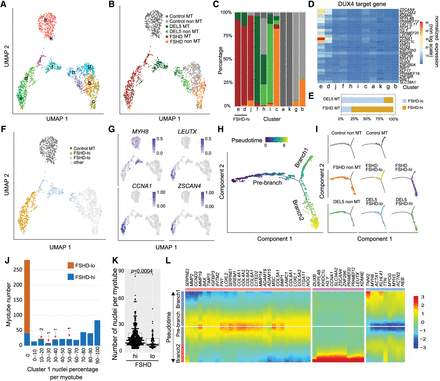
Clustering and pseudotime analysis identify a developmental bifurcation in the myoblast population, associated with a distinct reduction in myotube volume. (A) UMAP plots of MERFISH data from 987 myotubes and 629 nonmyotube regions. The plots are colored according to the clusters determined by the shared nearest neighbor (SNN) algorithm after batch correction using the Harmony algorithm. (B) UMAP from A colored by cell types, including the myotubes and nonmyotube regions of the differentiated control, FSHD and DEL5 cells. (C) Bar plot of the proportion of cell types in each cluster from the UMAP, colored by annotated cell types as in B. The myotubes for FSHD-hi and FSHD-lo are determined by DUX4 target gene expression levels. (D) Average expression profiles of the DUX4 target genes in each cluster. (E) The percentage of FSHD-hi and FSHD-lo cells in FSHD and DEL5 myotubes. (F) UMAP from A colored by designation of FSHD-hi or FSHD-lo versus control myotubes and remaining cells. (G) UMAP from A colored by normalized and scaled expression of example marker genes. (H) Trajectory analysis of batch-corrected MERFISH data using Monocle. The top 18 genes with the highest variability in expression within the myotube and nonmyotube regions are used to construct the pseudotime tree. Myotubes or nonmyotube regions on the tree are colored by pseudotime. (I) The pseudotime tree from H is colored by cell types and displayed individually. (J) Bar plot of the distribution of two types of myotubes, FSHD-hi and FSHD-lo, based on the percentage of cluster 1 nuclei in each myotube. The x-axis represents the range of cluster 1 nuclei percentages. The majority of FSHD-lo myotubes have no cluster 1 nuclei, with a few exceptions. To highlight these rare occurrences, the FSHD-lo myotube numbers (1 or 2) are indicated on the bars with red arrows. (K) The number of nuclei per myotube quantified in the FSHD-hi and FSHD-lo groups of FSHD myotubes. (L) The pseudotime heatmap displays the expression level of significantly changed genes (P < 0.01) in the two branches for comparison. Color key from blue to red indicates relative expression levels from low to high.











