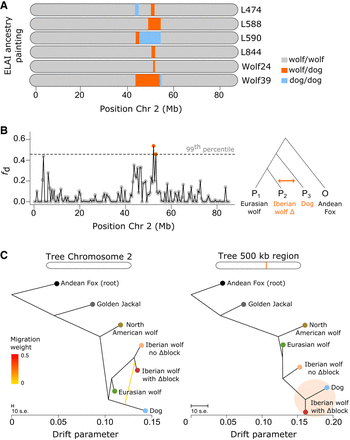
Signatures of excess allele sharing and phylogenetic discordance. (A) Representation of the Δblock on Chromosome 2 in six Iberian wolves using local ancestry analysis. Colors denote the attributed local ancestry: gray for homozygous wolf, orange for wolf/dog (heterozygous), and blue for homozygous dog. (B) Fraction of introgression (fd) across nonoverlapping 500 kb windows on Chromosome 2. The test followed the phylogenetic arrangement depicted on the right: Eurasian wolves as P1, Iberian wolves with the Δblock as P2, and dogs as P3, with the Andean fox as the outgroup. The dashed line indicates the threshold for outlier regions, with orange dots representing 500 kb windows surpassing the 99th percentile. (C) Population trees estimated on Treemix for Chromosome 2 and the 500 kb region ranked as the top window in the fd analysis, involving several canid species (Iberian, Eurasian, and North American wolves, dogs, Golden jackal, and Andean fox). The yellow arrow on the Chromosome 2 tree indicates migration (i.e., ancestry contribution) from dogs to Iberian wolves carrying the Δblock.











