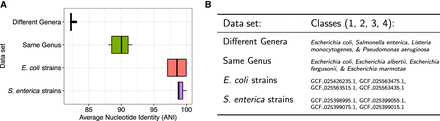
Summary of reference data sets of increasing difficulty for read classification. (A) Sequence homology, measured as average nucleotide identity (ANI) for all across-class pairs of sequences. ANI was estimated with fastANI (Jain et al. 2018). (B) List of the specific species and strains used for classes 1, 2, 3, and 4 for each of the four data sets. In the case of “different genera” and “same genus,” we used 10 genomes per class. In the case of “E. coli strains” and “S. enterica strains,” we used a single genome for each strain.











