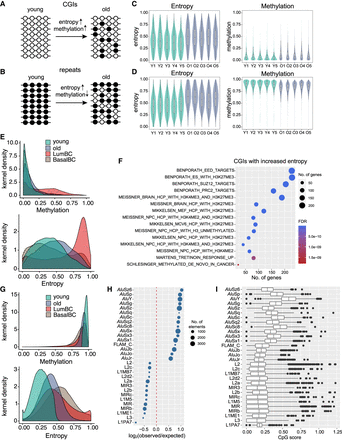
Increased methylation entropy with age in normal LEps predicts CpG island (CGI) methylation gain and Alu methylation loss in breast cancer. (A) Schematic depicting changes in entropy associated with stochastic methylation gain at CGIs. (B) Schematic depicting changes in entropy associated with stochastic methylation loss at fully methylated regions such as repeat elements. (C) Methylation and entropy levels at unmethylated CGIs with an increase in entropy. (D) Methylation and entropy levels at fully methylated regions with an increase in entropy. (E) Kernel density plots of methylation and entropy levels at unmethylated CGIs with an increase in entropy with age. Luminal breast cancer (LumBC) samples (red) show an increase in DNA methylation and entropy at these CGIs, whereas basal breast cancer (BasalBC) samples (brown) do not. (F) MSigDB gene sets enriched in CGI promoters that gain methylation entropy with age. BENPORATH gene sets represent PRC2 target genes identified in embryonic stem cells. (G) Kernel density plots of methylation and entropy levels at fully methylated regions with an increase in entropy with age. Both luminal breast cancer (red) and basal breast cancer samples (brown) show a decrease in DNA methylation with an increase in entropy at these regions. (H) TE subfamilies enriched at regions that show increased entropy with methylation loss. The size of the dot represents the number of elements. (I) Boxplot shows the CpG score of TEs that show stochastic methylation loss. TEs are grouped by subfamily.











