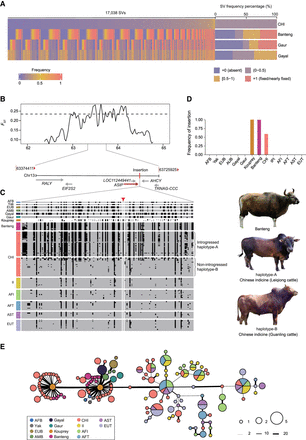
Candidate introgressed SV of Chinese indicine cattle and a case of adaptive introgressed haplotype associated with coat color. (A) The heatmap shows the frequency distribution of 3136 population-stratified SVs in Chinese indicine population and shared with the banteng, gaur, and gayal populations. Each vertical line represents an SV in the corresponding population, and the color ranging from dark blue to red represents frequency varying from zero to one. The bar plot at the right of the heatmap shows the percentage of the SV frequency in the corresponding population. (B) The schematic map indicates the selective signal (FST) and the genomic location on Chromosome 13 (62.37–64.83 Mb). The horizontal dashed line of SNPs (the 1% right tail of the empirical FST distribution) was identified as the selected regions for Chinese indicine cattle and Indian indicine cattle. (C) The haplotypes of the insertion and its flanking SNP in each cattle population. The missing genotypes of the ancient kouprey sample in the haplotype are indicated by blank blocks. (D) The frequency of the 6.3-kb insertion in each cattle group. (E) Haplotype network based on pairwise differences around the insertion region; the haplotypes with the insertion (introgressed haplotype-A) in Chinese indicine population are predominantly clustered with kouprey-like origin, whereas other haplotypes without the insertion (nonintrogressed haplotype-B) tend to cluster with domestic cattle population. (AFB) African buffalo; (EUB) European bison; (AMB) American bison; (CHI) Chinese indicine; (II) Indian indicine; (AFI) African indicine; (AFT) African taurine; (AST) Asian taurine; (EUT) European taurine.











