNA production from distal promoters. The discovery of the SV40 viral enhancer was a landmark event more than 40 years ago. The 72 bp DNA sequence encoding the SV40 enhancer increased the expression of a reporter gene by more than two orders of magnitude in a manner that was largely independent of its position and orientation (Banerji et al. 1981; Benoist and Chambon 1981; Gruss et al. 1981). Today we know of numerous examples in which enhancers regulate gene expression from long genomic distances (∼10–100 kb) and are occasionally found >1 Mb away from the target promoter (Lettice et al. 2003; Spitz et al. 2003; Antonellis et al. 2006; Ghiasvand et al. 2011; Hay et al. 2016; Gasperini et al. 2019). These examples raise numerous questions about how enhancers reliably find their target promoter within a crowded nucleus, the structure of interacting enhancer–promoter pairs, and the mechanistic basis by which they communicate.
Enhancers: functional elements characterized by a stereotyped chromatin architecture
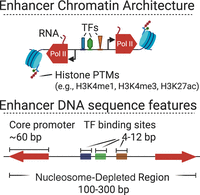
Enhancers are one variety of cis-regulatory element encoded in Metazoan genomes. They are characterized by a stereotyped pattern of transcription factor binding, transcription initiation, and chromatin organization (Fig. 1). Enhancers are often bound by one or more of the approximately 1800 transcription factors (Lambert et al. 2018). Transcription factors bind to short consensus DNA sequence motifs (∼4–12 bp) embedded in nucleosome-depleted regions ∼100–300 bp wide (He et al. 2010; Core et al. 2014; Scruggs et al. 2015; Grossman et al. 2018). Pioneer transcription factors, the first to recognize and bind to DNA even in the presence of nucleosomes, recruit histone remodeling complexes that move or displace nucleosomes, thereby creating nucleosome-free regions (Zaret and Carroll 2011). Transcriptional activators bind the nucleosome-free region and tag the flanking +1 and −1 nucleosomes with post-translational modifications, such as acetylation (H3K27ac) and methylation (H3K4me3 and H3K4me1) (Barski et al. 2007; Heintzman et al. 2007; Creyghton et al. 2010; Henriques et al. 2018).
Stereotypical features of chromatin organization at enhancers. Enhancers are units with a stereotypical organization composed of transcription factors, histones carrying characteristic post-translational modifications (PTMs), and Pol II (top) and DNA sequence codes (bottom). (Created in BioRender. Danko, C. [2025] https://www.biorender.com)
Transcription factors also promote the bidirectional initiation of RNA polymerase II (Pol II) at enhancers (De Santa et al. 2010; Kim et al. 2010; Hah et al. 2013; Andersson et al. 2014; Core et al. 2014). Pol II initiates bidirectionally, transcribes 20–60 bp, and enters a paused state, known as the promoter proximal pause (Fig. 1; Buckley et al. 2014; Jonkers et al. 2014; Henriques et al. 2018). At enhancers, these short and typically bidirectional transcription units encode enhancer-templated RNAs (eRNAs), which are terminated and rapidly degraded (De Santa et al. 2010; Kim et al. 2010).
Several lines of evidence indicate that either the act of transcription or the nascent eRNA has an important functional role. First, several studies have knocked down eRNAs, revealing a small but statistically measurable decrease in the expression of target genes (Lam et al. 2013; Melo et al. 2013; Schaukowitch et al. 2014). Second, depleting Pol II results in the immediate loss of chromatin marks indicative of enhancers, like H3K27ac (Martin et al. 2021; Wang et al. 2022), as well as enhancer–promoter loop interactions (Barshad et al. 2023; Zhang et al. 2023) (discussed in detail below). Third, removing core promoter elements, which control transcription initiation and pausing, from reporter constructs decreased enhancer activity (Tippens et al. 2020). Thus, although much work remains to be done in elucidating the role of eRNAs and other promoter and enhancer marks, existing data nevertheless support a model in which either the act of transcription or the sequence of the RNA is important for enhancer function.
The stereotyped chromatin and transcriptional environment at enhancers is similar to that observed at active promoters, raising questions about whether enhancers and promoters represent distinct classes of cis-regulatory element. In addition to evidence that enhancers initiate eRNA transcription from their own DNA sequence, thereby exhibiting promoter-like activity, several studies have shown that a subset of active promoters can function as enhancers to increase transcription at distant loci (Engreitz et al. 2016; Dao et al. 2017). These findings support a model in which cis-regulatory elements may encode both enhancer and promoter functions, depending on context (Andersson et al. 2015; Kim and Shiekhattar 2015; Andersson and Sandelin 2020).
Prevailing models of enhancer communication: structural bridge versus malleable hub
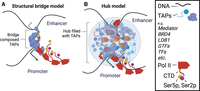
Because enhancer–promoter interactions are a critical feature of Metazoan gene regulation, much attention has been paid to the fundamental mechanisms that enable enhancer–promoter communication. Much of this discussion revolves around the architecture of enhancers’ interactions with their cognate promoters. For many years, the prevailing model proposed that enhancers and promoters come into close physical proximity in the nucleus, creating “loops” composed of the DNA polymer connecting them (Choi and Engel 1988; Robson et al. 2019; Schoenfelder and Fraser 2019). Classical looping models, which we collectively refer to as the structural bridge model, represent these enhancer–promoter interactions as a physical bridge consisting of highly stereotyped protein–protein interactions connecting the enhancer to its target promoter (Fig 2A; Yin and Wang 2014; Kim and Shiekhattar 2015; Petrenko et al. 2016).
Overview of the structural bridge and hub models. (A) The structural bridge model postulates a stereotyped bridge composed of protein–protein interactions between transcription-associated proteins (TAPs). (B) The hub model postulates a malleable hub filled with high local concentrations of TAPs. (Created in BioRender. Danko, C. [2025] https://www.biorender.com)
Despite some experimental support, however, several recent observations are not compatible with classical versions of the structural bridge model. An alternative model, which we refer to as the “hub” model, was proposed to address these limitations (Fig 2B; Di Giammartino et al. 2020; Bhat et al. 2021; Lim and Levine 2021; Hamamoto and Fukaya 2022). The hub model predicts that enhancer and promoter regions form malleable “hubs,” which consist of clusters of transcription-associated proteins (TAPs), such as Pol II, mediator, transcription factors, the candidate enhancer–promoter architectural protein YY1, and transcriptional coactivators and corepressors. Hubs are postulated to contain high local concentrations of TAPs; as a result, most TAPs are concentrated into specific nuclear subcompartments that comprise a relatively small portion of the nucleoplasm. The individual protein–protein or protein–nucleic acid interactions supporting the recruitment of TAPs to a hub are proposed to be relatively weak but numerous and supported by a high local concentration of a variety of interacting factors within the hub.
Variations on the hub model place different constraints on the relative physical position of the enhancer and promoter DNA. Some theoretical models postulate the transmission of regulatory information over large physical distances, including action-at-a-distance, which suggests allostery between transcriptional regulators within a liquid droplet space (Bialek et al. 2019), and the transcription factor activity gradient model, which suggests a gradient-directed flow of activated transcription factors from enhancers to their target promoters, which are spatially close (Karr et al. 2022). Notably, these models allow for fairly large distances, on the order of hundreds of nanometers, between enhancers and their target promoters. Readers interested in exploring these additional models are directed to a recent review by Yang and Hansen (2024) for additional discussion.
It is perhaps worth noting that the hub model has deep roots in transcription models that have been discussed in the literature for decades. The hub model is similar to the concept of transcription factories, which were first proposed after findings that nascent RNA production occurs in a few hundred focal sites within each nucleus (Jackson et al. 1993; Iborra et al. 1996; for review, see Sutherland and Bickmore 2009). In a sense, the hub concept advances these ideas by proposing that these transcription factories are the sites in which enhancer–promoter communication occurs. Finally, the hub model is also similar to the more recently proposed liquid–liquid phase separation models for transcriptional activation (Hnisz et al. 2017), but we note that a general hub model does not necessarily require the stringent biophysical definition of phase separation to be met precisely.
Transcriptional regulatory hubs may be just one of many distinct varieties of nuclear subcompartment (for reviews, see Bhat et al. 2021; Belmont 2022). Classic examples of nuclear subcompartments include the nucleolus (McClintock 1934), nuclear speckles (Swift 1959; Beck 1961), and dense heterochromatin located near the nuclear membrane (Pickersgill et al. 2006; Guelen et al. 2008; Kind and van Steensel 2010), each of which have unique physical and functional characteristics that facilitate their biological role. More recent work reports clusters of TAPs, including Pol II and mediator, that have been observed in living cells and may be direct observations of a transcriptional hub (Cho et al. 2018).
Throughout the remainder of this review, we will examine key findings about the dynamics, structural properties, and proteins involved in mediating enhancer–promoter communication. We will place these findings within the context of both the structural bridge and hub models. It is important to note that these two models are not mutually exclusive; we hope our review will outline the key evidence for each model and present a path toward understanding the fundamental biology of enhancer–promoter communication.
TAPs have properties that facilitate hub formation
Both the structural bridge and hub models postulate that interactions between TAPs, such as transcription factors, Pol II, mediator, cohesin, and a wide variety of transcriptional coactivator and corepressor proteins and protein complexes, have important roles in enhancer–promoter communication. Because these factors also regulate transcription, their roles in enhancer–promoter communication are closely intertwined with their functions in the Pol II transcription cycle.
Protein–protein interactions between TAPs are mediated through highly specialized protein domains. One of the most important protein domain classes for the formation of transcription hubs is intrinsically disordered regions (IDRs). IDRs contain amino acid motifs that facilitate protein–protein interactions and thereby facilitate membership in a transcription hub (Boija et al. 2018; Sabari et al. 2018). For instance, transcription factors often contain IDRs that facilitate interactions between the transcription factor and a variety of cofactors, chromatin remodelers, and other flavors of TAP (Frietze and Farnham 2011). These protein–protein interactions facilitate localization within a hub and are therefore critical for many aspects of transcription factor function. For instance, despite not having a direct role in DNA binding, IDRs in the Saccharomyces cerevisiae transcription factors Yap1 and Msn2 were nevertheless critical determinants of which of the various DNA sequence motifs each transcription factor bound in vivo (Brodsky et al. 2020). One attractive model to explain these observations is that the IDRs are required to localize transcription factors into a hub. By analogy, IDRs serve as a postal code that directs the transcription factor to one of the many nuclear subcompartments. Once in the transcriptional regulatory hub, the DNA-binding domains of the transcription factor identify the subset of high-affinity targets in DNA that are accessible to the hub (Bhat et al. 2021). This model is attractive because it explains how transcription factors identify their complement of DNA binding sites in organisms with large genomes more quickly than can be explained under more simplistic diffusion models (Jana et al. 2021).
Like transcription factors, core transcription proteins, including RNA Pol II, mediator, general transcription factors, and the coactivators and corepressors that reorganize chromatin, also have IDRs or other domains that facilitate hub membership. RNA Pol II has several domains and macromolecule associations that could be involved. First, the largest subunit of Pol II (RPB1) contains a C-terminal heptad repeat domain that is an IDR that contributes to macromolecular clustering (Banani et al. 2017; Boija et al. 2018; Cho et al. 2018; Chong et al. 2018; Sabari et al. 2018; Strom et al. 2018). Second, the nascent RNA emerging from the Pol II exit channel may also contribute to clustering. RNA can facilitate hub membership either through the m6A RNA modification (Barutcu et al. 2019; Nair et al. 2019) or by interactions between RNA and transcription factors, mediator, NELF, or other transcriptional coactivators (Sigova et al. 2015; Vos et al. 2016; Bose et al. 2017; Oksuz et al. 2023). Third, reports indicate that secondary structures like G-quadruplexes or R-loops can link eRNAs to a target promoter (Hegyi 2015; Akiki et al. 2024). Like Pol II, the mediator complex also forms clusters both on its own (Zamudio et al. 2019) and in conjunction with Pol II via networks of interactions between tyrosine and proline amino acids (Flores-Solis et al. 2023). Similar interactions underlie the recruitment of nucleosome remodelers, including the BAF complex (Patil et al. 2023), BRD4 (Sabari et al. 2018), and components of the Pol II preinitiation complex (PIC) (Wei et al. 2020).
Given the importance of TAPs in forming both hubs and/ or a structural bridge, it stands to reason that these proteins should play an important role in establishing enhancer–promoter interactions. Indeed, recent studies using high-resolution chromosome conformation capture (3C) methods have found that depletion of both Pol II (Barshad et al. 2023; Zhang et al. 2023) and mediator (Kagey et al. 2010; Ramasamy et al. 2023) impact enhancer–promoter contact frequency. Likewise, multiple lines of evidence indicate that transcription factors are also important for shaping enhancer–promoter interactions. Changes in transcription factor binding between different conditions are associated with changes in the contact frequency between enhancer–promoter pairs (Vakoc et al. 2005; Grubert et al. 2015; Rubin et al. 2017; Johanson et al. 2018; Ray et al. 2019; Portillo-Ledesma et al. 2023). Moreover, recently developed Micro-C variants that allow single-base resolution for loop discovery have found that loop ends are frequently adjacent to transcription factor binding sites (Hua et al. 2021). Others have found that chromatin is organized into smaller, localized microcompartments, which may be direct observations of transcriptional hubs (Rowley et al. 2017; Goel et al. 2023). As transcription factors have a key role in establishing the chromatin environment characteristic of an enhancer, these associations between transcription factors and enhancer–promoter contract are likely to be causal.
Collectively, these studies demonstrate the importance of TAPs in mediating enhancer–promoter contacts and communication.
CTCF and cohesin: indirect players in enhancer–promoter loop formation
CTCF and cohesin are two proteins that have long been postulated to be involved in a structural bridge connecting enhancer and promoter regions (Yin and Wang 2014; Kim and Shiekhattar 2015; Petrenko et al. 2016). However, emerging evidence indicates that the role both protein complexes play in transcription and enhancer–promoter loop formation is indirect.
CTCF and cohesin are key molecules in the formation of topological associated domains (TADs) through a molecular process called loop extrusion (Sanborn et al. 2015; Fudenberg et al. 2016). Cohesin is a protein complex that forms a ring structure around DNA (Merkenschlager and Nora 2016). Once the ring is formed, cohesin has an integrated motor that pulls DNA through the ring by consuming ATP for energy (Ganji et al. 2018; Vian et al. 2018). As DNA is pulled through the cohesin complex, two points of DNA are held by cohesin in very close proximity, forming a loop that can be resolved using both 3C and microscopy-based approaches. Because of these loops and the continual nature of the extrusion process, this model earned the name “loop extrusion.”
CTCF is a highly conserved protein with a zinc finger domain that binds DNA in a sequence-specific manner and changes the dynamics of loop extrusion near its binding site. Early models postulated that CTCF blocked loop extrusion through a steric mechanism by capturing the cohesin complex through protein–protein interactions, a bit like a cork fits into the neck of a bottle (Rao et al. 2014; Tang et al. 2015). More recent work has argued that CTCF blocks cohesin by causing tension in the DNA polymer that is incompatible with extrusion past the CTCF binding site (Davidson et al. 2023). Regardless of the mechanism, all authors agree that CTCF limits loop extrusion in a directional manner, such that the extruding complex is prevented from extending past the CTCF binding site only in one orientation.
Loop extrusion involves the dynamic activity of the cohesin complex over relatively long timescales. Once loaded onto DNA, cohesin can remain associated with a locus for extended periods while continuously extruding DNA to form loops. Direct estimates of residence time are available when cohesin is interacting with CTCF, where it remains stably bound for ∼30 min (Hansen et al. 2017). Halting the process of loop extrusion through the degradation of RAD21, a key protein in the cohesin complex, results in the near immediate loss of most TADs, based on in situ Hi-C data (Rao et al. 2017), as the loops held together by cohesin diffuse apart.
Loop extrusion has several indirect impacts on enhancer–promoter loop formation. First, cumulative evidence from both sides of a raging scientific debate indicate that TADs boundaries reduce, but do not completely eliminate, the effect of an enhancer on a target promoter (for a more detailed summary of literature on this topic, see Symmons et al. 2014; for review, see Cavalheiro et al. 2021; Zuin et al. 2022). Second, loop extrusion may also play a role in increasing the chances that distally located enhancer–promoter pairs will find one another in the crowded nucleus. Promoters are often located near the end of a TAD, and these TAD boundaries are often associated with an in situ Hi-C pattern known as a “stripe,” which represents the promoter forming high frequency interactions across the TAD (Vian et al. 2018). These frequent interactions effectively change the problem of enhancer–promoter discovery from a three-dimensional search in the nucleus to a guided one-dimensional search in which loop extrusion assists the promoter in interacting with enhancers within the stripe. Indeed, NIPBL cohesin loading factor (NIPBL), encoded by Nipbl, loads the cohesin complex onto DNA near promoters and enhancers (Kagey et al. 2010), facilitating this mechanism. Finally, even when promoters are not directly adjacent to the end of the TAD, the process of loop extrusion could nevertheless constrain the three-dimensional diffusion to a smaller space, by preventing diffusion further apart than the current constraints placed by the loop. Indeed, these mechanisms may explain why cohesin depletion has a larger impact on enhancer–promoter pairs located further from one another along the chromatin fiber (Guckelberger et al. 2024).
Despite indirect effects, however, the evidence suggests that loop extrusion does not play a direct role in enhancer–promoter loop formation. The primary evidence for this point is the development of degron systems that rapidly deplete RAD21 (Rao et al. 2017). Following 6 h of RAD21 depletion, both TADs and CTCF-cohesin-dependent loop domains were lost (Rao et al. 2017). Surprisingly, however, measurements of gene transcription using PRO-seq found that transcriptional changes were relatively small in magnitude (less than about 1.5-fold) at the same 6 h of RAD21 depletion (Rao et al. 2017). This suggests a surprisingly modest impact of cohesin on transcriptional regulation.
Transcription also feeds back to impact loop extrusion in several ways. First, cohesin loading is impacted by depletion of both Pol II and mediator (Zhang et al. 2021; Ramasamy et al. 2023), consistent with the aforementioned importance of enhancer–promoter DNA for NIPBL binding (Kagey et al. 2010). Additionally, RNA Pol II is a large protein complex that prevents cohesin's extrusion past an engaged Pol II (Banigan et al. 2023; Zhang et al. 2023). As Pol II takes ∼5–10 min to transcribe a median-length gene (assuming an elongation rate of ∼1.5–4 kb/min) (Danko et al. 2013; Jonkers et al. 2014), the active elongation complex can push the cohesin machinery across tens of thousands of bases of DNA for timescales that are on the same order of magnitude as the interaction between cohesin and CTCF.
These lines of evidence indicate that cohesin and CTCF are unlikely to form a stable structural bridge whose primary function is to facilitate enhancer–promoter interactions. Instead, both cohesin and CTCF play a primary role in loop extrusion, which exerts indirect effects on transcriptional hubs or loops.
Distances between enhancer–promoter DNA during interactions
A key difference between the structural bridge and hub models concerns the distance between enhancer and promoter DNA. The structural bridge model predicts that the enhancer and promoter come into close physical proximity, such that the distance between them can be spanned by a bridge formed through protein–protein interactions. However, recent work using imaging methods has called these relatively short distances between interacting DNA into question.
Several of the key observations that led the community to challenge the structural bridge model were imaging studies that measured the distance separating enhancer–promoter pairs in a variety of model systems. These studies revealed that enhancers and promoters are frequently on the order of hundreds of nanometers apart, even during transcriptional activation (Chen et al. 2018; Alexander et al. 2019; Benabdallah et al. 2019). In one particularly interesting example, the distance between the sonic hedgehog (Shh) promoter and a developmental enhancer increased following Shh gene activation (Benabdallah et al. 2019). These long distances, if estimated correctly, are almost certainly not compatible with the formation of a physical protein–protein bridge. However, numerous technical issues make it challenging to measure precise distances using imaging methods (as noted below).
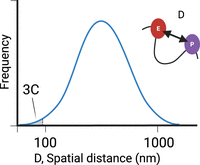
In contrast to the results of imaging studies, 3C-based genomic studies suggest that enhancers and promoters come into close proximity in the nucleus. 3C-based methods such as in situ Hi-C (Rao et al. 2014) and Micro-C (Hsieh et al. 2020; Krietenstein and Rando 2022) measure the frequency of ligation between DNA sequences that are close together in 3D space. These 3C methods are only sensitive to the tail of the distribution of molecular distances between two loci, because they only capture interactions that are close enough for the free ends of DNA to be cross-linked and ligated to one another (Fig. 3; Fudenberg and Imakaev 2017). The exact proximity between 3C-based ligation events remains unknown (see discussion below).
Enhancer promoter distance measurements. The probability density function (PDF) represents the distribution of distances between candidate promoter–enhancer pairs. Imaging methods can theoretically measure the entire distribution, whereas 3C-based methods only measure the lower tail. Distribution drawn based on data from Chen et al. (2018). (Created in BioRender. Danko, C. [2025] https://www.biorender.com)
Multiple studies demonstrate associations between 3C contact frequency and enhancer–promoter function that appear to indicate that DNA sequences come close together at some point during transcriptional activation. Many studies have now shown that 3C methods can predict the functional impact of enhancers on a target gene (Fulco et al. 2019; Ray et al. 2019; Zuin et al. 2022; Barshad et al. 2023). Likewise, changes in either the composition or frequency of enhancer–promoter loop interactions are associated with the activation or repression of target promoters (Rubin et al. 2017; Mateo et al. 2019; Beagan et al. 2020; Titus et al. 2024). One such study introduced an elegant system that ectopically creates an enhancer–promoter loop interaction in the presence of blue light, without obvious changes in activity of transcription factors or other proteins associated with the enhancer or promoter (Kim et al. 2019). This study noted a small but significant increase in the expression of a target gene after enhancer–promoter loop formation (Kim et al. 2019), indicating that loops likely cause increased target gene expression.
Such perturbation experiments, combined with newer and higher-resolution 3C variants, have shown that functionally active enhancer–promoter pairs have higher contact frequencies compared with nonfunctional pairs and that these differences are most prominent close to the enhancer and promoter. A recent study found that enhancers that elicit a functional impact on the expression of its target promoter had ∼30% higher contact frequency compared with enhancer–promoter pairs with no evidence of functional impact (Barshad et al. 2023). Moreover, by using nucleosome resolution Micro-C data, the authors showed that differences between functional and nonfunctional pairs were most prominent between the first two nucleosomes (+1 and +2) of the enhancer and promoter, adjacent to the transcription start site (Barshad et al. 2023). These results suggest that enhancers come very close to their target promoters during at least one point of enhancer activation.
It is important to note that relatively close enhancer–promoter distances do not necessarily require a structural bridge model. Although some proponents of the hub model have argued that enhancers and promoters may not be able to come close together owing to issues of molecular crowding within a hub (Lim and Levine 2021), we do not see any strong theoretical reasons to believe that enhancer–promoter DNA must remain far apart from one another. Rather, the results of 3C studies discussed above may reflect the formation of transcriptional hubs in which some functional enhancer–promoter pairs spend a small amount of time at shorter distances than measured by imaging studies.
There are several other factors that complicate distinguishing the hub and bridge models. First, imaging studies have observed considerable variability in the diameter of transcription factories across distinct loci (Iborra et al. 1996; Eskiw et al. 2008; Eskiw and Fraser 2011), potentially raising the prospect that the few loci measured in microscopy studies are uncharacteristically large. Second, microscopy studies capture the average interaction distances occurring within a larger hub but miss interactions occurring on a transient physical bridge, which may be more directly related to a transcriptional burst (see additional discussion of that matter below). Third, it is difficult to determine how large a structural bridge would have to be based on current data. Mediator is a remarkably large protein complex: CryoEM structures show the distance from DNA bound by the Pol II PIC to the most distant edge of the mediator complex is ∼20 nm, whereas the whole structure spans at least 30 nm at its widest point (Rengachari et al. 2021). Moreover, because enhancers are transcribed, it is plausible that separate Pol II and mediator complexes are present at enhancers and promoters, which could potentially increase the distance spanned by a structural bridge considerably further. Finally, although Pol II and mediator are outstanding candidates for a structural bridge owing to their importance for transcription, it is also possible that additional proteins may be involved in these interactions that extend a structural bridge further than anticipated.
Dynamics of enhancer–promoter interactions
Early versions of the structural bridge model envisioned a physical bridge that was stable for relatively long durations. In contrast, however, multiple lines of evidence indicate that enhancer–promoter interactions are extremely transient. One study used a piggyBac transposon system to place a single enhancer–promoter pair in hundreds of different relative positions and orientations. Kinetic models best matched these experimental observations when enhancer–promoter interactions occurred much faster than other regulatory steps, typically lasting for a timescale of seconds or less (Zuin et al. 2022). More recently, live-cell imaging systems simultaneously tracking the position of the Sox2 gene, its enhancer, and a Pol II hub showed that Sox2 burst size increased when all three units are transiently close together, relative to their typical distances, for as little as just a few imaging frames (Du et al. 2024). We note that although the absolute distances observed by Du et al. (2024) during transcriptional bursts are large, these distances could be influenced by the challenges designing and preparing probes for imaging studies that plague all imaging approaches (discussed below).
Live cell imaging measurements show considerable variation in how long-lived different transcription hubs are in the nucleus. Measurements of Pol II cluster residence times have found a mean of ∼5–12 sec (Cisse et al. 2013; Cho et al. 2016; Du et al. 2024); however, some clusters, including those located near stronger “super-enhancers,” persist for >2 min (Cho et al. 2016; Du et al. 2024). Studies using 3C-based measurements also find evidence for considerable variation in contact frequency across the genome. Loci located near highly active enhancers, like super-enhancers in the MYC locus, have much higher 3C contact frequency than the average enhancer–promoter pair (Barshad et al. 2023).
Taken together, current literature supports a “kiss-and-go” model, in which most enhancer–promoter interactions are transient and sufficient to induce a transcriptional burst.
Reconciling distance and time measurements between 3C and imaging methods
The results from imaging and genomic studies appear challenging to reconcile at first glance. It is important to keep in mind precisely what each approach measures, as well as the technical limitations, when interpreting results. In this section, we summarize the most fundamental limitations of 3C methods before discussing limitations in imaging approaches. We close with how these technical limitations influence our understanding of the distance and dynamics of enhancer–promoter interaction.
The first challenge with interpreting 3C methods is that they do not directly measure distance between interacting loci. Rather, 3C methods can detect only the contacts between DNA sequences in very close physical apposition, at the lower tail of the distribution two loci might occupy in cells (Fig. 3). How far apart two DNA sequences need to be for detection by 3C methods is not known precisely, but it may be possible to obtain rough estimates. The most conservative model, which assumes that contacts can be obtained if N-terminal nucleosome tails are cross-linked, will still only detect DNA closer than 100 nm. However, most modern Hi-C and Micro-C data sets use in situ ligation protocols performed in intact nuclei. Performing ligation reactions in situ likely makes the distance less sensitive to more distal contacts for two reasons: First, many blunt ends are available for ligation, resulting in competition for ligation events that favor the detection of more accessible, shorter contacts. Second, a network of cross-links likely prevents diffusion from ligating more distal DNA compared with older Hi-C protocols that perform ligations in dilute conditions. Taken together, these considerations most likely mean that DNA contacts observed in a 3C method are much closer than 100 nm. Although the effects of cross-linking on nuclear architecture remains an important caveat (see discussion below), it is possible that distances may be on the order of tens of nanometers. Candidate contacts between loci further than this distance threshold would not be detected in a 3C library.
A second limitation of 3C methods is that contact maps reflect a time-weighted average of interactions over a population of cells and 3D chromatin conformations adopted by two loci. These averages include both times when enhancer–promoter communication is occurring and when loci lie dormant. Above, we note reasonably strong evidence that enhancer–promoter residence times are relatively short. Such short residence times may explain why the effect of enhancer–promoter contacts is a relatively minor component of the signal obtained in 3C maps.
Although imaging studies can directly measure distance without a bias for shorter interactions, a variety of technical factors complicate distance measurements by imaging methods. Imaging studies require probes to measure DNA. Often probes hybridize a DNA carrying a fluorescent tag to nuclear DNA to identify the position of the enhancer and/or promoter. These probes often lie thousands of bases from the target regulatory region. Moreover, in many studies the regulatory region itself is coarsely defined, often as a ∼1–20 kb window that contains one or more enhancer units. The placement of the probes and the coarse-grained resolution of the enhancer unit make it challenging to interpret distance even in a microscopy experiment. Why? The main reason is that DNA is a long polymer: 1000 bp of naked DNA stretches ∼340 nm. Even considering that chromatin may be compacted by an order of magnitude in a living nucleus, a probe located a few kilobases from the precise ∼110 bp active unit of an enhancer (or promoter) could still lie tens to hundreds of nanometers away from its intended target. The potential for error further increases as investigators adapt these techniques to measure the distance between two (or more) loci, each of which has its own measurement error. These experimental difficulties make measuring precise enhancer–promoter distances challenging, and they require extreme care to be taken in the interpretation of the resulting distances.
Another important challenge shared by many imaging and most 3C studies is that current approaches rely on cross-linking with formaldehyde. Although ostensibly this step is devised to preserve the structure of cells through highly disruptive enzymatic steps, it has been widely reported to alter protein binding and nuclear structure in ways that may obscure accurate measurements (for a detailed explanations, see Belmont 2014; Gavrilov et al. 2015). To investigate the impact of formaldehyde cross-linking on chromatin structures, new methods have been introduced that omit or substantially change fixation techniques in 3C assays. First, the introduction of in situ Hi-C in 2014 introduced a new protocol in which ligation is performed prior to fragmenting nuclei. Performing in situ Hi-C or 4C without formaldehyde fixation generally recovers the same features as full Hi-C maps generated using more conventional cell preparation strategies (Rao et al. 2014; Brant et al. 2016). Second, DamC, an assay that measures DNA proximity by recruiting a DNA methylase to a target anchor (Redolfi et al. 2019), omits both formaldehyde cross-linking and ligation, yet resolves very similar structures as 4C and Hi-C. Third, recent studies introduced alternative cross-linkers that directly cross-link DNA to DNA (You et al. 2021) and recovered the same features identified using Hi-C. Overall, these approaches generally provide support for the structures first observed using conventional Hi-C data, including TADs and promoter–enhancer loops. Although these initial experiments appear to confirm structures identified using Hi-C, continuing to develop creative new methods that provide alternatives to formaldehyde cross-linking is an essential next step for the continued development of the field.
These considerations, along with the relatively short residence times of communication, make us unable to determine precisely how far apart enhancer–promoter pairs lie during transcriptional activation. However, associations between enhancer–promoter function and 3C contact frequency seem incompatible with models in which distances are larger than ∼100 nm during the entire transcriptional activation cycle. These findings are potentially compatible with the formation of a transitory structural bridge model and imply that distances between functionally interacting enhancers and promoters may not be as large as some proponents of the hub model would expect. Nevertheless, it remains challenging to distinguish between a structural bridge and variants of hub models where direct contact is important.
Future outlook and open questions
Although significant progress has been made in understanding the dynamics, distances, and models of enhancer–promoter communication, several key questions and debates remain unresolved. First, when during the transcription activation cycle does communication happen? The prevailing model suggests that communication occurs during a transcriptional burst; however, it is also possible that communication immediately precedes a burst, and interactions are released before the burst completes. Moreover, just as enhancer–promoter interactions can influence the transcription cycle, transcription can also influence contacts, as seen by recent studies showing that Pol II in general (Zhang et al. 2023), and even Pol II pausing (Barshad et al. 2023; Penagos-Puig et al. 2023), can affect enhancer–promoter interactions. Determining when communication occurs during the transcription activation cycle could provide critical insights into the regulation of gene expression and the mechanisms underlying transcriptional bursts. Second, do interactions better resemble a structural bridge or hub? The two models are not mutually exclusive. Although the literature appears to provide more and more support for hubs of some variety, evidence also suggests that enhancer–promoter pairs come into close apposition, and it remains possible that hubs support the formation of a transient protein–protein bridge during transcriptional activation. Understanding how elements of the structural bridge and hub models might coexist or function together could reshape our understanding of the molecular mechanisms governing enhancer–promoter communication.
These open questions highlight the urgent need for new techniques that can more accurately capture the spatial and temporal dynamics of enhancer–promoter interactions. Existing methods are not adequate for measuring enhancer–promoter communication. Imaging approaches risk changing the system they are measuring, probing neighboring DNA and thereby reducing accuracy, or ignoring rare events at the tail end of a distance distribution. Conversely, 3C methods do not directly measure distances, and short residence times ensure enhancer–promoter communication is a minor part of 3C signal. These considerations highlight the need for new methods that more precisely measure enhancer–promoter interactions in real time. Potential approaches include advanced live-cell imaging, innovative single-molecule techniques, or refinements to 3C-based methods that directly measure distances or specifically target loci while in active states. These new methods will be essential for advancing our understanding of enhancer-mediated gene regulation.
Competing interest statement
The authors declare no competing interests.
Acknowledgments
We thank members of the Danko laboratory, John Lis, Abdullah Ozer, James Lewis, and many other scientists at Cornell and around the world for thoughtful conversations that have shaped our thoughts on this topic. This work was supported by the National Human Genome Research Institute (R01-HG010346).
Footnotes
-
Article and publication date are at https://www.genome.org/cgi/doi/10.1101/gr.278389.123.
This article is distributed exclusively by Cold Spring Harbor Laboratory Press for the first six months after the full-issue publication date (see https://genome.cshlp.org/site/misc/terms.xhtml). After six months, it is available under a Creative Commons License (Attribution-NonCommercial 4.0 International), as described at http://creativecommons.org/licenses/by-nc/4.0/.