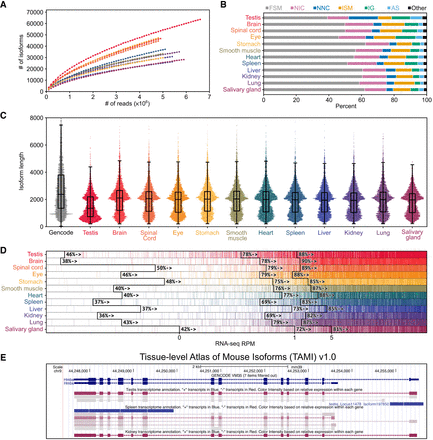
Characterization of tissue-level transcriptomes. (A) Isoform saturation curves for each tissue. (B) Isoform category distributions for each tissue as determined by SQANTI3 (full splice match [FSM], novel in catalog [NIC], novel not in catalog [NNC], incomplete splice match [ISM], intergenic [IG], antisense [AS]). (C) Isoform length distribution for each tissue compared to GENCODE vM30 basic protein-coding transcripts. (D) For each tissue, genes are rank-ordered based on their expression level in Illumina RNA-seq data. Genes are marked by a vertical colored line if at least a single isoform is assigned to them in the respective tissue; 0, 1, and 5 RPM levels in RNA-seq data are indicated by black lines. The percentage of genes expressed higher than that RPM with at least one isoform assigned to them is shown adjacent to those lines. (E) Screenshot of the testis, spleen, and kidney tracks from the TAMI trackhub as displayed on the UCSC Genome Browser.











