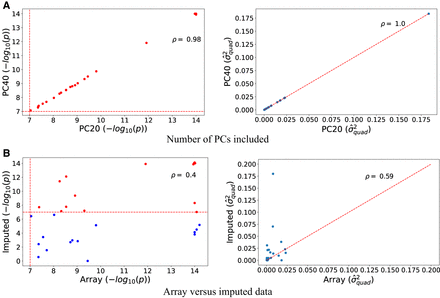
Robustness analysis. We report the Spearman's correlation (ρ) between estimators of 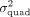 obtained by QuadKAST under different scenarios. (A) We report the correlation of the negative log10 P-values and the estimates of
obtained by QuadKAST under different scenarios. (A) We report the correlation of the negative log10 P-values and the estimates of 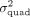 for all the significant epistatic trait-gene pairs when we include the top 20 PCs (default choice) versus the top 40 PCs
into the set of covariates. (B) We report the correlation of the P-values and the variance component estimates between the array data set and the imputed data set where the imputed data set
contains 4,824,392 SNPs whereas the array data set contains 459,792 SNPs. For ease of visualization,
for all the significant epistatic trait-gene pairs when we include the top 20 PCs (default choice) versus the top 40 PCs
into the set of covariates. (B) We report the correlation of the P-values and the variance component estimates between the array data set and the imputed data set where the imputed data set
contains 4,824,392 SNPs whereas the array data set contains 459,792 SNPs. For ease of visualization, 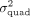 estimates of more than 0.2 have been excluded from the display.
estimates of more than 0.2 have been excluded from the display.











