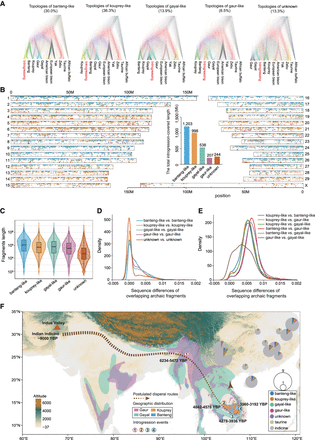
The genomic landscape of introgression in 20 Chinese indicine partially phased genomes. (A) DensiTree (Bouckaert 2010) visualization of five tree topologies: banteng-like, kouprey-like, gayal-like, gaur-like, and unknown origin. The value in brackets is the percentage of the same topologies. (B) The genome-wide distribution of archaic fragments in 20 Chinese indicine haplotypes, colored according to the topologies: banteng-like (blue), kouprey-like (golden), gayal-like (green), gaur-like (purple), and unknown (dark orange). The inset shows the total introgression-covered length of each origin. (C) Length distribution of banteng-like, kouprey-like, gayal-like, gaur-like, and unknown archaic fragments. The sequence differences between overlapping archaic fragments from the same origin are shown in D (KDE plot) and from different origins are shown in E. (F) The postulated dispersal routes of Chinese indicine cattle and admixture with wild Bos species. Introgression events with different wild Asian Bos species are marked with numbers in colored circles. The pie charts show the geographical distribution and genetic make-up of 15 Chinese indicine cattle breeds (45 samples), with each circle sized proportionally to the number of samples per breed.











