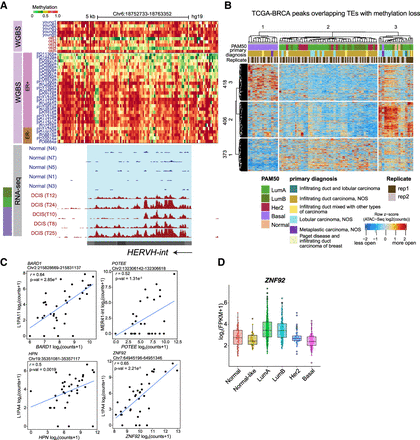
Transposable elements (TEs) with age-dependent methylation loss are activated in breast cancer. (A) A representative HERVH element with variable loss of methylation in old samples and breast cancer (BC) samples. The estrogen receptor (ER) expression status of the tumors is shown for the BC WGBS tracks. The bottom tracks are RNA-seq tracks from DCIS samples showing cryptic transcription of the HERVH element. All RNA-seq tracks are scaled to the same value, and the PAM50 subtype is indicated on the left. (B) Heatmap showing signals across 74 TCGA-BRCA samples for ATAC-seq peaks that overlap TEs hypomethylated with age. ATAC-seq peaks (rows) and TCGA BRCA samples (columns) are split into three clusters, each using k-means clustering. (C) Dot plot of RNA-seq counts at indicated TE and gene pairs that are correlated across DCIS samples. Each dot represents an individual sample. Spearman's correlation coefficients and P-values are indicated. (D) Expression of ZNF92 in normal tissues and five PAM50 subtypes of breast tumors from the TCGA BRCA collection (P-value luminal A [LumA] vs. basal and luminal B [LumB] vs. basal; P-value < 2.2 × 10−16, using a two-tailed Wilcoxon signed-rank test).











