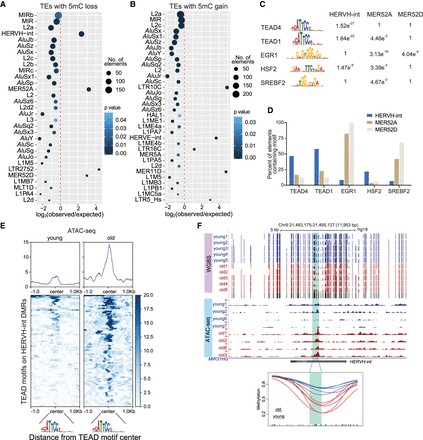
Age-dependent methylation changes occur at TE subfamilies that contain lineage-specific transcription factor binding sites. (A,B) TE subfamilies enriched at DMRs with methylation loss (A) and methylation gain (B). The size of the dot represents the number of elements with methylation loss or gain. All TEs with more than 10 copies overlapping DMRs are shown for completeness. (C) Transcription factor binding motifs enriched in HERVH-int, MER52A, and MER52D elements with methylation loss. The P-values for enrichment over scrambled sequences are shown. (D) Percentage of elements containing indicated transcription factor motifs across TEs with methylation loss. (E) ATAC-seq profile at TEAD motifs within HERVH-int elements showing methylation loss with age. (F) Genome Browser tracks show a representative example of a HERVH-int element losing methylation in old samples. The bottom tracks are from ATAC-seq data showing a gain of accessibility at the DMR. The bottom panel shows kernel-smoothed line plots of the highlighted region.











