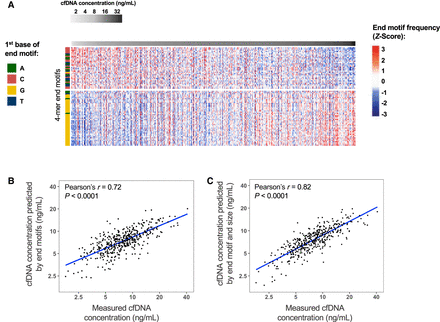
End motifs with significant correlation to cfDNA concentration. The frequencies of all 256 end motifs (4-mer) were correlated to the cfDNA concentration. The resulting analysis revealed 34 negatively correlated end motifs to cfDNA concentration and 45 positively correlated end motifs. (A) Heatmap analysis with rows indicating a particular 4-mer motif, in which the first base of the motif highlighted by a specific color in the left-most column (A, C, G, and T are colored by green, red, yellow, and blue, respectively). Each column indicates a plasma DNA sample from one subject. The frequency z-score, calculated for each end motif, is shown by the color scale. A support vector regression (SVR) model was trained with fragmentomic features to predict cfDNA concentration, using end motif frequencies (B) and both end motif and size profiles (C).











