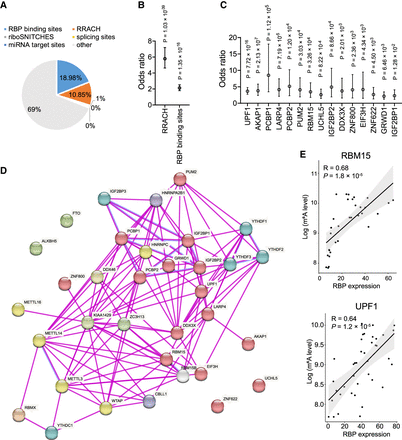
Prediction of trans-factors involved in ASm6A regulation. (A) Proportion of the putative functional variants for ASm6As that overlap with RNA-related features, including RBP binding sites, RRACH sequence, miRNA target sites, riboSNITCHES variants, and splicing sites. (B) Enrichment of the putative functional variants for ASm6As versus random control variants in RRACH sequence and RBP binding sites by Fisher's exact test. (C) Enrichment of the putative functional variants for ASm6As in binding sites of different RBPs by Fisher's exact test. (D) Protein–protein interactions between the RBPs in C and known m6A writers, readers, and erasers. The red sphere indicates the RBPs in C. (E) Correlation between RBP expression and aggregative m6A level of its targets for RBM15 (top) and UPF1 (bottom).











