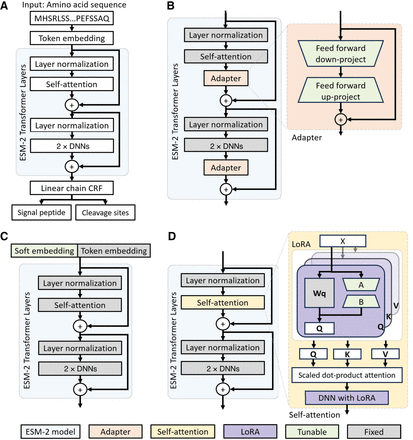
The architectures for the ESM-2 model and PEFT-SP using different PEFT modules. The light green modules are tunable during training, whereas the gray modules are fixed. (A) The ESM-2 backbone model uses amino acid sequences for SP and CS prediction. (B) PEFT-SP using adapter tuning contains a bottleneck architecture. (C) PEFT-SP using prompt tuning appends soft embedding into token embedding. (D) PEFT-SP using LoRA adds trainable rank decomposition matrices into the self-attention layer.











