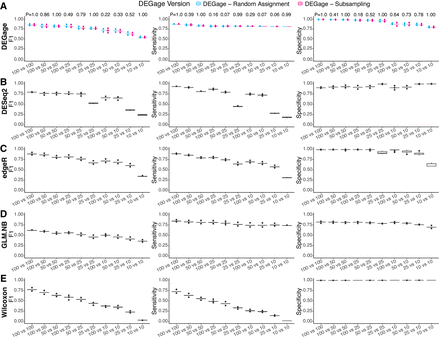
Performance evaluation of DEGage on imbalanced data sets. (A) Comparison of DEGage's performance between the random assignment and the subsampling strategy. F1, sensitivity, and specificity scores were calculated for 10 combinations of balanced and imbalanced data sets. Each boxplot represents the scores across 10 replicates. The P-values were calculated using a two-sided Student's t-test. Performance of DESeq2 (B), edgeR (C), GLM.NB (D), and Wilcoxon test (E) on imbalanced data sets. Detailed scores are presented in Supplemental Tables S4–S6.











