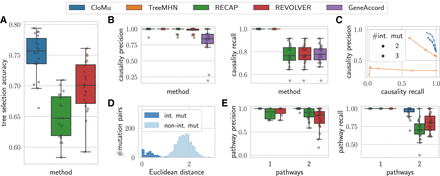
CloMu outperforms existing methods on several prediction tasks on simulated data with known ground truth. (A) Tree selection accuracy measures the ability to correctly identify the ground-truth tree from a set 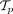 of possible trees generated from simulated bulk sequencing for each patient p. (B) Causality precision and recall measure the ability to determine positive causal relationships between ordered pairs of mutations.
Panels A and B show results for simulations I-a. (C) On simulations I-b and I-c, causality precision and recall measure the ability to identify causation and inhibition between
pairs of mutations in the presence of mutation interchangeability. (D) On simulations II, interchangeability detection shows that the latent representations generated by our model are meaningful,
accurately encapsulating mutation similarity. (E) On simulations II, pathway precision and recall measure the ability to determine evolutionary pathways in the presence of
both interchangeability and complex multimutation interactions.
of possible trees generated from simulated bulk sequencing for each patient p. (B) Causality precision and recall measure the ability to determine positive causal relationships between ordered pairs of mutations.
Panels A and B show results for simulations I-a. (C) On simulations I-b and I-c, causality precision and recall measure the ability to identify causation and inhibition between
pairs of mutations in the presence of mutation interchangeability. (D) On simulations II, interchangeability detection shows that the latent representations generated by our model are meaningful,
accurately encapsulating mutation similarity. (E) On simulations II, pathway precision and recall measure the ability to determine evolutionary pathways in the presence of
both interchangeability and complex multimutation interactions.











