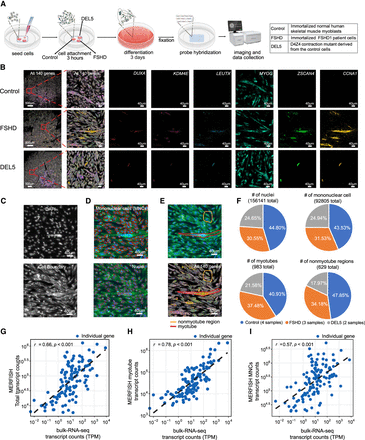
Spatial transcriptomic profiling of differentiated control and FSHD patient/mutant myoblasts by MERFISH. (A) Schematic diagram of the sample preparation process for MERFISH experiments and the genotypes used in the study. (B) Spatial distribution of detected genes by MERFISH. (Column 1) Illustration of the original sample image (cropped). All 140 genes used in the study are illustrated. (Column 2) The magnified example region corresponds to the red square area on the left. (Columns 3–8) Distribution of six example genes in the magnified area. (C) Staining results of the magnified region in B. (Top) DAPI staining of the nuclei in the region; (bottom) cell boundary staining of the region that illustrates the cytoplasm. (D) Automatically segmented mononuclear cells (MNCs) and nuclei inside the magnified region of B. The background is the overlay of cell boundary staining and DAPI staining. (Top) Detected MNCs inside the area. MNCs are circled with red lines. (Bottom) Detected nuclei within the region. Nuclei are circled with red lines. (E) Manually labeled myotube and nonmyotube regions (top), and the corresponding spatial transcriptome illustration with all 140 genes (bottom) in the magnified region in B. (F) Composition of nuclei, MNCs, myotube, and nonmyotube of the different genotypes in this study. (G–I) Comparison of MERFISH results with bulk RNA-seq of control myoblasts at day 3 of differentiation. Comparisons between the log-transformed total transcript counts from MERFISH and total transcript counts of all myocytes from bulk RNA-seq (G), intra-myotube transcript counts from MERFISH and total counts from bulk RNA-seq (H), and intra-MNC transcript counts from MERFISH and total counts from bulk RNA-seq (I). Overall MERFISH detections show a high level of correlation with the detection results from bulk RNA-seq and scRNA-seq methods (G) Pearson's r = 0.66, P < 0.001; (H) Pearson's r = 0.78, P < 0.001; (I) Pearson's r = 0.57, P < 0.001; N = 140 genes.











