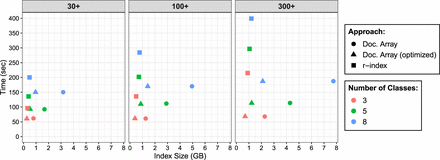
Query time and index size when performing document listing queries using the document array profiles and the r-index. We varied the size of the database increasing from 30 bacterial genomes to 300 bacterial genomes. For each species/class, we would simulate nanopore reads at 95% accuracy and extract 1 million maximal-exact matches (MEMs) to query the data-structures. Therefore, for the three-class, five-class, and eight-class indexes, we queried them with 3 million, 5 million, and 8 million MEMs, respectively. This explains why the query time would increase for the indexes containing more classes.











