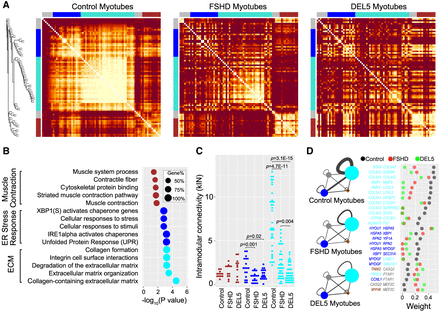
Weighted gene coexpression network analysis (WGCNA) of the non-DUX4-targets reveals gene network disruptions in FSHD and DEL5 myotubes. (A) Heatmaps depict the topological overlap matrix (TOM) among the 77 non-DUX4-target genes in the analysis. This matrix indicates similarity between genes by analyzing the extent to which individual genes coexpress with the same subset of genes (including each other). This results in a more robust similarity measure than pure correlation matrices. Lighter colors represent higher overlap, and darker colors represent lower overlap. The three heatmaps are plotted with the same order of genes based on the gene dendrogram of control myotubes. The modules of control are shown along the left side and the top. (B) The bubble plot shows Gene Ontology (GO) enrichment analysis of genes in the modules of control using the online tool g:Profiler. The bubble colors correspond to the module colors in A. The x-axis displays log10-adjusted P-values used Bonferroni correction and a threshold of 0.05. Bubble size indicates, in each term, the percentage of total genes in the given module. (C) The intramodular k-values of the genes in the blue and turquoise modules, identified in control myotubes, are decreased significantly in the FSHD and DEL5 myotubes. (D) The intramodular and intermodular connections of genes change significantly in FSHD and DEL5 myotubes compared with the control. The width of the edges and self-loops is proportional to sum of the weights (topological overlap values) of the edges, thresholded at 0.1. The module size indicates the number of genes within it. Some gene pairs are shown on the right panel as examples.











