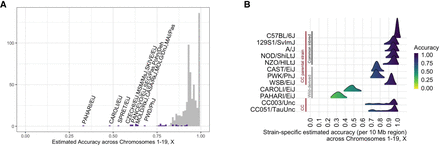
Imputation accuracy. (A) Histogram of estimated strain-specific imputation accuracies across Chromosomes 1–19 and X for 657 strains. Wild-derived strain accuracies are colored purple. Strains with accuracies below 0.70 are annotated. (B) Ridgeline plots of the strain-specific distribution of imputation accuracies across Chromosomes 1–19 and X (all 657 strains in Supplemental Fig. S1). Shown are the common inbred CC founders (C57BL/6J, 129S1/SvImJ, A/J, NOD/ShiLtJ, NZO/HILtJ), wild-derived CC founders (CAST/EiJ, PWK/PhJ, WSB/EiJ), two CC strains (CC003/Unc, and CC051/TauUnc), and two more-distant wild-derived strains (CAROLI/EiJ, PAHARI/EiJ).











