Table 1.
Significant epistatic trait-gene pairs
| Trait | Chr | Gene | Start (Mb) | End (Mb) | −log10P-value | −log10P-value (40 PC) | −log10P-value (Imputed) |
|---|---|---|---|---|---|---|---|
| Alanine aminotransferase | 22 | PNPLA3 | 44.320 | 44.342 | ≥13 | ≥13 | ≥13 |
| 22 | SAMM50 | 44.351 | 44.392 | 8.02 | 7.97 | 6.61 | |
| Apolipoprotein B | 19 | BCAM | 45.312 | 45.324 | 9.03 | 9.02 | 2.84 |
| 19 | APOE | 45.410 | 45.413 | ≥13 | ≥13 | ≥13 | |
| Aspartate aminotransferase | 22 | PNPLA3 | 44.320 | 44.342 | 11.92 | 11.91 | ≥13 |
| Creatinine | 1 | DNAJC16 | 15.856 | 15.895 | 7.05 | 7.07 | 6.42 |
| Direct bilirubin | 2 | SAG | 234.218 | 234.256 | 7.56 | 7.55 | 3.43 |
| 2 | DGKD | 234.263 | 234.378 | ≥13 | ≥13 | 7.02 | |
| 2 | USP40 | 234.386 | 234.474 | ≥13 | ≥13 | 4.51 | |
| 2 | UGT1A8 | 234.526 | 234.681 | ≥13 | ≥13 | ≥13 | |
| Eosinophil count | 11 | PRG3 | 57.144 | 57.148 | ≥13 | ≥13 | 5.17 |
| HDL cholesterol | 16 | CETP | 56.996 | 57.018 | 9.81 | 9.87 | 5.13 |
| Hemoglobin A1c | 10 | HK1 | 71.048 | 71.161 | 7.71 | 7.68 | 1.55 |
| LDL direct | 19 | APOE | 45.410 | 45.413 | 8.91 | 8.88 | 9.37 |
| Lipoprotein(a) | 6 | IGF2R | 160.390 | 160.526 | ≥13 | ≥13 | 8.31 |
| 6 | SLC22A2 | 160.638 | 160.680 | 8.54 | 8.53 | 7.77 | |
| 6 | SLC22A3 | 160.769 | 160.872 | ≥13 | ≥13 | 4.13 | |
| 6 | LPA | 160.953 | 161.085 | ≥13 | ≥13 | ≥13 | |
| 6 | PLG | 161.123 | 161.174 | 8.54 | 8.54 | 12.11 | |
| 6 | AGPAT4 | 161.558 | 161.653 | ≥13 | ≥13 | 3.81 | |
| Mean corpuscular hemoglobin | 22 | TMPRSS6 | 37.462 | 37.500 | 8.72 | 8.75 | 2.69 |
| Mean sphered cell volume | 1 | OR10Z1 | 158.576 | 158.577 | 7.39 | 7.4 | 7.71 |
| 1 | SPTA1 | 158.581 | 158.656 | 9.31 | 9.32 | 7.21 | |
| Monocyte count | 22 | ADA2 | 17.662 | 17.691 | 8.8 | 8.8 | 2.97 |
| Platelet distribution width | 20 | TUBB1 | 57.595 | 57.600 | ≥13 | ≥13 | ≥13 |
| 20 | EDN3 | 57.876 | 57.900 | 9.45 | 9.5 | 0.01 | |
| SHBG | 17 | TNK1 | 7.286 | 7.292 | 7.37 | 7.35 | 2.42 |
| Urate | 1 | DNAJC16 | 15.856 | 15.895 | 8.26 | 8.27 | 11.42 |
| 4 | SLC2A9 | 9.828 | 10.028 | ≥13 | ≥13 | ≥13 | |
| 4 | WDR1 | 10.077 | 10.118 | ≥13 | ≥13 | ≥13 | |
| 4 | MEPE | 88.756 | 88.768 | 7.36 | 7.28 | 0.57 | |
| Urea | 1 | MUC1 | 155.159 | 155.163 | 8.33 | 8.34 | 7.14 |
-
We report trait-gene pairs with statistically significant epistatic effects (
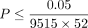 accounting for the number of genes and traits tested). The −log10 P-values were reported in entry −log10 P-value, with a precision level bounded by 13. We report the P-values at these trait-gene pairs when the top 40 PCs were regressed out (−log10 P-value [40 PC]) and when analyzing imputed genotypes (−log10 P-value [Imputed]).
accounting for the number of genes and traits tested). The −log10 P-values were reported in entry −log10 P-value, with a precision level bounded by 13. We report the P-values at these trait-gene pairs when the top 40 PCs were regressed out (−log10 P-value [40 PC]) and when analyzing imputed genotypes (−log10 P-value [Imputed]).











