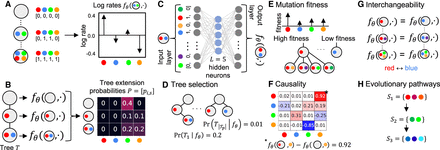
Overview of CloMu. (A) Using the independent clonal evolution assumption, our model determines a log rate 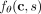 of any clone c ∈ {0, 1}m acquiring a mutation s ∈ [m]. (B) This in turn enables us to compute probabilities P = [pi,s] that the next mutation to occur on a tree T is mutation s at node/clone ci. The resulting Independent Clonal Evolution problem seeks model parameters θ that maximize the data probability
of any clone c ∈ {0, 1}m acquiring a mutation s ∈ [m]. (B) This in turn enables us to compute probabilities P = [pi,s] that the next mutation to occur on a tree T is mutation s at node/clone ci. The resulting Independent Clonal Evolution problem seeks model parameters θ that maximize the data probability  of a cohort of trees for n patients. (C) CloMu represents
of a cohort of trees for n patients. (C) CloMu represents 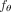 using a low-parameter, two-layer neural network that is trained via reinforcement learning. We use the model for five prediction
tasks: (D) tree selection for each patient, (E) determination of mutation fitness, (F) causality inference for pairs of mutations, (G) identification of interchangeable mutations, and (H) identification of their evolutionary pathways.
using a low-parameter, two-layer neural network that is trained via reinforcement learning. We use the model for five prediction
tasks: (D) tree selection for each patient, (E) determination of mutation fitness, (F) causality inference for pairs of mutations, (G) identification of interchangeable mutations, and (H) identification of their evolutionary pathways.











