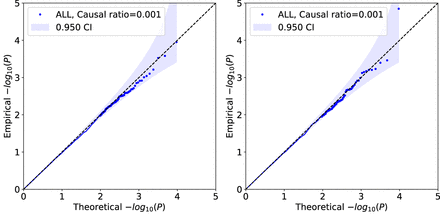
Calibration of QuadKAST when causal variants are imperfectly tagged. We simulated an additive phenotype using imputed genotypes over N = 50 K individuals, 4,824,392 SNPs (of which we randomly selected 1% of the SNPs to be causal). We then tested QuadKAST on the SNPs genotyped on the UKB array with sets defined by protein-coding genes. We applied QuadKAST with (left) and without (right) self-interactions.











