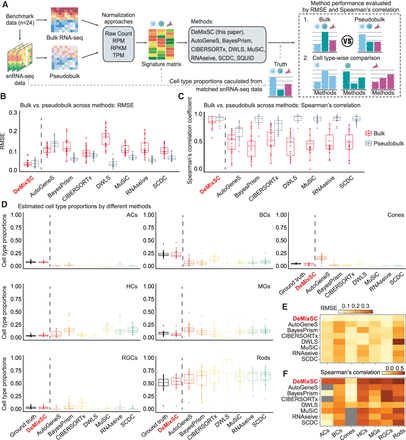
Comparing the estimation accuracy of DeMixSC to existing deconvolution methods. (A) Workflow for the deconvolution benchmarking design. We use benchmark data from retinal samples. The cell count proportions for each cell type are used as ground truth for the corresponding tissue samples. We assess the deconvolution performance of DeMixSC and seven existing methods for both bulk and pseudobulk mixtures. In addition to the raw counts, we also test RPM, RPKM, and TPM. The deconvolution performance is assessed by RMSE and Spearman's correlation coefficient. Note the results by SQUID are discussed in the text only. (B,C) Boxplots showing the deconvolution performance of eight deconvolution methods for the bulk and pseudobulk data. RMSE and Spearman's correlation coefficient values are calculated across seven major cell types for each sample, with gray denoting pseudobulk and red denoting real bulk. Smaller RMSEs or larger Spearman's correlations indicate a higher accuracy in proportion estimation. (D) Boxplots showing the distributions of deconvolution estimates at the cell type level for all 24 retinal samples. Each color corresponds to a given deconvolution method, with black denoting the ground truth, and each panel corresponds to a given cell type. (E,F), An overview of deconvolution performance at the cell type level across the eight methods using RMSE and Spearman's correlation coefficient, respectively. Lighter colors correspond to lower RMSE or Spearman's correlation coefficient values. Gray indicates NA.











