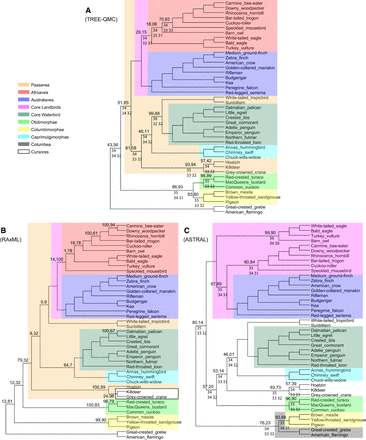
Avian UCE data. (A) Tree estimated from UCE gene trees using TREE-QMC-n2/wQFM. (B) Tree estimated from concatenated UCE alignment using RAxML. (C) Tree estimated from UCE gene trees using ASTRAL-III/FASTRAL. Above the branch, we show support values X, Y, where X is estimated using ASTRAL's local posterior probability (multiplied by 100) and Y is computed using RAxML's bootstrap support for B and using MLBS for A and C. Support values are only shown when X is <100. Below the branch, we show the quartet support (the two values below it correspond to quartet support for the two alternative resolutions of the branch). Taxa outside of Neoaves are not shown as all methods recovered the same topology outside of Neoaves.











