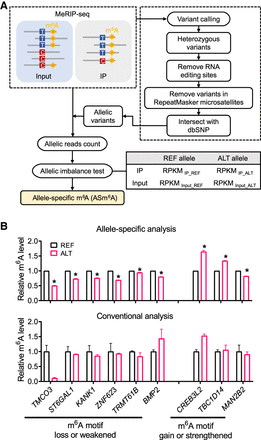
ASm6As analysis pipeline generation and evaluation. (A) Pipeline for ASm6As identification. Allelic variants were identified by variant calling and stringent filtering. Allelic reads were then counted from sequencing data of input and IP, and ASm6As were predicted using a Fisher's exact test. (B) Comparison between allele-specific analysis and conventional analysis. In the m6A motif loss or weakened group, the potential m6A motifs (reference to alternative allele) were found in TMCO3 (AAACT to AAATT), ST6GAL1 (GGACT to GGGCT), KANK1 (GAACT to GAGCT), ZNF623 (GGACC to AGACC), TRMT61B (GAACT to GAGCT), and BMP2 (AGACT to AGTCT). In the m6A motif gain or strengthen group, they were found in CREB3L2 (GCACT to GAACT), TBC1D14 (CGACT to GGACT), and MAN2B2 (GAGCA to GAACA). The relative m6A level was calculated by dividing the reads of IP by input, then normalized to the reference allele. Data are mean ± SD from two independent experiments. In the allele-specific analysis, (*) P < 0.05 in both replicates as assessed by Fisher's exact test.











